

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.5 + phase: 0
(221 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
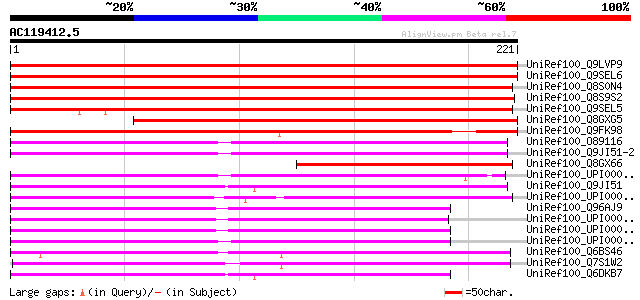
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LVP9 Vesicle transport v-SNARE 13 [Arabidopsis thali... 339 3e-92
UniRef100_Q9SEL6 Vesicle transport v-SNARE 11 [Arabidopsis thali... 338 6e-92
UniRef100_Q8S0N4 Putative vesicle transport v-SNARE protein [Ory... 323 3e-87
UniRef100_Q8S9S2 Similar to vesicle transport protein [Oryza sat... 301 1e-80
UniRef100_Q9SEL5 Vesicle transport v-SNARE 12 [Arabidopsis thali... 286 2e-76
UniRef100_Q8GXG5 Putative vesicle transport protein [Arabidopsis... 256 3e-67
UniRef100_Q9FK98 V-SNARE AtVTI1a-like protein [Arabidopsis thali... 175 7e-43
UniRef100_O89116 Vesicle transport through interaction with t-SN... 133 4e-30
UniRef100_Q9JI51-2 Splice isoform 1 of Q9JI51 [Rattus norvegicus] 129 4e-29
UniRef100_Q8GX66 Putative v-SNARE AtVTI1b [Arabidopsis thaliana] 126 4e-28
UniRef100_UPI00003606EC UPI00003606EC UniRef100 entry 125 1e-27
UniRef100_Q9JI51 Vesicle transport through interaction with t-SN... 124 2e-27
UniRef100_UPI00003C2589 UPI00003C2589 UniRef100 entry 120 3e-26
UniRef100_Q96AJ9 Vesicle transport through interaction with t-SN... 120 3e-26
UniRef100_UPI000036E95D UPI000036E95D UniRef100 entry 118 1e-25
UniRef100_UPI00003AE3EF UPI00003AE3EF UniRef100 entry 117 2e-25
UniRef100_UPI00004364AB UPI00004364AB UniRef100 entry 111 2e-23
UniRef100_Q6BS46 Similar to CA4631|CaVTI1 Candida albicans CaVTI... 111 2e-23
UniRef100_Q7S1W2 Hypothetical protein [Neurospora crassa] 110 3e-23
UniRef100_Q6DKB7 VTI1A protein [Xenopus laevis] 109 6e-23
>UniRef100_Q9LVP9 Vesicle transport v-SNARE 13 [Arabidopsis thaliana]
Length = 221
Score = 339 bits (869), Expect = 3e-92
Identities = 166/221 (75%), Positives = 196/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FE YERQYCE+SANLSKKCT+A AL+GEQKKQ +SE+KSG++EAEAL++KMDLEAR
Sbjct: 1 MSQGFERYERQYCEISANLSKKCTSAIALDGEQKKQNLSEIKSGVEEAEALVKKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMADT+TASADQR
Sbjct: 61 NLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMADTLTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH TLHGVDDN+G
Sbjct: 121 SRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHETLHGVDDNVG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 181 KSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 221
>UniRef100_Q9SEL6 Vesicle transport v-SNARE 11 [Arabidopsis thaliana]
Length = 221
Score = 338 bits (867), Expect = 6e-92
Identities = 166/221 (75%), Positives = 195/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+VF+GYERQYCELSA+LSKKC++A +L+GEQKKQK+SE+KSG++ AE LIRKMDLEAR
Sbjct: 1 MSDVFDGYERQYCELSASLSKKCSSAISLDGEQKKQKLSEIKSGLENAEVLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLRE+KSDLNN K+EVK+I SG LN +ARDELLE+GMADT TASADQR
Sbjct: 61 TLPPNLKSSLLVKLREFKSDLNNFKTEVKRITSGQLNAAARDELLEAGMADTKTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM STERL +T +RVKDSRRTM+ETEE+GVSILQDLH QRQSLL AH TLHGVDDNIG
Sbjct: 121 ARLMMSTERLGRTTDRVKDSRRTMMETEEIGVSILQDLHGQRQSLLRAHETLHGVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T+M+RRMNKNKW IG I++ L+ AI ILYFKL K
Sbjct: 181 KSKKILTDMTRRMNKNKWTIGAIIIALIAAIFIILYFKLTK 221
>UniRef100_Q8S0N4 Putative vesicle transport v-SNARE protein [Oryza sativa]
Length = 221
Score = 323 bits (827), Expect = 3e-87
Identities = 157/219 (71%), Positives = 195/219 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS VFEGYERQYCE+SA+L +KCT A+AL+GE+KKQK+SE++SG++EAE+LIRKMDLEAR
Sbjct: 1 MSEVFEGYERQYCEVSASLFRKCTTASALDGEKKKQKLSEIQSGVEEAESLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
S+QP++K LLAKLREYKSDLNNLKSE+K+I + N + R+ELLESG+ADT+ AS DQR
Sbjct: 61 SLQPSVKAGLLAKLREYKSDLNNLKSELKRISAPNARQATREELLESGLADTLAASTDQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM +TERLN++ +++K+SRRT+LETEELGVSILQDLH QRQSLLHAH TLHGVDDN+G
Sbjct: 121 GRLMMTTERLNQSNDKIKESRRTILETEELGVSILQDLHQQRQSLLHAHTTLHGVDDNVG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KSKKI+ MS+RM++NKWIIG I+ LV+AI+ ILYFKL
Sbjct: 181 KSKKILAAMSKRMDRNKWIIGGIIAALVLAILLILYFKL 219
>UniRef100_Q8S9S2 Similar to vesicle transport protein [Oryza sativa]
Length = 221
Score = 301 bits (770), Expect = 1e-80
Identities = 145/220 (65%), Positives = 186/220 (83%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
M++VFEGYERQYCE+SA+L++KCTAA+AL GE+ KQK SE+KSGID AEALI+KMDLEAR
Sbjct: 1 MADVFEGYERQYCEISASLARKCTAASALQGEKLKQKASEIKSGIDGAEALIKKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ QP+++ LLAKLREYKSDLNNLK +K++ +GN +R+ELLESGMA+T+ SADQ+
Sbjct: 61 NQQPSVRAGLLAKLREYKSDLNNLKGTLKRVTTGNAQQGSREELLESGMAETLGVSADQK 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RL+ TE+ NKT +R++DS RTMLETE+LGVS+LQDLH QR+ L+HAH TL VDDNIG
Sbjct: 121 SRLLRITEKQNKTTDRIRDSHRTMLETEDLGVSLLQDLHQQRERLIHAHGTLDNVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLV 220
KS++IM M RRM++NKWIIG I+ +LV+ I+ ILYFK V
Sbjct: 181 KSRRIMGAMVRRMDRNKWIIGFIIALLVLVILVILYFKFV 220
>UniRef100_Q9SEL5 Vesicle transport v-SNARE 12 [Arabidopsis thaliana]
Length = 223
Score = 286 bits (733), Expect = 2e-76
Identities = 144/221 (65%), Positives = 182/221 (82%), Gaps = 2/221 (0%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATAL-NGEQKKQKVSE-VKSGIDEAEALIRKMDLE 58
MS+VFEGYERQYCELS NLS+KC +A+ L NGE+KK K++E +KSGIDEA+ LIRKMDLE
Sbjct: 1 MSDVFEGYERQYCELSTNLSRKCHSASVLSNGEEKKGKIAEQIKSGIDEADVLIRKMDLE 60
Query: 59 ARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASAD 118
ARS+QP+ K V L+KLREYKSDLN LK E K++ S + PS+R+EL+ESGMAD SAD
Sbjct: 61 ARSLQPSAKAVCLSKLREYKSDLNQLKKEFKRVSSADAKPSSREELMESGMADLHAVSAD 120
Query: 119 QRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDN 178
QR RL S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD
Sbjct: 121 QRGRLAMSVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDA 180
Query: 179 IGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
I KSKK++T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 181 IDKSKKVLTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 221
>UniRef100_Q8GXG5 Putative vesicle transport protein [Arabidopsis thaliana]
Length = 167
Score = 256 bits (654), Expect = 3e-67
Identities = 126/167 (75%), Positives = 147/167 (87%)
Query: 55 MDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMT 114
MDLEAR++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMADT+T
Sbjct: 1 MDLEARNLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMADTLT 60
Query: 115 ASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHG 174
ASADQR+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH TLHG
Sbjct: 61 ASADQRSRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHETLHG 120
Query: 175 VDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
VDDN+GKSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 121 VDDNVGKSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 167
>UniRef100_Q9FK98 V-SNARE AtVTI1a-like protein [Arabidopsis thaliana]
Length = 212
Score = 175 bits (444), Expect = 7e-43
Identities = 98/222 (44%), Positives = 142/222 (63%), Gaps = 11/222 (4%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MSNVF Y++QY ELS NLSKKC+ A +L G +KK+K+SE+ ++ AE LI KMD A
Sbjct: 1 MSNVFHEYDQQYSELSINLSKKCSLAFSLKGGEKKEKLSEITYDLENAEKLISKMDHAAS 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTAS-ADQ 119
++ PNIK +LL KL+E KS L L++E+K+ S NL + R+E+LE+ A ADQ
Sbjct: 61 NLPPNIKSILLEKLKESKSSLKRLRNEIKRNTSENLKVTTREEVLEAEKAVCKDMDLADQ 120
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R+RLM STE L +T E +KDS+R +LETE +G+SIL++L Q++SL ++ LH +DD +
Sbjct: 121 RSRLMKSTEGLVRTREMIKDSQRKLLETENIGISILENLQRQKESLQNSQAMLHEIDDTV 180
Query: 180 GKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
+S+ I+ ++ + + II FKLVK
Sbjct: 181 KESRSIVRSIKIKE----------FFTVTAPIIIYFLFKLVK 212
>UniRef100_O89116 Vesicle transport through interaction with t-SNAREs homolog 1A [Mus
musculus]
Length = 217
Score = 133 bits (334), Expect = 4e-30
Identities = 67/217 (30%), Positives = 128/217 (58%), Gaps = 5/217 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRS-----RIAYSDEVRNELLGDAGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A + L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARDRLRDADANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
KS +I+T M RR+ +N+ ++ + +++V+AI+ + F
Sbjct: 176 KSSRILTGMLRRIIQNRILLVILGIIVVIAILTAIAF 212
>UniRef100_Q9JI51-2 Splice isoform 1 of Q9JI51 [Rattus norvegicus]
Length = 217
Score = 129 bits (325), Expect = 4e-29
Identities = 66/217 (30%), Positives = 126/217 (57%), Gaps = 5/217 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ + L+A ++ K + L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSADFEGYEQDFAVLTAEITSKISRVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRS-----RIAYSDEVRNELLGDAGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDRERIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
KS +I+T M RR+ +N+ ++ + +++V+ I+ + F
Sbjct: 176 KSSRILTGMLRRIIQNRILLVILGIIVVITILTAITF 212
>UniRef100_Q8GX66 Putative v-SNARE AtVTI1b [Arabidopsis thaliana]
Length = 97
Score = 126 bits (317), Expect = 4e-28
Identities = 59/94 (62%), Positives = 79/94 (83%)
Query: 126 STERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKI 185
S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD I KSKK+
Sbjct: 2 SVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDAIDKSKKV 61
Query: 186 MTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
+T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 62 LTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 95
>UniRef100_UPI00003606EC UPI00003606EC UniRef100 entry
Length = 217
Score = 125 bits (313), Expect = 1e-27
Identities = 70/219 (31%), Positives = 127/219 (57%), Gaps = 10/219 (4%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FE YE+ + ++A ++ K L+GE+K Q V V ++E L+ +MDLEAR
Sbjct: 1 MSSDFEAYEQDFGTITAEITNKIGRIPKLSGEEKTQLVLNVDKQLEEVRELLEQMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ + + ++L+ YK ++ L+ + K+ + DE+ + D ++S +QR
Sbjct: 61 EIPIQSRAMYNSRLKSYKQEVEKLEKDFKRS-----RIAYSDEVRNELLGDDASSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TE+L ++ R++ + +ETE++G IL +LH+ R+ + A L D N+G
Sbjct: 116 AHLLDNTEKLERSSRRLQAGYQIAVETEQVGQEILTNLHTDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRRMNKNK---WIIGCIVLVLVVAIIAILY 216
KS +I+T M RR+ +N+ +I+G I+LV + I+AIL+
Sbjct: 176 KSSRILTGMLRRIIQNRVLIFILGAIILVTI--ILAILF 212
>UniRef100_Q9JI51 Vesicle transport through interaction with t-SNAREs homolog 1A
[Rattus norvegicus]
Length = 224
Score = 124 bits (310), Expect = 2e-27
Identities = 65/220 (29%), Positives = 127/220 (57%), Gaps = 4/220 (1%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ + L+A ++ K + L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSADFEGYEQDFAVLTAEITSKISRVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL---ESGMADTMTASA 117
+ P +G+ ++R YK ++ L+++ K+ + R+ELL + + +
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRSRIA-YSDEVRNELLGDAGNSSENQLIKLR 119
Query: 118 DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDD 177
++R L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D
Sbjct: 120 EERAHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDRERIQRARERLRETDA 179
Query: 178 NIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
N+GKS +I+T M RR+ +N+ ++ + +++V+ I+ + F
Sbjct: 180 NLGKSSRILTGMLRRIIQNRILLVILGIIVVITILTAITF 219
>UniRef100_UPI00003C2589 UPI00003C2589 UniRef100 entry
Length = 224
Score = 120 bits (301), Expect = 3e-26
Identities = 74/230 (32%), Positives = 123/230 (53%), Gaps = 18/230 (7%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS +F+ Y +L + +K A + NGE KK + + ++EA+ +I +MD+E +
Sbjct: 1 MSELFDSYASDLEQLKQGIRQKLDQAKSQNGEAKKSTLRRAEMELEEADEVISQMDIEVQ 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSAR-----------DELLESGM 109
+++ ++R +K DL L +V+ L+PS+R DE +
Sbjct: 61 GFPQSVRSRFSVQIRGFKQDLQTLTQQVR----AGLSPSSRGSRGAYDAAYADEETDLEA 116
Query: 110 ADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAH 169
AD TA+ R RL+ +T L R+++S R LETE+LG IL+DL SQR+ + ++
Sbjct: 117 ADGATAN---RQRLLHATASLEDGTRRLEESNRLALETEDLGADILRDLRSQREQIENSR 173
Query: 170 NTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
NTL D +I +S + ++ M RR + K + I++ LV+ I+ ILY KL
Sbjct: 174 NTLRQADQSIDRSSRTLSKMIRRAKQQKLVTVGIIVCLVLLILLILYNKL 223
>UniRef100_Q96AJ9 Vesicle transport through interaction with t-SNAREs homolog 1A
[Homo sapiens]
Length = 203
Score = 120 bits (301), Expect = 3e-26
Identities = 62/192 (32%), Positives = 111/192 (57%), Gaps = 5/192 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA+ L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEAKELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKR-----SRIAYSDEVRNELLGDDGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRR 192
KS +I+T M RR
Sbjct: 176 KSSRILTGMLRR 187
>UniRef100_UPI000036E95D UPI000036E95D UniRef100 entry
Length = 186
Score = 118 bits (296), Expect = 1e-25
Identities = 61/191 (31%), Positives = 110/191 (56%), Gaps = 5/191 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA+ L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEAKELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKR-----SRIAYSDEVRNELLGDDGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSR 191
KS +I+T M R
Sbjct: 176 KSSRILTGMLR 186
>UniRef100_UPI00003AE3EF UPI00003AE3EF UniRef100 entry
Length = 187
Score = 117 bits (293), Expect = 2e-25
Identities = 59/192 (30%), Positives = 112/192 (57%), Gaps = 5/192 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FE YE+ + L+A+++ + L G++KKQ V+ ++ ++EA L+ +M+LE R
Sbjct: 1 MSSDFESYEQDFAVLTADITGRIGRVPKLLGDEKKQMVANIEKQLEEARELLEQMELEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ +++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSSRMRSYKQEMGKLEADFKR-----SRIAYSDEVRNELLGDDGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRR 192
KS +I+T M RR
Sbjct: 176 KSSRILTGMLRR 187
>UniRef100_UPI00004364AB UPI00004364AB UniRef100 entry
Length = 187
Score = 111 bits (277), Expect = 2e-23
Identities = 61/192 (31%), Positives = 106/192 (54%), Gaps = 5/192 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FE YE+ + L+A ++ K L GE+K Q V V ++E L+ +MDLE R
Sbjct: 1 MSSDFEAYEQDFGTLTAEITNKIGRIPKLAGEEKTQLVLNVDKQLEEVRELMEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ + + ++L+ YK ++ L+ + K+ + DE+ + D ++S QR
Sbjct: 61 EIPIQSRAMYNSRLKSYKQEMEKLEKDFKRS-----RIAYSDEVRNELLGDDGSSSETQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G IL +LHS R+ + + + L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQVGQEILANLHSDREKIQRSRDRLRETDANLG 175
Query: 181 KSKKIMTNMSRR 192
KS +I+T M RR
Sbjct: 176 KSSRILTGMLRR 187
>UniRef100_Q6BS46 Similar to CA4631|CaVTI1 Candida albicans CaVTI1 v-SNARE
[Debaryomyces hansenii]
Length = 220
Score = 111 bits (277), Expect = 2e-23
Identities = 70/222 (31%), Positives = 119/222 (53%), Gaps = 8/222 (3%)
Query: 1 MSNVFEGYERQY-CELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEA 59
MS +F YE L SK ++A +++KQ + +++ DE ++ +M +E
Sbjct: 1 MSELFATYESDLQLALQEAKSKLAQISSADGKDERKQFLRAIETATDECLEVLDQMSIEV 60
Query: 60 RSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASA-- 117
+++ N + K+R+YKS + + K +KK+ L+ + EL + D +
Sbjct: 61 QNLPSNQRSTYNTKIRQYKSQVEDSKDRLKKL----LDDQDKYELFGNRYTDDDSHDDIH 116
Query: 118 -DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVD 176
QR +L+ + L +T ER++DS+R LETE +G +IL DL SQR+ + + NTL D
Sbjct: 117 DSQRKQLLNNNSSLERTSERLRDSQRIALETENIGGNILNDLRSQREQITGSRNTLMTAD 176
Query: 177 DNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
+ KS K + +MSRR+ NK+I I+ VL++ I +L K
Sbjct: 177 GYVDKSIKTLKSMSRRLTANKFISYAIIAVLILLIFLVLASK 218
>UniRef100_Q7S1W2 Hypothetical protein [Neurospora crassa]
Length = 228
Score = 110 bits (274), Expect = 3e-23
Identities = 64/222 (28%), Positives = 125/222 (55%), Gaps = 11/222 (4%)
Query: 2 SNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEARS 61
+ +F+ YE +Y + A+L +K + L GE +K +S + ++EA L+ +M++E ++
Sbjct: 11 TELFKHYETEYQLVQADLVQKLDQISELTGEPRKAALSTAERALEEAGELLDQMNMEKQN 70
Query: 62 MQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASA---- 117
+ + + + +LR+Y+SD++ + +++ + + R L D +S+
Sbjct: 71 VPSSSRTAINKRLRDYRSDVDKYRRKLQTLATD------RSALFGGRYTDNPGSSSGDRH 124
Query: 118 -DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVD 176
+QR +L++ T+RL+++ +R+K S+ ETE +G S L DLH QR+ + H L+ +
Sbjct: 125 FEQRQQLLSGTDRLDRSTQRLKASQALAAETEAIGASTLADLHRQREVIEHTTTILYESE 184
Query: 177 DNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
+ +S K + M+RRM N+ I I+ VLV+ IIA++ K
Sbjct: 185 GYVDRSVKTLKGMARRMATNRIITIAIITVLVLLIIAVIVSK 226
>UniRef100_Q6DKB7 VTI1A protein [Xenopus laevis]
Length = 194
Score = 109 bits (272), Expect = 6e-23
Identities = 60/195 (30%), Positives = 107/195 (54%), Gaps = 4/195 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ Y L+A ++ + L G++KKQ + V+ ++EA L+ +MDLE R
Sbjct: 1 MSAEFEGYEQDYAVLTAEITGRVGKIPRLLGDEKKQMILNVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL---ESGMADTMTASA 117
+ + + ++R YK +L L ++ K+ + R+ELL + +
Sbjct: 61 EIPVQSRAMYSNRMRSYKQELGKLDADFKRSRIA-YSDEVRNELLGDDGNSSESQLIKLR 119
Query: 118 DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDD 177
++R L+ +TERL ++ R++ + +ETE++G +IL++L R+ + A L D
Sbjct: 120 EERAHLLDNTERLERSSRRLEAGYQVAVETEQIGQNILENLSQDREKIQRARERLRETDA 179
Query: 178 NIGKSKKIMTNMSRR 192
N+GKS +++T M RR
Sbjct: 180 NLGKSSRVLTGMLRR 194
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 310,164,646
Number of Sequences: 2790947
Number of extensions: 11237582
Number of successful extensions: 47109
Number of sequences better than 10.0: 797
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 691
Number of HSP's that attempted gapping in prelim test: 46590
Number of HSP's gapped (non-prelim): 1095
length of query: 221
length of database: 848,049,833
effective HSP length: 123
effective length of query: 98
effective length of database: 504,763,352
effective search space: 49466808496
effective search space used: 49466808496
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 72 (32.3 bits)
Medicago: description of AC119412.5


