

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.3 + phase: 0
(194 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
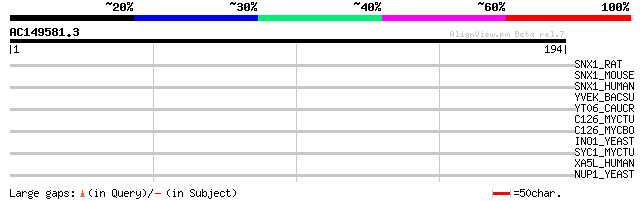
Score E
Sequences producing significant alignments: (bits) Value
SNX1_RAT (Q99N27) Sorting nexin 1 32 0.71
SNX1_MOUSE (Q9WV80) Sorting nexin 1 32 0.71
SNX1_HUMAN (Q13596) Sorting nexin 1 32 0.71
YVEK_BACSU (P71050) Hypothetical protein yveK 31 2.1
YT06_CAUCR (Q9A4D0) Hypothetical UPF0276 protein CC2906 30 2.7
C126_MYCTU (P63711) Putative cytochrome P450 126 (EC 1.14.-.-) 30 2.7
C126_MYCBO (P63712) Putative cytochrome P450 126 (EC 1.14.-.-) 30 2.7
INO1_YEAST (P11986) Inositol-3-phosphate synthase (EC 5.5.1.4) (... 30 4.6
SYC1_MYCTU (P96862) Cysteinyl-tRNA synthetase 1 (EC 6.1.1.16) (C... 29 6.0
XA5L_HUMAN (Q9Y247) XAP-5-like protein 29 7.8
NUP1_YEAST (P20676) Nucleoporin NUP1 (Nuclear pore protein NUP1) 29 7.8
>SNX1_RAT (Q99N27) Sorting nexin 1
Length = 522
Score = 32.3 bits (72), Expect = 0.71
Identities = 27/95 (28%), Positives = 44/95 (45%), Gaps = 3/95 (3%)
Query: 98 LLGMSEVDVRVEVRSESAGHISY--PTLKRVYEHHLTEARRLEEPQTREKLQERGRKRAI 155
L+GM++V V E S SA + L+R + + L++P RE L++ RA+
Sbjct: 216 LIGMTKVKVGKE-DSSSAEFLEKRRAALERYLQRIVNHPTMLQDPDVREFLEKEELPRAV 274
Query: 156 GKWSWGGMALAYLYDYLDDSVILNNRTMAGSTILF 190
G + G L +++ D+V M S I F
Sbjct: 275 GTQALSGAGLLKMFNKATDAVSKMTIKMNESDIWF 309
>SNX1_MOUSE (Q9WV80) Sorting nexin 1
Length = 522
Score = 32.3 bits (72), Expect = 0.71
Identities = 27/95 (28%), Positives = 44/95 (45%), Gaps = 3/95 (3%)
Query: 98 LLGMSEVDVRVEVRSESAGHISY--PTLKRVYEHHLTEARRLEEPQTREKLQERGRKRAI 155
L+GM++V V E S SA + L+R + + L++P RE L++ RA+
Sbjct: 216 LIGMTKVKVGKE-DSSSAEFLEKRRAALERYLQRIVNHPTMLQDPDVREFLEKEELPRAV 274
Query: 156 GKWSWGGMALAYLYDYLDDSVILNNRTMAGSTILF 190
G + G L +++ D+V M S I F
Sbjct: 275 GTQALSGAGLLKMFNKATDAVSKMTIKMNESDIWF 309
>SNX1_HUMAN (Q13596) Sorting nexin 1
Length = 522
Score = 32.3 bits (72), Expect = 0.71
Identities = 27/95 (28%), Positives = 44/95 (45%), Gaps = 3/95 (3%)
Query: 98 LLGMSEVDVRVEVRSESAGHISY--PTLKRVYEHHLTEARRLEEPQTREKLQERGRKRAI 155
L+GM++V V E S SA + L+R + + L++P RE L++ RA+
Sbjct: 216 LIGMTKVKVGKE-DSSSAEFLEKRRAALERYLQRIVNHPTMLQDPDVREFLEKEELPRAV 274
Query: 156 GKWSWGGMALAYLYDYLDDSVILNNRTMAGSTILF 190
G + G L +++ D+V M S I F
Sbjct: 275 GTQTLSGAGLLKMFNKATDAVSKMTIKMNESDIWF 309
>YVEK_BACSU (P71050) Hypothetical protein yveK
Length = 234
Score = 30.8 bits (68), Expect = 2.1
Identities = 28/101 (27%), Positives = 43/101 (41%), Gaps = 18/101 (17%)
Query: 75 IHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEA 134
I+ + DHD A + A L+ + EVD R+ V+ H L+EA
Sbjct: 117 INVAVQDHDPAKAAEIANTLVNKF--EKEVDERMNVQGV---------------HILSEA 159
Query: 135 RRLEEPQTREKLQERGRKRAIGKWSWGGMALAYLYDYLDDS 175
+ E P + + R A G GG+ LA+ +LDD+
Sbjct: 160 KASESPMIKPA-RLRNMVMAFGAAVMGGITLAFFLHFLDDT 199
>YT06_CAUCR (Q9A4D0) Hypothetical UPF0276 protein CC2906
Length = 280
Score = 30.4 bits (67), Expect = 2.7
Identities = 12/27 (44%), Positives = 17/27 (62%)
Query: 15 WEPVEGFDLHDLIYTGYSTVTHAMICA 41
W VEGF+ HDL+ Y+ A++CA
Sbjct: 100 WTGVEGFNSHDLLPVPYTEEAMAVVCA 126
>C126_MYCTU (P63711) Putative cytochrome P450 126 (EC 1.14.-.-)
Length = 414
Score = 30.4 bits (67), Expect = 2.7
Identities = 28/97 (28%), Positives = 37/97 (37%), Gaps = 9/97 (9%)
Query: 4 LGKPEAGLRWFWEPVE-GFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTI 62
LG PE W +E VE GFD + G T + T + L+ G+
Sbjct: 162 LGVPETDRHWLFEAVEPGFD-----FRGSRRATMPRLNVEDAGSRLYTYALELIAGKRAE 216
Query: 63 PLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLL 99
P DDM L + + I D D D + L LL
Sbjct: 217 PADDM---LSVVANATIDDPDAPALSDAELYLFFHLL 250
>C126_MYCBO (P63712) Putative cytochrome P450 126 (EC 1.14.-.-)
Length = 414
Score = 30.4 bits (67), Expect = 2.7
Identities = 28/97 (28%), Positives = 37/97 (37%), Gaps = 9/97 (9%)
Query: 4 LGKPEAGLRWFWEPVE-GFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTI 62
LG PE W +E VE GFD + G T + T + L+ G+
Sbjct: 162 LGVPETDRHWLFEAVEPGFD-----FRGSRRATMPRLNVEDAGSRLYTYALELIAGKRAE 216
Query: 63 PLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLL 99
P DDM L + + I D D D + L LL
Sbjct: 217 PADDM---LSVVANATIDDPDAPALSDAELYLFFHLL 250
>INO1_YEAST (P11986) Inositol-3-phosphate synthase (EC 5.5.1.4)
(Myo-inositol-1-phosphate synthase) (MI-1-P synthase)
(IPS)
Length = 536
Score = 29.6 bits (65), Expect = 4.6
Identities = 23/74 (31%), Positives = 35/74 (47%), Gaps = 7/74 (9%)
Query: 57 VGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVR----- 111
VG+ + +D+ ++ L + H RI H+ + A L+I LL M+E RV +
Sbjct: 411 VGDSKVAMDEYYSELMLGGHNRISIHNVCEDSLLATALIIDLLVMTEFCTRVSYKKVDPV 470
Query: 112 SESAGHIS--YPTL 123
E AG YP L
Sbjct: 471 KEDAGKFENFYPVL 484
>SYC1_MYCTU (P96862) Cysteinyl-tRNA synthetase 1 (EC 6.1.1.16)
(Cysteine--tRNA ligase 1) (CysRS 1)
Length = 469
Score = 29.3 bits (64), Expect = 6.0
Identities = 22/69 (31%), Positives = 34/69 (48%), Gaps = 11/69 (15%)
Query: 81 DHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEP 140
DHD A+ AI M+ +LG +D R E R E++ ++ + L +A E
Sbjct: 375 DHDGALRSASAIRAMMGILGCDPLDQRWESRDETSAALAAVDV-------LVQA----EL 423
Query: 141 QTREKLQER 149
Q REK +E+
Sbjct: 424 QNREKAREQ 432
>XA5L_HUMAN (Q9Y247) XAP-5-like protein
Length = 325
Score = 28.9 bits (63), Expect = 7.8
Identities = 26/94 (27%), Positives = 40/94 (41%), Gaps = 21/94 (22%)
Query: 106 VRVEVRSESAGHISYPTLKRVYE-------------HHLTEARRLEEPQTREKLQERGRK 152
V E++S + G ++ +K E HL E +RL++ + RE+ Q R RK
Sbjct: 56 VEAELKSSTVGLVTLNDMKARQEALVRERERQLAKRQHLEE-QRLQQERQREQEQRRERK 114
Query: 153 RAIGKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
R I L++ D LDD AG+
Sbjct: 115 RKIS-------CLSFALDDLDDQADAAEARRAGN 141
>NUP1_YEAST (P20676) Nucleoporin NUP1 (Nuclear pore protein NUP1)
Length = 1076
Score = 28.9 bits (63), Expect = 7.8
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 8/58 (13%)
Query: 107 RVEVRSESAGHISY--------PTLKRVYEHHLTEARRLEEPQTREKLQERGRKRAIG 156
RVE RS S+ S+ P+ K+V+ +L+ A LEE + L RKR G
Sbjct: 13 RVEKRSFSSTLKSFFTNPNKKRPSSKKVFSSNLSYANHLEESDVEDTLHVNKRKRVSG 70
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,285,821
Number of Sequences: 164201
Number of extensions: 922926
Number of successful extensions: 2190
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2185
Number of HSP's gapped (non-prelim): 11
length of query: 194
length of database: 59,974,054
effective HSP length: 104
effective length of query: 90
effective length of database: 42,897,150
effective search space: 3860743500
effective search space used: 3860743500
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149581.3


