

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
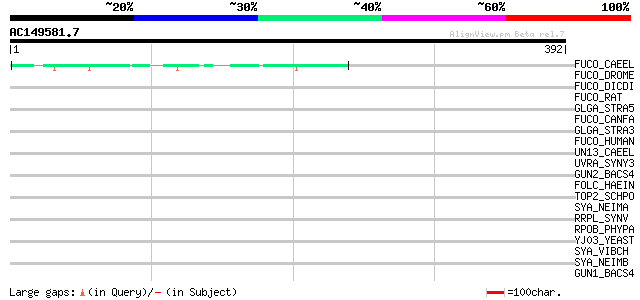
Score E
Sequences producing significant alignments: (bits) Value
FUCO_CAEEL (P49713) Putative alpha-L-fucosidase precursor (EC 3.... 45 4e-04
FUCO_DROME (Q9VTJ4) Putative alpha-L-fucosidase precursor (EC 3.... 43 0.001
FUCO_DICDI (P10901) Alpha-L-fucosidase precursor (EC 3.2.1.51) (... 43 0.002
FUCO_RAT (P17164) Tissue alpha-L-fucosidase precursor (EC 3.2.1.... 38 0.051
GLGA_STRA5 (Q8E079) Glycogen synthase (EC 2.4.1.21) (Starch [bac... 36 0.15
FUCO_CANFA (P48300) Tissue alpha-L-fucosidase precursor (EC 3.2.... 36 0.15
GLGA_STRA3 (Q8E5V5) Glycogen synthase (EC 2.4.1.21) (Starch [bac... 35 0.33
FUCO_HUMAN (P04066) Tissue alpha-L-fucosidase precursor (EC 3.2.... 34 0.56
UN13_CAEEL (P27715) Phorbol ester/diacylglycerol-binding protein... 33 0.96
UVRA_SYNY3 (P73412) UvrABC system protein A (UvrA protein) (Exci... 32 2.8
GUN2_BACS4 (P06565) Endoglucanase B (EC 3.2.1.4) (Endo-1,4-beta-... 31 4.8
FOLC_HAEIN (P43775) Folylpolyglutamate synthase (EC 6.3.2.17) (F... 31 4.8
TOP2_SCHPO (P08096) DNA topoisomerase II (EC 5.99.1.3) 31 6.2
SYA_NEIMA (Q9JTG4) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 31 6.2
RRPL_SYNV (P31332) RNA polymerase beta subunit (EC 2.7.7.48) (La... 31 6.2
RPOB_PHYPA (P60283) DNA-directed RNA polymerase beta chain (EC 2... 31 6.2
YJ03_YEAST (P47104) Hypothetical 154.9 kDa protein in CPR7-PET19... 30 8.1
SYA_VIBCH (Q56648) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 30 8.1
SYA_NEIMB (Q9JYG6) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 30 8.1
GUN1_BACS4 (P06566) Endoglucanase A (EC 3.2.1.4) (Endo-1,4-beta-... 30 8.1
>FUCO_CAEEL (P49713) Putative alpha-L-fucosidase precursor (EC
3.2.1.51) (Alpha-L-fucoside fucohydrolase)
Length = 407
Score = 44.7 bits (104), Expect = 4e-04
Identities = 58/255 (22%), Positives = 97/255 (37%), Gaps = 49/255 (19%)
Query: 2 ILTAKHHDGFCLWPSKYTKHSVISSKWQN----GKGDVVKEFVNAATDKGIDVGIYLS-- 55
+ T+KHH+GF +WPS+ S W + K D+V E +A + G+Y S
Sbjct: 130 VFTSKHHEGFTMWPSR------TSWNWNSMDIGPKRDIVGELRDAFKKTDVHFGLYFSQF 183
Query: 56 ----PWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDW 111
P D ++ + Q+ +++ KY + +W DG D SD
Sbjct: 184 EWFHPMFLDDGKFNTTFYPEQVSYPQMIDIVTKY-NPEVVWSDGEWDK---------SDD 233
Query: 112 FSMVKE----LQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLN 167
+ KE L +S + + RW TGT G G + + +
Sbjct: 234 YWKAKEFLAWLYNSSPVKDQVVVNDRW---GTGTMGK-----------HGGFMTYSDHYD 279
Query: 168 TGDPKGTDWLPAECDVSIRPGWFWHKSESPKKLS---DLLDIYYKSVGRNCVLLLNVPPN 224
G W C + W + +++ ++++ +++ N LLLNV PN
Sbjct: 280 PGKLLEKKW--ENCMTLDKHSWGNRRDMKASEVNTAYEIIEQLARTIACNGNLLLNVGPN 337
Query: 225 TTGLISENDAHRLKE 239
G I RL+E
Sbjct: 338 MHGQIPAIFEDRLEE 352
>FUCO_DROME (Q9VTJ4) Putative alpha-L-fucosidase precursor (EC
3.2.1.51) (Alpha-L-fucoside fucohydrolase)
Length = 494
Score = 43.1 bits (100), Expect = 0.001
Identities = 34/108 (31%), Positives = 54/108 (49%), Gaps = 20/108 (18%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQN----GKGDVVKEFVNA-ATDKGIDVGIYLS 55
++LT+KHHDGF LWPSK S W + K D+VKE A + + G+Y S
Sbjct: 129 VVLTSKHHDGFTLWPSKN------SYGWNSMDVGPKRDIVKELAAAIRKESDLRFGLYYS 182
Query: 56 PWDRHDSRYGHD---LLYNEYYL-----AQLQELLKKYQDVREIWFDG 95
++ + + D LL ++++ + EL+++Y IW DG
Sbjct: 183 LFEWFNPLWTDDKLHLLMQQHFVERKVRPEQMELVQQYLP-EIIWSDG 229
>FUCO_DICDI (P10901) Alpha-L-fucosidase precursor (EC 3.2.1.51)
(Alpha-L-fucoside fucohydrolase)
Length = 461
Score = 42.7 bits (99), Expect = 0.002
Identities = 30/109 (27%), Positives = 54/109 (49%), Gaps = 21/109 (19%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKW---QNGKG-DVVKEFVNAATDKGIDVGIYLSP 56
++LT+KHH+G+ LW S+ S W + G G D+V E + + G+ +G+Y S
Sbjct: 122 VVLTSKHHEGYTLWNSEQ------SWNWNSVETGPGIDIVGELTKSVKNMGLHMGLYHSL 175
Query: 57 WDRHDSRYGHD----------LLYNEYYLAQLQELLKKYQDVREIWFDG 95
++ + Y D + +E + QL++++ Y+ IW DG
Sbjct: 176 FEWFNPLYLADAETGKNPTTQVYVDEILMKQLKDIVTTYEP-ELIWADG 223
>FUCO_RAT (P17164) Tissue alpha-L-fucosidase precursor (EC 3.2.1.51)
(Alpha-L-fucosidase I) (Alpha-L-fucoside fucohydrolase)
Length = 462
Score = 37.7 bits (86), Expect = 0.051
Identities = 31/103 (30%), Positives = 52/103 (50%), Gaps = 11/103 (10%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
++LTAKHH+GF WPS + + +SK D+V E A + I G+Y S ++
Sbjct: 128 VVLTAKHHEGFTNWPSAVSWN--WNSKDVGPHRDLVGELGAAVRKRNIRYGLYHSLFEWF 185
Query: 61 DSRYGHDL---LYNEYYLA-----QLQELLKKYQDVREIWFDG 95
Y D L +++++ +L +L+ +Y+ IW DG
Sbjct: 186 HPLYLLDKKNGLKTQHFVSTKTMPELYDLVNRYKP-DLIWSDG 227
>GLGA_STRA5 (Q8E079) Glycogen synthase (EC 2.4.1.21) (Starch
[bacterial glycogen] synthase)
Length = 476
Score = 36.2 bits (82), Expect = 0.15
Identities = 25/83 (30%), Positives = 38/83 (45%), Gaps = 15/83 (18%)
Query: 31 GKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVRE 90
G GDV+ + + KG DV + + +D D ++G Q++ L+ Y DV
Sbjct: 18 GLGDVIGALPKSLSKKGHDVAVVMPYYDMVDQKFGD----------QIENLMYFYTDVG- 66
Query: 91 IW---FDGAKDPRAQNVTYYFSD 110
W + G K NVT+YF D
Sbjct: 67 -WRHQYVGVKRLSQDNVTFYFID 88
>FUCO_CANFA (P48300) Tissue alpha-L-fucosidase precursor (EC
3.2.1.51) (Alpha-L-fucosidase I) (Alpha-L-fucoside
fucohydrolase)
Length = 465
Score = 36.2 bits (82), Expect = 0.15
Identities = 30/107 (28%), Positives = 48/107 (44%), Gaps = 19/107 (17%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNG----KGDVVKEFVNAATDKGIDVGIYLSP 56
++LT KHH+GF WPS +S W + D+V E A + I G+Y S
Sbjct: 132 VVLTTKHHEGFTNWPSS------VSWNWNSNDVGPHRDLVGELGRALRKRNIRYGLYHSL 185
Query: 57 WDRHDSRYGHDLLYN---EYY-----LAQLQELLKKYQDVREIWFDG 95
+ Y D N +++ + +L +L+ +Y+ IW DG
Sbjct: 186 LEWFHPLYLLDKKNNFKTQFFVRAKTMPELYDLVNRYEP-DLIWSDG 231
>GLGA_STRA3 (Q8E5V5) Glycogen synthase (EC 2.4.1.21) (Starch
[bacterial glycogen] synthase)
Length = 476
Score = 35.0 bits (79), Expect = 0.33
Identities = 25/83 (30%), Positives = 38/83 (45%), Gaps = 15/83 (18%)
Query: 31 GKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVRE 90
G GDV+ + + KG DV + + +D D ++G Q++ L+ Y DV
Sbjct: 18 GLGDVIGALPKSLSKKGHDVVVVMPYYDMVDQKFGD----------QIENLMYFYTDVG- 66
Query: 91 IW---FDGAKDPRAQNVTYYFSD 110
W + G K NVT+YF D
Sbjct: 67 -WRHQYVGVKRLSQDNVTFYFID 88
>FUCO_HUMAN (P04066) Tissue alpha-L-fucosidase precursor (EC
3.2.1.51) (Alpha-L-fucosidase I) (Alpha-L-fucoside
fucohydrolase)
Length = 461
Score = 34.3 bits (77), Expect = 0.56
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLS 55
++LT KHH+GF WPS + + +SK D+V E A + I G+Y S
Sbjct: 127 VVLTTKHHEGFTNWPSPVSWN--WNSKDVGPHRDLVGELGTALRKRNIRYGLYHS 179
>UN13_CAEEL (P27715) Phorbol ester/diacylglycerol-binding protein
unc-13 (Uncoordinated protein 13)
Length = 2155
Score = 33.5 bits (75), Expect = 0.96
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 3/71 (4%)
Query: 231 ENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGG---KEGGFGPENMLDSDHLWSYWT 287
++D H+ E F + + +R + S GG + G DS H W Y +
Sbjct: 457 QDDTHQYDEVDRGSRVSFTRTPSVDRTDRPSESGGGFYDEMSESGRPGRPDSHHNWRYDS 516
Query: 288 PREDDKEKDHW 298
+E+D EKD+W
Sbjct: 517 IQEEDNEKDNW 527
>UVRA_SYNY3 (P73412) UvrABC system protein A (UvrA protein)
(Excinuclease ABC subunit A)
Length = 970
Score = 32.0 bits (71), Expect = 2.8
Identities = 24/84 (28%), Positives = 39/84 (45%), Gaps = 14/84 (16%)
Query: 50 VGIYLSPWDRHDSRYGHDLLYN--EYYLAQLQELLKK----------YQDVREIWFDGAK 97
V + ++PW D+ Y LLY+ +++ QLQ KK Y EIWF+G
Sbjct: 333 VYVAIAPWSEKDNSYYLSLLYSLGQHFDFQLQTPWKKLTKEQKEIILYGTEEEIWFEG-- 390
Query: 98 DPRAQNVTYYFSDWFSMVKELQSS 121
+ R +N Y+ + + LQ +
Sbjct: 391 ESRYRNKQGYYRRFAGALNILQKN 414
>GUN2_BACS4 (P06565) Endoglucanase B (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Cellulase)
Length = 409
Score = 31.2 bits (69), Expect = 4.8
Identities = 21/71 (29%), Positives = 33/71 (45%), Gaps = 1/71 (1%)
Query: 36 VKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDG 95
VKE V A + ID+GIY+ D H +Y E E+ + Y D + ++
Sbjct: 104 VKEKVKEAVEAAIDLGIYVI-IDWHILSDNDPNIYKEEAKDFFDEMSELYGDYPNVIYEI 162
Query: 96 AKDPRAQNVTY 106
A +P +VT+
Sbjct: 163 ANEPNGSDVTW 173
>FOLC_HAEIN (P43775) Folylpolyglutamate synthase (EC 6.3.2.17)
(Folylpoly-gamma-glutamate synthetase) (FPGS)
Length = 437
Score = 31.2 bits (69), Expect = 4.8
Identities = 30/110 (27%), Positives = 49/110 (44%), Gaps = 19/110 (17%)
Query: 23 VISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELL 82
VI+ NGKG + + G+ VG+Y SP H L YNE Q Q+L
Sbjct: 50 VITVGGTNGKGTTCRLLETILLNHGLRVGVYSSP---------HLLRYNERVRIQNQDLP 100
Query: 83 KKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQSSINIFSDAGPDV 132
+ + D + + +++TY+ FS + S++++F A DV
Sbjct: 101 DEAHTASFAFID---ENKTESLTYF---EFSTL----SALHLFKQAKLDV 140
>TOP2_SCHPO (P08096) DNA topoisomerase II (EC 5.99.1.3)
Length = 1485
Score = 30.8 bits (68), Expect = 6.2
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 2/63 (3%)
Query: 30 NGKGDVVKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVR 89
NG+G +K + T D+ Y S DRH +Y H + + L ++ KK DVR
Sbjct: 646 NGRGWKIKYYKGLGTSDHDDMKSYFSDLDRH-MKYFHAMQEKDAELIEM-AFAKKKADVR 703
Query: 90 EIW 92
+ W
Sbjct: 704 KEW 706
>SYA_NEIMA (Q9JTG4) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 874
Score = 30.8 bits (68), Expect = 6.2
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Query: 256 RYVKVSSQRGGKEGGFGPENMLDSDHLWSYW--TPREDDKEKDHWIEIWGNDGSLRFN 311
+YV + + G G GP + + DH W P +++ D WIEIW N ++FN
Sbjct: 162 KYVSDNFWQMGDTGPCGPCSEIFYDHGEEIWGGIPGSPEEDGDRWIEIW-NCVFMQFN 218
>RRPL_SYNV (P31332) RNA polymerase beta subunit (EC 2.7.7.48) (Large
structural protein) (L protein)
Length = 2116
Score = 30.8 bits (68), Expect = 6.2
Identities = 26/83 (31%), Positives = 39/83 (46%), Gaps = 18/83 (21%)
Query: 19 TKHSVISS-------KWQNGKGDVVKEFVNAATDKGID-----VGIYLSPWDRH---DSR 63
TK+SV S KW N + D + F+ DKG+D +G+Y P +R +R
Sbjct: 519 TKNSVFDSTKRRGILKWLNEQSDKIYNFLMRIDDKGLDEDDCIIGLY--PKEREMKTKAR 576
Query: 64 YGHDLLYN-EYYLAQLQELLKKY 85
+ + Y Y+ +ELL KY
Sbjct: 577 FFSLMSYKLRMYVTSTEELLGKY 599
>RPOB_PHYPA (P60283) DNA-directed RNA polymerase beta chain (EC
2.7.7.6) (PEP) (Plastid-encoded RNA polymerase beta
subunit) (RNA polymerase beta subunit)
Length = 1085
Score = 30.8 bits (68), Expect = 6.2
Identities = 28/108 (25%), Positives = 43/108 (38%), Gaps = 10/108 (9%)
Query: 165 YLNTGDPKGTDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPN 224
++ TGD P E + S+R + K L + I + C L VP N
Sbjct: 746 WVETGDVLVGKLTPQETEESLR-------APEGKLLQAIFGIQVTNAKETC---LKVPLN 795
Query: 225 TTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFG 272
G + + KE S + I H IA+ R ++V + G+ G G
Sbjct: 796 GKGRVIDVIWISKKENSSNYEKIIHVYIAQKRKIQVGDKVAGRHGNKG 843
>YJ03_YEAST (P47104) Hypothetical 154.9 kDa protein in CPR7-PET191
intergenic region
Length = 1357
Score = 30.4 bits (67), Expect = 8.1
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 2/57 (3%)
Query: 52 IYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVREIWF--DGAKDPRAQNVTY 106
I LS DRH + + Y Y + Q+L+KK++D+ + F D +DP + Y
Sbjct: 917 ILLSSIDRHITSWNRAREYRIAYWIKEQDLVKKFEDIAKYEFSKDDKRDPSRCAIFY 973
>SYA_VIBCH (Q56648) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 860
Score = 30.4 bits (67), Expect = 8.1
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 16/68 (23%)
Query: 252 IAENRYVKVSSQRGGKE---------GGFGP-----ENMLD-SDHLWSYWTPREDDKEKD 296
I +R +++ ++GGK+ G GP E D DH+W P +++ D
Sbjct: 147 IPADRIIRIGDKKGGKKFDSDNFWQMGDTGPCGPCTEIFYDHGDHIWG-GPPGSPEEDGD 205
Query: 297 HWIEIWGN 304
+IEIW N
Sbjct: 206 RFIEIWNN 213
>SYA_NEIMB (Q9JYG6) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 874
Score = 30.4 bits (67), Expect = 8.1
Identities = 21/65 (32%), Positives = 32/65 (48%), Gaps = 6/65 (9%)
Query: 252 IAENRYVKVSSQ---RGGKEGGFGPENMLDSDHLWSYW--TPREDDKEKDHWIEIWGNDG 306
I +N+ K +S + G G GP + + DH W P +++ D WIEIW N
Sbjct: 155 IGDNKGAKYASDNFWQMGDTGPCGPCSEIFYDHGEEIWGGIPGSPEEDGDRWIEIW-NCV 213
Query: 307 SLRFN 311
++FN
Sbjct: 214 FMQFN 218
>GUN1_BACS4 (P06566) Endoglucanase A (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Cellulase)
Length = 488
Score = 30.4 bits (67), Expect = 8.1
Identities = 21/71 (29%), Positives = 31/71 (43%), Gaps = 1/71 (1%)
Query: 36 VKEFVNAATDKGIDVGIYLSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDG 95
VKE V A + ID+GIY+ D H +Y E E+ Y D + ++
Sbjct: 102 VKEKVKEAVEAAIDLGIYVI-IDWHILSDNDPNIYKEEAKEFFDEMSALYGDYPNVIYEI 160
Query: 96 AKDPRAQNVTY 106
A +P NV +
Sbjct: 161 ANEPNGHNVRW 171
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,831,467
Number of Sequences: 164201
Number of extensions: 2274650
Number of successful extensions: 4963
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 4946
Number of HSP's gapped (non-prelim): 32
length of query: 392
length of database: 59,974,054
effective HSP length: 112
effective length of query: 280
effective length of database: 41,583,542
effective search space: 11643391760
effective search space used: 11643391760
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149581.7


