

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.10 + phase: 2 /partial
(349 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
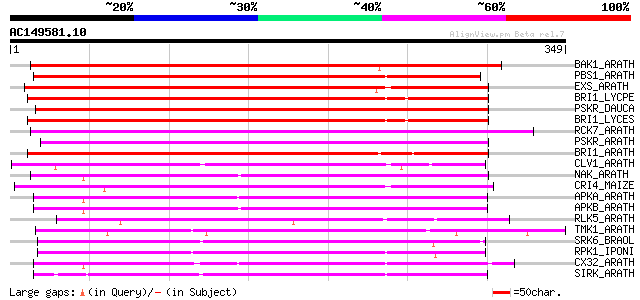
Score E
Sequences producing significant alignments: (bits) Value
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 251 1e-66
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 235 1e-61
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 230 3e-60
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 227 3e-59
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 226 5e-59
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 226 7e-59
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 226 9e-59
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 224 3e-58
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 222 1e-57
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 214 3e-55
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 211 2e-54
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 204 2e-52
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 203 5e-52
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 203 6e-52
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 202 1e-51
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 199 9e-51
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 196 6e-50
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 192 1e-48
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 191 2e-48
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 182 8e-46
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 251 bits (642), Expect = 1e-66
Identities = 132/300 (44%), Positives = 198/300 (66%), Gaps = 4/300 (1%)
Query: 14 ERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKT-MTAKAEMEFAVE 72
+R++L+EL A++NF N +G GGFG VY G+ + G +AVKRLK T E++F E
Sbjct: 275 KRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTE 334
Query: 73 VEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSIT 132
VE++ H+NLL LRGF ERL+VY YM+N S+ + L + S LDWP+R I
Sbjct: 335 VEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIA 394
Query: 133 VGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKG 192
+G+A GLAYLH +P IIHRD+KA+N+LLD EF+A V DFG AKL+ +H+TT V+G
Sbjct: 395 LGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRG 454
Query: 193 TLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIE--KLPGGIKRDIVQWVTPYVQ 250
T+G++APEY GK SE DV+ +G++LLE+I+ ++ + +L ++ WV ++
Sbjct: 455 TIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLK 514
Query: 251 KGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLK-DGVSKRKKE 309
+ + + D L+GN+ E+++ +I +A+ CT SSP +RP M EVV L+ DG+++R +E
Sbjct: 515 EKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLEGDGLAERWEE 574
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 235 bits (599), Expect = 1e-61
Identities = 129/285 (45%), Positives = 178/285 (62%), Gaps = 5/285 (1%)
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQT-SKGVEIAVKRLKTMTAKAEMEFAVEVE 74
+ +EL AT NFH D +GEGGFG VY G+ S G +AVK+L + EF VEV
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVL 133
Query: 75 VLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVG 134
+L + H NL+ L G+ A GD+RL+VY++M SL HLH LDW RM I G
Sbjct: 134 MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAG 193
Query: 135 AAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG-VSHLTTRVKGT 193
AA+GL +LH +ANP +I+RD K+SN+LLD F K++DFG AKL P G SH++TRV GT
Sbjct: 194 AAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGT 253
Query: 194 LGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIE-KLPGGIKRDIVQWVTP-YVQK 251
GY APEYAM G+++ DVYSFG++ LE+I+ +K I+ ++P G ++++V W P + +
Sbjct: 254 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHG-EQNLVAWARPLFNDR 312
Query: 252 GVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVV 296
F +ADP+LKG F L + +A C RP + +VV
Sbjct: 313 RKFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVV 357
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 230 bits (587), Expect = 3e-60
Identities = 122/296 (41%), Positives = 182/296 (61%), Gaps = 7/296 (2%)
Query: 10 DYPWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEF 69
+ P + L +++ AT++F + N IG+GGFG+VY +AVK+L + EF
Sbjct: 899 EQPLLKVRLGDIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREF 958
Query: 70 AVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRM 129
E+E LG+V+H NL+ L G+ + +E+L+VY+YM N SL L Q +LDW +R+
Sbjct: 959 MAEMETLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRL 1018
Query: 130 SITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTR 189
I VGAA GLA+LHH PHIIHRDIKASN+LLD +F+ KVADFG A+LI A SH++T
Sbjct: 1019 KIAVGAARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTV 1078
Query: 190 VKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKP----IEKLPGGIKRDIVQWV 245
+ GT GY+ PEY + + DVYSFG++LLE+++ K+P ++ GG ++V W
Sbjct: 1079 IAGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGG---NLVGWA 1135
Query: 246 TPYVQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
+ +G + DP L ++ IA+ C +P KRP+M++V++ LK+
Sbjct: 1136 IQKINQGKAVDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKALKE 1191
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 227 bits (579), Expect = 3e-59
Identities = 121/293 (41%), Positives = 183/293 (62%), Gaps = 5/293 (1%)
Query: 12 PWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAV 71
P + T +LL ATN FH D+ +G GGFG VY Q G +A+K+L ++ + + EF
Sbjct: 872 PLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTA 931
Query: 72 EVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSI 131
E+E +G+++H+NL+ L G+ G+ERL+VY+YM SL LH + + L+WP R I
Sbjct: 932 EMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKI 991
Query: 132 TVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLT-TRV 190
+GAA GLA+LHH PHIIHRD+K+SNVLLD +A+V+DFG A+L+ A +HL+ + +
Sbjct: 992 AIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTL 1051
Query: 191 KGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQ 250
GT GY+ PEY + S DVYS+G++LLE+++ K+P + G ++V WV +
Sbjct: 1052 AGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFG-DNNLVGWVKLHA- 1109
Query: 251 KGVFKHIADPK-LKGNFDLE-QLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
KG + D + LK + +E +L + +A C D KRP+MI+V+ K+
Sbjct: 1110 KGKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKE 1162
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 226 bits (577), Expect = 5e-59
Identities = 117/285 (41%), Positives = 173/285 (60%)
Query: 17 TLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVL 76
+L ++L++T++F+Q N IG GGFG VY G ++A+KRL T + + EF EVE L
Sbjct: 732 SLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQAEVETL 791
Query: 77 GRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAA 136
R +H NL+ L G+ +++L++Y YM N SL LH ++ LDW R+ I GAA
Sbjct: 792 SRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGAA 851
Query: 137 EGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTLGY 196
EGLAYLH PHI+HRDIK+SN+LL F A +ADFG A+LI +H+TT + GTLGY
Sbjct: 852 EGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTDLVGTLGY 911
Query: 197 LAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKH 256
+ PEY + DVYSFG++LLE+++ ++P++ RD++ WV +
Sbjct: 912 IPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRESE 971
Query: 257 IADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
I DP + E++ V+ IA RC +P RP+ ++V WL++
Sbjct: 972 IFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLEN 1016
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 226 bits (576), Expect = 7e-59
Identities = 121/293 (41%), Positives = 182/293 (61%), Gaps = 5/293 (1%)
Query: 12 PWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAV 71
P + T +LL ATN FH D+ +G GGFG VY Q G +A+K+L ++ + + EF
Sbjct: 872 PLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTA 931
Query: 72 EVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSI 131
E+E +G+++H+NL+ L G+ G+ERL+VY+YM SL LH + L+WP R I
Sbjct: 932 EMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKI 991
Query: 132 TVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLT-TRV 190
+GAA GLA+LHH PHIIHRD+K+SNVLLD +A+V+DFG A+L+ A +HL+ + +
Sbjct: 992 AIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTL 1051
Query: 191 KGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQ 250
GT GY+ PEY + S DVYS+G++LLE+++ K+P + G ++V WV +
Sbjct: 1052 AGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFG-DNNLVGWVKLHA- 1109
Query: 251 KGVFKHIADPK-LKGNFDLE-QLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
KG + D + LK + +E +L + +A C D KRP+MI+V+ K+
Sbjct: 1110 KGKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKE 1162
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 226 bits (575), Expect = 9e-59
Identities = 128/320 (40%), Positives = 183/320 (57%), Gaps = 4/320 (1%)
Query: 14 ERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEI-AVKRLKTMTAKAEMEFAVE 72
+ +T +EL AT NF D +GEGGFG V+ G K ++ A+K+L + EF VE
Sbjct: 89 QTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVE 148
Query: 73 VEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSIT 132
V L H NL+ L GF A GD+RL+VY+YM SL HLH + LDW RM I
Sbjct: 149 VLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIA 208
Query: 133 VGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG-VSHLTTRVK 191
GAA GL YLH P +I+RD+K SN+LL ++Q K++DFG AK+ P+G +H++TRV
Sbjct: 209 AGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTRVM 268
Query: 192 GTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTP-YVQ 250
GT GY AP+YAM G+++ D+YSFG++LLE+I+ +K I+ +++V W P +
Sbjct: 269 GTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLFKD 328
Query: 251 KGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD-GVSKRKKE 309
+ F + DP L+G + + L + I+ C P RP + +VV L SK
Sbjct: 329 RRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLALNFLASSKYDPN 388
Query: 310 IPNLSNNKGHDEENDENYEE 329
P+ S+ K D + EE
Sbjct: 389 SPSSSSGKNPSFHRDRDDEE 408
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 224 bits (570), Expect = 3e-58
Identities = 115/282 (40%), Positives = 169/282 (59%)
Query: 20 ELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVLGRV 79
+LL +TN+F Q N IG GGFG VY G ++A+K+L + E EF EVE L R
Sbjct: 726 DLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREFEAEVETLSRA 785
Query: 80 RHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGL 139
+H NL+ LRGF ++RL++Y YM N SL LH + LL W R+ I GAA+GL
Sbjct: 786 QHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRIAQGAAKGL 845
Query: 140 AYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTLGYLAP 199
YLH +PHI+HRDIK+SN+LLD F + +ADFG A+L+ +H++T + GTLGY+ P
Sbjct: 846 LYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDLVGTLGYIPP 905
Query: 200 EYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKHIAD 259
EY + DVYSFG++LLE+++ K+P++ RD++ WV + + D
Sbjct: 906 EYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHESRASEVFD 965
Query: 260 PKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
P + + +++ V+ IA C +P +RP+ ++V WL D
Sbjct: 966 PLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSWLDD 1007
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 222 bits (566), Expect = 1e-57
Identities = 118/293 (40%), Positives = 184/293 (62%), Gaps = 5/293 (1%)
Query: 12 PWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAV 71
P + T +LL+ATN FH D+ IG GGFG VY G +A+K+L ++ + + EF
Sbjct: 867 PLRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDREFMA 926
Query: 72 EVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSI 131
E+E +G+++H+NL+ L G+ GDERL+VY++M SL LH + L+W R I
Sbjct: 927 EMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKI 986
Query: 132 TVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLT-TRV 190
+G+A GLA+LHH +PHIIHRD+K+SNVLLD +A+V+DFG A+L+ A +HL+ + +
Sbjct: 987 AIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTL 1046
Query: 191 KGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQ 250
GT GY+ PEY + S DVYS+G++LLE+++ K+P + P ++V WV + +
Sbjct: 1047 AGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDS-PDFGDNNLVGWVKQHAK 1105
Query: 251 KGVFKHIADPKL-KGNFDLE-QLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
+ + DP+L K + LE +L + +AV C D +RP+M++V+ K+
Sbjct: 1106 LRI-SDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFKE 1157
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 214 bits (545), Expect = 3e-55
Identities = 122/311 (39%), Positives = 182/311 (58%), Gaps = 18/311 (5%)
Query: 2 VDDKKNIRDYPWERYTLKELLRATNN----FHQDNKIGEGGFGSVYWGQTSKGVEIAVKR 57
++ KKN + W+ ++L + + ++N IG+GG G VY G V++A+KR
Sbjct: 662 MNKKKNQKSLAWKLTAFQKLDFKSEDVLECLKEENIIGKGGAGIVYRGSMPNNVDVAIKR 721
Query: 58 LKTM-TAKAEMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQ 116
L T +++ F E++ LGR+RH++++ L G+ A D L++Y+YM N SL LHG
Sbjct: 722 LVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGYVANKDTNLLLYEYMPNGSLGELLHGS 781
Query: 117 LASDCLLDWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFA 176
L W R + V AA+GL YLHH+ +P I+HRD+K++N+LLD++F+A VADFG A
Sbjct: 782 KGGH--LQWETRHRVAVEAAKGLCYLHHDCSPLILHRDVKSNNILLDSDFEAHVADFGLA 839
Query: 177 K-LIPAGVSHLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPG 235
K L+ S + + G+ GY+APEYA KV E DVYSFG++LLE+I+ KKP+ +
Sbjct: 840 KFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIAGKKPVGEFGE 899
Query: 236 GIKRDIVQWV-------TPYVQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDK 288
G+ DIV+WV T + I DP+L G + L + V IA+ C +
Sbjct: 900 GV--DIVRWVRNTEEEITQPSDAAIVVAIVDPRLTG-YPLTSVIHVFKIAMMCVEEEAAA 956
Query: 289 RPSMIEVVEWL 299
RP+M EVV L
Sbjct: 957 RPTMREVVHML 967
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 211 bits (538), Expect = 2e-54
Identities = 116/300 (38%), Positives = 179/300 (59%), Gaps = 13/300 (4%)
Query: 14 ERYTLKELLRATNNFHQDNKIGEGGFGSVYWG----------QTSKGVEIAVKRLKTMTA 63
+ ++L EL AT NF D+ +GEGGFG V+ G + G+ IAVKRL
Sbjct: 54 KNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGF 113
Query: 64 KAEMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLL 123
+ E+ E+ LG++ H NL+ L G+ + RL+VY++M+ SL HL + L
Sbjct: 114 QGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPL 173
Query: 124 DWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGV 183
W R+ + +GAA GLA+LH+ A P +I+RD KASN+LLD+ + AK++DFG A+ P G
Sbjct: 174 SWNTRVRMALGAARGLAFLHN-AQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPMGD 232
Query: 184 -SHLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIV 242
SH++TRV GT GY APEY G +S DVYSFG++LLE++S ++ I+K + ++V
Sbjct: 233 NSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHNLV 292
Query: 243 QWVTPYV-QKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKD 301
W PY+ K + DP+L+G + L + + ++A+ C RP+M E+V+ +++
Sbjct: 293 DWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIVKTMEE 352
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 204 bits (520), Expect = 2e-52
Identities = 120/306 (39%), Positives = 183/306 (59%), Gaps = 8/306 (2%)
Query: 4 DKKNIRDYPWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRL--KTM 61
D ++++ + ++ +EL +AT F +D+++G+G F V+ G G +AVKR +
Sbjct: 481 DMEDLKIRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASD 540
Query: 62 TAKAEMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLAS-D 120
K+ EF E+++L R+ H +LL L G+ G ERL+VY++M++ SL HLHG+ +
Sbjct: 541 VKKSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLK 600
Query: 121 CLLDWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIP 180
L+W RR++I V AA G+ YLH A P +IHRDIK+SN+L+D + A+VADFG + L P
Sbjct: 601 KRLNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGP 660
Query: 181 A-GVSHLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIE-KLPGGIK 238
A + L+ GTLGYL PEY ++ DVYSFG++LLEI+S +K I+ + G
Sbjct: 661 ADSGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEG-- 718
Query: 239 RDIVQWVTPYVQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEW 298
+IV+W P ++ G I DP L DLE LK + +A +C RPSM +V
Sbjct: 719 -NIVEWAVPLIKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTA 777
Query: 299 LKDGVS 304
L+ ++
Sbjct: 778 LEHALA 783
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 203 bits (517), Expect = 5e-52
Identities = 114/297 (38%), Positives = 175/297 (58%), Gaps = 13/297 (4%)
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWG----------QTSKGVEIAVKRLKTMTAKA 65
++ EL AT NF D+ +GEGGFG V+ G + G+ IAVK+L +
Sbjct: 56 FSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGWQG 115
Query: 66 EMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDW 125
E+ EV LG+ H++L+ L G+ + RL+VY++M SL HL + L W
Sbjct: 116 HQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLSW 175
Query: 126 PRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG-VS 184
R+ + +GAA+GLA+LH + +I+RD K SN+LLD+E+ AK++DFG AK P G S
Sbjct: 176 KLRLKVALGAAKGLAFLH-SSETRVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPIGDKS 234
Query: 185 HLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQW 244
H++TRV GT GY APEY G ++ DVYSFG++LLE++S ++ ++K +R++V+W
Sbjct: 235 HVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERNLVEW 294
Query: 245 VTPY-VQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLK 300
PY V K + D +L+ + +E+ V +++RC + RP+M EVV L+
Sbjct: 295 AKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHLE 351
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 203 bits (516), Expect = 6e-52
Identities = 115/297 (38%), Positives = 174/297 (57%), Gaps = 13/297 (4%)
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWG----------QTSKGVEIAVKRLKTMTAKA 65
+T EL AT NF D+ +GEGGFGSV+ G + GV IAVK+L +
Sbjct: 57 FTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQDGWQG 116
Query: 66 EMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDW 125
E+ EV LG+ H NL+ L G+ + RL+VY++M SL HL + + L W
Sbjct: 117 HQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQPLSW 176
Query: 126 PRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG-VS 184
R+ + +GAA+GLA+LH+ A +I+RD K SN+LLD+E+ AK++DFG AK P G S
Sbjct: 177 TLRLKVALGAAKGLAFLHN-AETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTGDKS 235
Query: 185 HLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQW 244
H++TR+ GT GY APEY G ++ DVYS+G++LLE++S ++ ++K ++ +V+W
Sbjct: 236 HVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQKLVEW 295
Query: 245 VTPYV-QKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLK 300
P + K + D +L+ + +E+ V +A+RC RP+M EVV L+
Sbjct: 296 ARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHLE 352
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 202 bits (514), Expect = 1e-51
Identities = 117/298 (39%), Positives = 170/298 (56%), Gaps = 16/298 (5%)
Query: 30 QDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEME----------FAVEVEVLGRV 79
+ N IG G G VY + G +AVK+L + E FA EVE LG +
Sbjct: 685 EKNVIGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTI 744
Query: 80 RHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGL 139
RHK+++ L + GD +L+VY+YM N SL LHG +L WP R+ I + AAEGL
Sbjct: 745 RHKSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGL 804
Query: 140 AYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAK---LIPAGVSHLTTRVKGTLGY 196
+YLHH+ P I+HRD+K+SN+LLD+++ AKVADFG AK + + + + G+ GY
Sbjct: 805 SYLHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCGY 864
Query: 197 LAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKH 256
+APEY +V+E D+YSFG++LLE+++ K+P + G +D+ +WV + K +
Sbjct: 865 IAPEYVYTLRVNEKSDIYSFGVVLLELVTGKQPTDSELG--DKDMAKWVCTALDKCGLEP 922
Query: 257 IADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGVSKRKKEIPNLS 314
+ DPKL F E++ VI I + CT P RPSM +VV L++ PN S
Sbjct: 923 VIDPKLDLKFK-EEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQEVSGAVPCSSPNTS 979
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 199 bits (506), Expect = 9e-51
Identities = 127/350 (36%), Positives = 190/350 (54%), Gaps = 20/350 (5%)
Query: 17 TLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKT--MTAKAEMEFAVEVE 74
+++ L TNNF DN +G GGFG VY G+ G +IAVKR++ + K EF E+
Sbjct: 577 SIQVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIA 636
Query: 75 VLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCL--LDWPRRMSIT 132
VL +VRH++L+ L G+ G+E+L+VY+YM +L HL + + + L L W +R+++
Sbjct: 637 VLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLF-EWSEEGLKPLLWKQRLTLA 695
Query: 133 VGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKG 192
+ A G+ YLH A+ IHRD+K SN+LL + +AKVADFG +L P G + TR+ G
Sbjct: 696 LDVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRIAG 755
Query: 193 TLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWV-TPYVQK 251
T GYLAPEYA+ G+V+ DVYSFG++L+E+I+ +K +++ +V W Y+ K
Sbjct: 756 TFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINK 815
Query: 252 -GVFKHIADPKLKGNFDLEQLKSVIMIAV---RCTDSSPDKRPSMIEVVEWLKDGVSKRK 307
FK D + + D E L SV +A C P +RP M V L V K
Sbjct: 816 EASFKKAIDTTI--DLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSLVELWK 873
Query: 308 KEIPNLSNNKGHDEEND--------ENYEEFVTMQSNNLKILSDNDRRKM 349
N + G D + + YE ++S+ +L D +M
Sbjct: 874 PSDQNPEDIYGIDLDMSLPQALKKWQAYEGRSDLESSTSSLLPSLDNTQM 923
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 196 bits (499), Expect = 6e-50
Identities = 110/290 (37%), Positives = 169/290 (57%), Gaps = 10/290 (3%)
Query: 18 LKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVLG 77
++ +++AT NF NK+G+GGFG VY G+ G EIAVKRL + + EF EV ++
Sbjct: 518 METVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEVTLIA 577
Query: 78 RVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAE 137
R++H NL+ + G GDE++++Y+Y+ N SL ++L G+ L+W R IT G A
Sbjct: 578 RLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRS-KLNWNERFDITNGVAR 636
Query: 138 GLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTT-RVKGTLGY 196
GL YLH ++ IIHRD+K SN+LLD K++DFG A++ + T +V GT GY
Sbjct: 637 GLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGTYGY 696
Query: 197 LAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKH 256
++PEYAM+G SE DV+SFG+++LEI+S KK + D++ +V ++G
Sbjct: 697 MSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGRALE 756
Query: 257 IADPKLKGN-------FDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWL 299
I DP + + F +++ I I + C + RP+M VV W+
Sbjct: 757 IVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVV-WM 805
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 192 bits (488), Expect = 1e-48
Identities = 113/289 (39%), Positives = 167/289 (57%), Gaps = 9/289 (3%)
Query: 18 LKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLK-TMTAKAEMEFAVEVEVL 76
L ++L AT N + IG+G G++Y S AVK+L T + E+E +
Sbjct: 806 LNKVLEATENLNDKYVIGKGAHGTIYKATLSPDKVYAVKKLVFTGIKNGSVSMVREIETI 865
Query: 77 GRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAA 136
G+VRH+NL+ L F+ + LI+Y YM N SL LH + LDW R +I VG A
Sbjct: 866 GKVRHRNLIKLEEFWLRKEYGLILYTYMENGSLHDILH-ETNPPKPLDWSTRHNIAVGTA 924
Query: 137 EGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHL-TTRVKGTLG 195
GLAYLH + +P I+HRDIK N+LLD++ + ++DFG AKL+ + + + V+GT+G
Sbjct: 925 HGLAYLHFDCDPAIVHRDIKPMNILLDSDLEPHISDFGIAKLLDQSATSIPSNTVQGTIG 984
Query: 196 YLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWV-TPYVQKGVF 254
Y+APE A S DVYS+G++LLE+I+ KK ++ G + DIV WV + + Q G
Sbjct: 985 YMAPENAFTTVKSRESDVYSYGVVLLELITRKKALDPSFNG-ETDIVGWVRSVWTQTGEI 1043
Query: 255 KHIADPKLKGNF----DLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWL 299
+ I DP L +EQ+ + +A+RC + DKRP+M +VV+ L
Sbjct: 1044 QKIVDPSLLDELIDSSVMEQVTEALSLALRCAEKEVDKRPTMRDVVKQL 1092
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 191 bits (486), Expect = 2e-48
Identities = 119/315 (37%), Positives = 174/315 (54%), Gaps = 20/315 (6%)
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWG----------QTSKGVEIAVKRLKTMTAKA 65
Y +L AT NF D+ +G+GGFG VY G + G+ +A+KRL + + +
Sbjct: 74 YNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQG 133
Query: 66 EMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDW 125
E+ EV LG + H+NL+ L G+ E L+VY++M SL +HL + + W
Sbjct: 134 FAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFRR---NDPFPW 190
Query: 126 PRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGV-S 184
R+ I +GAA GLA+LH +I+RD KASN+LLD+ + AK++DFG AKL PA S
Sbjct: 191 DLRIKIVIGAARGLAFLH-SLQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKS 249
Query: 185 HLTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIE-KLPGGIKRDIVQ 243
H+TTR+ GT GY APEY G + DV++FG++LLEI++ K P G + +V
Sbjct: 250 HVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRG-QESLVD 308
Query: 244 WVTPYV-QKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDG 302
W+ P + K K I D +KG + + + I + C + P RP M EVVE L+
Sbjct: 309 WLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVLEH- 367
Query: 303 VSKRKKEIPNLSNNK 317
+ +PN S+ K
Sbjct: 368 -IQGLNVVPNRSSTK 381
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 182 bits (463), Expect = 8e-46
Identities = 106/286 (37%), Positives = 169/286 (59%), Gaps = 7/286 (2%)
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
+ E++ TNNF + IG+GGFG VY G + G ++AVK L +A+ EF EV++
Sbjct: 564 FKYSEVVNITNNF--ERVIGKGGFGKVYHGVIN-GEQVAVKVLSEESAQGYKEFRAEVDL 620
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGA 135
L RV H NL L G+ + +++Y+YM+N +L +L G+ + +L W R+ I++ A
Sbjct: 621 LMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSF--ILSWEERLKISLDA 678
Query: 136 AEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPA-GVSHLTTRVKGTL 194
A+GL YLH+ P I+HRD+K +N+LL+ + QAK+ADFG ++ G ++T V G++
Sbjct: 679 AQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAGSI 738
Query: 195 GYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVF 254
GYL PEY +++E DVYS G++LLE+I+ + I K I V + G
Sbjct: 739 GYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTE-KVHISDHVRSILANGDI 797
Query: 255 KHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLK 300
+ I D +L+ +D+ + IA+ CT+ + +RP+M +VV LK
Sbjct: 798 RGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMELK 843
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,620,184
Number of Sequences: 164201
Number of extensions: 1818954
Number of successful extensions: 8472
Number of sequences better than 10.0: 1641
Number of HSP's better than 10.0 without gapping: 1157
Number of HSP's successfully gapped in prelim test: 485
Number of HSP's that attempted gapping in prelim test: 5184
Number of HSP's gapped (non-prelim): 1844
length of query: 349
length of database: 59,974,054
effective HSP length: 111
effective length of query: 238
effective length of database: 41,747,743
effective search space: 9935962834
effective search space used: 9935962834
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC149581.10


