

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.1 - phase: 0
(411 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
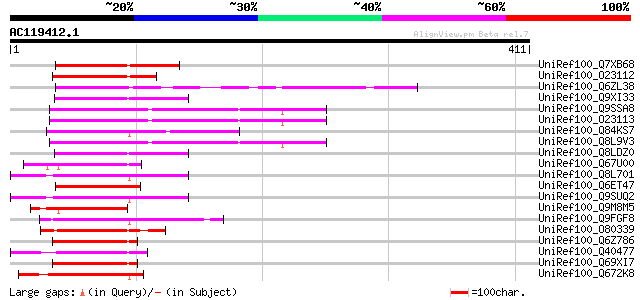
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XB68 Putative AP2 domain containing protein RAP2.11 ... 124 6e-27
UniRef100_O23112 AP2 domain containing protein RAP2.11 [Arabidop... 117 5e-25
UniRef100_Q6ZL38 Putative AP2 domain containing protein [Oryza s... 108 4e-22
UniRef100_Q9XI33 F9L1.31 protein [Arabidopsis thaliana] 90 1e-16
UniRef100_Q9SSA8 T18A20.14 protein [Arabidopsis thaliana] 89 2e-16
UniRef100_O23113 AP2 domain containing protein RAP2.12 [Arabidop... 89 2e-16
UniRef100_Q84KS7 AP2/ERF-domain protein [Solanum tuberosum] 89 3e-16
UniRef100_Q8L9V3 AP2 domain containing protein, putative [Arabid... 88 4e-16
UniRef100_Q8LDZ0 Putative ethylene responsive element [Arabidops... 87 7e-16
UniRef100_Q67U00 Ethylene-binding protein-like [Oryza sativa] 87 9e-16
UniRef100_Q8L701 Putative Ap2 domain protein [Arabidopsis thaliana] 87 1e-15
UniRef100_Q6ET47 Putative ethylene responsive element binding fa... 86 2e-15
UniRef100_Q9SUQ2 Putative Ap2 domain protein [Arabidopsis thaliana] 86 2e-15
UniRef100_Q9M8M5 Hypothetical protein T21F11.9 [Arabidopsis thal... 86 2e-15
UniRef100_Q9FGF8 Similarity to transcription factor [Arabidopsis... 86 2e-15
UniRef100_O80339 Ethylene-responsive transcription factor 3 [Ara... 86 2e-15
UniRef100_Q6Z786 Putative ethylene response factor [Oryza sativa] 85 4e-15
UniRef100_Q40477 Ethylene-responsive transcription factor 4 [Nic... 84 6e-15
UniRef100_Q69XI7 Putative ethylene response factor 1 [Oryza sativa] 84 7e-15
UniRef100_Q672K8 Ethylene response factor 2 [Gossypium barbadense] 84 7e-15
>UniRef100_Q7XB68 Putative AP2 domain containing protein RAP2.11 [Oryza sativa]
Length = 266
Score = 124 bits (310), Expect = 6e-27
Identities = 61/98 (62%), Positives = 78/98 (79%), Gaps = 2/98 (2%)
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPC 96
+FVGVRQRPSGRWVAEIKDT QKIR+WLGTF+TAEEAARAYDEAACLLRG+NTRTNF
Sbjct: 40 KFVGVRQRPSGRWVAEIKDTTQKIRMWLGTFETAEEAARAYDEAACLLRGSNTRTNF--A 97
Query: 97 SQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKY 134
+Q++ L+S+I +L + +++M +FS++ Y
Sbjct: 98 TQAAPDSPLASRIRTILTHKKLKKSMPQPTITFSTAVY 135
>UniRef100_O23112 AP2 domain containing protein RAP2.11 [Arabidopsis thaliana]
Length = 255
Score = 117 bits (294), Expect = 5e-25
Identities = 59/82 (71%), Positives = 68/82 (81%), Gaps = 2/82 (2%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
+ +FVGVRQRPSG+WVAEIKDT QKIR+WLGTF+TAEEAARAYDEAACLLRG+NTRTNF
Sbjct: 21 KTKFVGVRQRPSGKWVAEIKDTTQKIRMWLGTFETAEEAARAYDEAACLLRGSNTRTNF- 79
Query: 95 PCSQSSTSPALSSKITNLLLQR 116
+ + LS KI NLL Q+
Sbjct: 80 -ANHFPNNSQLSLKIRNLLHQK 100
>UniRef100_Q6ZL38 Putative AP2 domain containing protein [Oryza sativa]
Length = 315
Score = 108 bits (269), Expect = 4e-22
Identities = 94/287 (32%), Positives = 126/287 (43%), Gaps = 43/287 (14%)
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPC 96
+FVGVRQRPSGRWVAEIKDT QKIR+WLGTF+TA+ AARAYDEAA LLRGA RTNF P
Sbjct: 50 KFVGVRQRPSGRWVAEIKDTTQKIRMWLGTFETADAAARAYDEAARLLRGAEARTNFAP- 108
Query: 97 SQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRANDHDRQ 156
+ S L+ +I +L + ++ + S + GSP +++
Sbjct: 109 -RISPDCPLAVRIRGILHHKKLKK---------ARSAAAATAGSPGAASKKRSTT----- 153
Query: 157 EMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDTAQFDYISSSFESCLTENN 216
+ + T N + D + S SSS +SC +
Sbjct: 154 -----------AAAAAATPTITTTSNSNSDGAGS------ACGGSSSSSSSTDSC----D 192
Query: 217 GCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNNDVSVDVPDFRFLDDIAPTSYYSPFE 276
G + + +E D EE + F LD A +P
Sbjct: 193 GAVKQGGGGGGAPTDASEVYRPDFVHAGAEEFDSWMFDTAFGPFPELDSFAAVDAVTPPP 252
Query: 277 IAEEMEEQMVENESFCDEPSMIRAVMKRMKYERKFSASLYAFNGIPE 323
E ES P + A +R+K ER+ SASLYA NG+ E
Sbjct: 253 ATASPE------ESSAGTPPVEMAEFERIKVERRISASLYAMNGLQE 293
>UniRef100_Q9XI33 F9L1.31 protein [Arabidopsis thaliana]
Length = 199
Score = 90.1 bits (222), Expect = 1e-16
Identities = 49/106 (46%), Positives = 63/106 (59%), Gaps = 1/106 (0%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
K+F GVRQR G WVAEI+ + K R+WLGTF+TAEEAARAYDEAA L+ G N +TNF P
Sbjct: 5 KKFRGVRQRHWGSWVAEIRHPLLKRRIWLGTFETAEEAARAYDEAAVLMSGRNAKTNF-P 63
Query: 96 CSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSP 141
+ ++T K + ++S SS S+K SP
Sbjct: 64 LNNNNTGETSEGKTDISASSTMSSSTSSSSLSSILSAKLRKCCKSP 109
>UniRef100_Q9SSA8 T18A20.14 protein [Arabidopsis thaliana]
Length = 358
Score = 89.0 bits (219), Expect = 2e-16
Identities = 69/222 (31%), Positives = 109/222 (49%), Gaps = 5/222 (2%)
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + ++ G+RQRP G+W AEI+D + R+WLGTF TAEEAARAYD AA +RG+ +
Sbjct: 119 RKRKNQYRGIRQRPWGKWAAEIRDPREGARIWLGTFKTAEEAARAYDAAARRIRGSKAKV 178
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRAN 151
NF + + S S K L + + + N N S + +S + E + Q N
Sbjct: 179 NFPEENMKANSQKRSVKAN--LQKPVAKPNPNPSPALVQNSNISFENMCFMEEKHQVSNN 236
Query: 152 DHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDTAQFDYISSSFESC 211
++++ M VDA + FS DQ ++ D ++ S+ I I+++ +
Sbjct: 237 NNNQFGMTNSVDAGCNGYQYFSSDQGSNSF-DCSEFGWSDQAPITPDISSAVINNNNSAL 295
Query: 212 LTE--NNGCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNN 251
E N K M+ E+ NN +S D +ED +N
Sbjct: 296 FFEEANPAKKLKSMDFETPYNNTEWDASLDFLNEDAVTTQDN 337
>UniRef100_O23113 AP2 domain containing protein RAP2.12 [Arabidopsis thaliana]
Length = 317
Score = 89.0 bits (219), Expect = 2e-16
Identities = 69/222 (31%), Positives = 109/222 (49%), Gaps = 5/222 (2%)
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + ++ G+RQRP G+W AEI+D + R+WLGTF TAEEAARAYD AA +RG+ +
Sbjct: 78 RKRKNQYRGIRQRPWGKWAAEIRDPREGARIWLGTFKTAEEAARAYDAAARRIRGSKAKV 137
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRAN 151
NF + + S S K L + + + N N S + +S + E + Q N
Sbjct: 138 NFPEENMKANSQKRSVKAN--LQKPVAKPNPNPSPALVQNSNISFENMCFMEEKHQVSNN 195
Query: 152 DHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDTAQFDYISSSFESC 211
++++ M VDA + FS DQ ++ D ++ S+ I I+++ +
Sbjct: 196 NNNQFGMTNSVDAGCNGYQYFSSDQGSNSF-DCSEFGWSDQAPITPDISSAVINNNNSAL 254
Query: 212 LTE--NNGCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNN 251
E N K M+ E+ NN +S D +ED +N
Sbjct: 255 FFEEANPAKKLKSMDFETPYNNTEWDASLDFLNEDAVTTQDN 296
>UniRef100_Q84KS7 AP2/ERF-domain protein [Solanum tuberosum]
Length = 264
Score = 88.6 bits (218), Expect = 3e-16
Identities = 56/158 (35%), Positives = 87/158 (54%), Gaps = 7/158 (4%)
Query: 30 GARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANT 89
G R+ + ++ G+RQRP G+W AEI+D + +RVWLGTF+TAE+AARAYDEAA +RG
Sbjct: 89 GPRQRKNKYRGIRQRPWGKWAAEIRDPQKGVRVWLGTFNTAEDAARAYDEAAKRIRGNKA 148
Query: 90 RTNF----WPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERR 145
+ NF P + TS + LLL+ + NN S + Y +P
Sbjct: 149 KLNFPAPSPPAKRQCTSTVAADPPPALLLESSNIISYNN--SPLMNFGYDVQSQTPYYPM 206
Query: 146 EQQRA-NDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLN 182
E A +D++ +E ++++ E+ S DQF+ ++
Sbjct: 207 EMPVASDDYELKEQISNLESFLELEPADSSDQFSGIVD 244
>UniRef100_Q8L9V3 AP2 domain containing protein, putative [Arabidopsis thaliana]
Length = 358
Score = 88.2 bits (217), Expect = 4e-16
Identities = 69/222 (31%), Positives = 109/222 (49%), Gaps = 5/222 (2%)
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + ++ G+RQRP G+W AEI+D + R+WLGTF TAEEAARAYD AA +RG+ +
Sbjct: 119 RKRKNQYRGIRQRPWGKWAAEIRDPREGARIWLGTFKTAEEAARAYDAAARRIRGSKAKV 178
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQRAN 151
NF + + S S K L + + + N N S + +S + E + Q N
Sbjct: 179 NFPEENLKANSQKRSVKAN--LQKPVAKPNPNPSPALVQNSNISFENMCFMEEKHQVSNN 236
Query: 152 DHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDTAQFDYISSSFESC 211
++++ M VDA + FS DQ ++ D ++ S+ I I+++ +
Sbjct: 237 NNNQFGMTNSVDAGCNGYQYFSSDQGSNSF-DCSEFGWSDQAPITPDISSAVINNNNSAL 295
Query: 212 LTE--NNGCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNN 251
E N K M+ E+ NN +S D +ED +N
Sbjct: 296 FFEEANPAKKLKSMDFETPYNNTEWDASLDFLNEDAVTTQDN 337
>UniRef100_Q8LDZ0 Putative ethylene responsive element [Arabidopsis thaliana]
Length = 199
Score = 87.4 bits (215), Expect = 7e-16
Identities = 48/106 (45%), Positives = 62/106 (58%), Gaps = 1/106 (0%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWP 95
K+F GVRQR G WVAEI+ + K R+WLGTF+TAEEAARAY EAA L+ G N +TNF P
Sbjct: 5 KKFRGVRQRHWGSWVAEIRHPLLKRRIWLGTFETAEEAARAYHEAAVLMSGRNAKTNF-P 63
Query: 96 CSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSP 141
+ ++T K + ++S SS S+K SP
Sbjct: 64 LNNNNTGETSEGKTDISASSTMSSSTSSSSLSSILSAKLRKCCKSP 109
>UniRef100_Q67U00 Ethylene-binding protein-like [Oryza sativa]
Length = 274
Score = 87.0 bits (214), Expect = 9e-16
Identities = 50/102 (49%), Positives = 61/102 (59%), Gaps = 10/102 (9%)
Query: 12 EEGSMVWDDMMKEAASL----GGARRVRKR-----FVGVRQRPSGRWVAEIKDTIQKIRV 62
EE M D AA+ G +RVR+R + GVR RP G+W AEI+D + +R
Sbjct: 95 EEAVMATDYAAAAAAAAVAGGSGGKRVRRRRKKNVYRGVRHRPWGKWAAEIRDPRRAVRK 154
Query: 63 WLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQSSTSPA 104
WLGTFDTAEEAARAYD AA RGA + NF PCS+ P+
Sbjct: 155 WLGTFDTAEEAARAYDRAALEFRGARAKLNF-PCSEPLPMPS 195
>UniRef100_Q8L701 Putative Ap2 domain protein [Arabidopsis thaliana]
Length = 343
Score = 86.7 bits (213), Expect = 1e-15
Identities = 53/143 (37%), Positives = 74/143 (51%), Gaps = 7/143 (4%)
Query: 1 MARKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKI 60
M K+K AV+ S V + + G K+F GVRQRP G+W AEI+D ++++
Sbjct: 90 MESKKKQKRAVKSESTVSPVVSATTTTTG-----EKKFRGVRQRPWGKWAAEIRDPLKRV 144
Query: 61 RVWLGTFDTAEEAARAYDEAACLLRGANTRTNF--WPCSQSSTSPALSSKITNLLLQRLK 118
R+WLGT++TAEEAA YD AA LRG + TNF P + + S + + K
Sbjct: 145 RLWLGTYNTAEEAAMVYDNAAIQLRGPDALTNFSVTPTTATEKKAPPPSPVKKKKKKNNK 204
Query: 119 ERNMNNSCSSFSSSKYTSLLGSP 141
+ + SS S S L SP
Sbjct: 205 SKKSVTASSSISRSSSNDCLCSP 227
>UniRef100_Q6ET47 Putative ethylene responsive element binding factor [Oryza sativa]
Length = 203
Score = 85.5 bits (210), Expect = 2e-15
Identities = 42/67 (62%), Positives = 50/67 (73%)
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPC 96
R+ GVR+RPSGR+ AEI+D +K +WLGTFD+AE AARAYD AA LRGA RTNF P
Sbjct: 12 RYRGVRRRPSGRYAAEIRDPAKKTPIWLGTFDSAEAAARAYDAAARSLRGAAARTNFPPS 71
Query: 97 SQSSTSP 103
+ SS P
Sbjct: 72 TSSSAPP 78
>UniRef100_Q9SUQ2 Putative Ap2 domain protein [Arabidopsis thaliana]
Length = 343
Score = 85.5 bits (210), Expect = 2e-15
Identities = 52/143 (36%), Positives = 74/143 (51%), Gaps = 7/143 (4%)
Query: 1 MARKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKI 60
M K++ AV+ S V + + G K+F GVRQRP G+W AEI+D ++++
Sbjct: 90 MESKKRQKRAVKSESTVSPVVSATTTTTG-----EKKFRGVRQRPWGKWAAEIRDPLKRV 144
Query: 61 RVWLGTFDTAEEAARAYDEAACLLRGANTRTNF--WPCSQSSTSPALSSKITNLLLQRLK 118
R+WLGT++TAEEAA YD AA LRG + TNF P + + S + + K
Sbjct: 145 RLWLGTYNTAEEAAMVYDNAAIQLRGPDALTNFSVTPTTATEKKAPPPSPVKKKKKKNNK 204
Query: 119 ERNMNNSCSSFSSSKYTSLLGSP 141
+ + SS S S L SP
Sbjct: 205 SKKSVTASSSISRSSSNDCLCSP 227
>UniRef100_Q9M8M5 Hypothetical protein T21F11.9 [Arabidopsis thaliana]
Length = 256
Score = 85.5 bits (210), Expect = 2e-15
Identities = 48/83 (57%), Positives = 55/83 (65%), Gaps = 10/83 (12%)
Query: 17 VWDDMMKEAASLGGARRVRKR------FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTA 70
V D + KE GG +R RK F+GVR+RP GRW AEI+D I + R WLGTFDTA
Sbjct: 93 VSDHLTKE----GGVKRGRKMPQKTGGFMGVRKRPWGRWSAEIRDRIGRCRHWLGTFDTA 148
Query: 71 EEAARAYDEAACLLRGANTRTNF 93
EEAARAYD AA LRG +TNF
Sbjct: 149 EEAARAYDAAARRLRGTKAKTNF 171
>UniRef100_Q9FGF8 Similarity to transcription factor [Arabidopsis thaliana]
Length = 391
Score = 85.5 bits (210), Expect = 2e-15
Identities = 57/160 (35%), Positives = 76/160 (46%), Gaps = 19/160 (11%)
Query: 24 EAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACL 83
E +S G + R+R+ GVRQRP G+W AEI+D + RVWLGTFD AE AARAYDEAA
Sbjct: 172 ETSSFSGDQP-RRRYRGVRQRPWGKWAAEIRDPFKAARVWLGTFDNAESAARAYDEAALR 230
Query: 84 LRGANTRTNF--------------WPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSF 129
RG + NF P Q++ S+ + L R +N
Sbjct: 231 FRGNKAKLNFPENVKLVRPASTEAQPVHQTAAQRPTQSRNSGSTTTLLPIRPASNQSVHS 290
Query: 130 SSSKYTSLLGSPEERREQQRANDHDRQEMQQQVDAYGEVS 169
+ L E R+QQ+ H QQ +D Y ++S
Sbjct: 291 QPLMQSYNLSYSEMARQQQQFQQHH----QQSLDLYDQMS 326
>UniRef100_O80339 Ethylene-responsive transcription factor 3 [Arabidopsis thaliana]
Length = 225
Score = 85.5 bits (210), Expect = 2e-15
Identities = 50/99 (50%), Positives = 62/99 (62%), Gaps = 9/99 (9%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
AA G + +R F GVR+RP GR+ AEI+D +K RVWLGTFD+AEEAARAYD AA L
Sbjct: 17 AAINGSVKEIR--FRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEEAARAYDSAARNL 74
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMN 123
RG +TNF P SS P NL +++ +N N
Sbjct: 75 RGPKAKTNF-PIDSSSPPP------PNLRFNQIRNQNQN 106
>UniRef100_Q6Z786 Putative ethylene response factor [Oryza sativa]
Length = 205
Score = 84.7 bits (208), Expect = 4e-15
Identities = 42/67 (62%), Positives = 52/67 (76%), Gaps = 1/67 (1%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
+K+F GVRQR G WV+EI+ + K RVWLGTF+TAEEAARAYDEAA L+ G N +TNF
Sbjct: 5 KKKFRGVRQRHWGSWVSEIRHPLLKRRVWLGTFETAEEAARAYDEAAVLMSGRNAKTNF- 63
Query: 95 PCSQSST 101
P ++ST
Sbjct: 64 PVQRNST 70
>UniRef100_Q40477 Ethylene-responsive transcription factor 4 [Nicotiana tabacum]
Length = 225
Score = 84.3 bits (207), Expect = 6e-15
Identities = 52/109 (47%), Positives = 64/109 (58%), Gaps = 12/109 (11%)
Query: 1 MARKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKI 60
MA K KVS +G V D +KE + GVR+RP GR+ AEI+D +K
Sbjct: 1 MAVKNKVSNGNLKGGNVKTDGVKEV-----------HYRGVRKRPWGRYAAEIRDPGKKS 49
Query: 61 RVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQSSTSPALSSKI 109
RVWLGTFDTAEEAA+AYD AA RG +TNF P + SP+ SS +
Sbjct: 50 RVWLGTFDTAEEAAKAYDTAAREFRGPKAKTNF-PSPTENQSPSHSSTV 97
>UniRef100_Q69XI7 Putative ethylene response factor 1 [Oryza sativa]
Length = 243
Score = 84.0 bits (206), Expect = 7e-15
Identities = 41/67 (61%), Positives = 53/67 (78%), Gaps = 1/67 (1%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
+K+F GVRQR G WV+EI+ + K RVWLGTF+TAEEAARAYDEAA L+ G N +TNF
Sbjct: 5 KKKFRGVRQRHWGSWVSEIRHPLLKRRVWLGTFETAEEAARAYDEAAILMSGRNAKTNF- 63
Query: 95 PCSQSST 101
P ++++T
Sbjct: 64 PVARNAT 70
>UniRef100_Q672K8 Ethylene response factor 2 [Gossypium barbadense]
Length = 198
Score = 84.0 bits (206), Expect = 7e-15
Identities = 47/102 (46%), Positives = 63/102 (61%), Gaps = 10/102 (9%)
Query: 8 SEAVEEGSMVWDDMMKEAASLGGARRVRKRFV-GVRQRPSGRWVAEIKDTIQKIRVWLGT 66
SE +++ S V + K +R RK F G+RQRP G+W AEI+D + +RVWLGT
Sbjct: 70 SETIQKTSQVTKKVEK-------TQRTRKNFYRGIRQRPWGKWAAEIRDPQKGVRVWLGT 122
Query: 67 FDTAEEAARAYDEAACLLRGANTRTNF--WPCSQSSTSPALS 106
++TAEEA RAYDEAA +RG + NF P + SPAL+
Sbjct: 123 YNTAEEATRAYDEAAKRIRGDKAKLNFPQTPTKNQAASPALT 164
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 684,605,480
Number of Sequences: 2790947
Number of extensions: 29647346
Number of successful extensions: 171226
Number of sequences better than 10.0: 2063
Number of HSP's better than 10.0 without gapping: 636
Number of HSP's successfully gapped in prelim test: 1491
Number of HSP's that attempted gapping in prelim test: 153809
Number of HSP's gapped (non-prelim): 11168
length of query: 411
length of database: 848,049,833
effective HSP length: 130
effective length of query: 281
effective length of database: 485,226,723
effective search space: 136348709163
effective search space used: 136348709163
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC119412.1


