

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
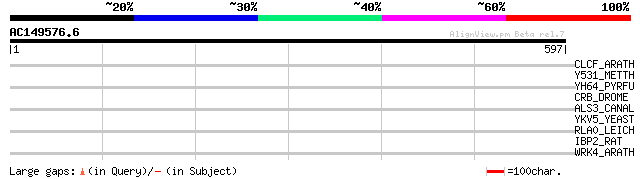
CLCF_ARATH (Q8RXR2) Chloride channel protein CLC-f (AtCLC-f) 35 0.43
Y531_METTH (O26631) Hypothetical protein MTH531 33 2.1
YH64_PYRFU (Q8U053) Hypothetical UPF0173 metal-dependent hydrola... 33 2.8
CRB_DROME (P10040) Crumbs protein precursor (95F) 33 2.8
ALS3_CANAL (O74623) Agglutinin-like protein 3 precursor 32 3.6
YKV5_YEAST (P28273) Hypothetical 140.4 kDa protein in URA1-DOA1 ... 32 6.2
RLA0_LEICH (P39096) 60S acidic ribosomal protein P0 32 6.2
IBP2_RAT (P12843) Insulin-like growth factor binding protein 2 p... 32 6.2
WRK4_ARATH (Q9XI90) Probable WRKY transcription factor 4 (WRKY D... 31 8.1
>CLCF_ARATH (Q8RXR2) Chloride channel protein CLC-f (AtCLC-f)
Length = 781
Score = 35.4 bits (80), Expect = 0.43
Identities = 28/101 (27%), Positives = 42/101 (40%), Gaps = 20/101 (19%)
Query: 412 WGGTANRGRLKLRVGQPPENW--------TSGVDLGRLLDLLE-LDLVTTNETLQDSGQE 462
W GT N G LR+ + + W T GV +G + LLE LD + + + Q G +
Sbjct: 161 WAGTPNEGAAWLRLQRLADTWHRILLIPVTGGVIVGMMHGLLEILDQIRQSNSSQRQGLD 220
Query: 463 QMNG-----------STAGIGSTVGESSPTVPIKEKLEESF 492
+ G T G G ++G P+V I + F
Sbjct: 221 FLAGIYPVIKAIQAAVTLGTGCSLGPEGPSVDIGKSCANGF 261
>Y531_METTH (O26631) Hypothetical protein MTH531
Length = 312
Score = 33.1 bits (74), Expect = 2.1
Identities = 41/163 (25%), Positives = 68/163 (41%), Gaps = 21/163 (12%)
Query: 220 LGAIVRSRTGNREVGFLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDD 279
+GA R R + L NR + D+ + H LP L A +R +
Sbjct: 10 IGAGNAGRPAARLLNHLNNRVLVNDV-----RELHELP--------LKAQKRIAEMEDEG 56
Query: 280 LWYGIFAGTNPETFVRADGAFIP--FAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSL 337
+ + F G + E + AD FI +D + ++ R GD+ + D+ +N L
Sbjct: 57 VMFR-FGGHSMEDILWADAVFISPNIPQDAPVRKMV---REAGDLVHITTSDIGRTLNEL 112
Query: 338 IGRQVIKVGRSSGLTTGTIMAYALEYNDEKGICF--LTDFLVV 378
IG ++ V + G TT T M + + + + F L D LV+
Sbjct: 113 IGLPMVGVAGTDGKTTTTNMIDHILSSRYRTVSFSSLQDSLVI 155
>YH64_PYRFU (Q8U053) Hypothetical UPF0173 metal-dependent hydrolase
PF1764
Length = 225
Score = 32.7 bits (73), Expect = 2.8
Identities = 20/61 (32%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 68 AAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRI 127
A D ANY KGV ET+G + G +E++ + A+HS ++S+G A G+ +
Sbjct: 71 AMYDLANYIAEKYKGV--ETIG-MNYGPTKVDEVEIVQVPAWHSSSDGKYSIGNACGYIV 127
Query: 128 R 128
+
Sbjct: 128 K 128
>CRB_DROME (P10040) Crumbs protein precursor (95F)
Length = 2139
Score = 32.7 bits (73), Expect = 2.8
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 4/61 (6%)
Query: 161 LEGPGGVWCDVDVVEFSYYG----APAPTPKEQLYTELADGLRGSDSCVGSGSQVASQET 216
L+GPG + + V+ +G P P L+T L D +D C+G G+ +S E
Sbjct: 229 LDGPGLQFVNNSTVQNVVFGHCPLTPGPCSDHDLFTRLPDNFCLNDPCMGHGTCSSSPEG 288
Query: 217 Y 217
Y
Sbjct: 289 Y 289
>ALS3_CANAL (O74623) Agglutinin-like protein 3 precursor
Length = 1119
Score = 32.3 bits (72), Expect = 3.6
Identities = 34/124 (27%), Positives = 51/124 (40%), Gaps = 9/124 (7%)
Query: 447 LDLVTTNETLQDSG----QEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHV 502
+D TT T Q + ++ T T +++P+VP E E N E+
Sbjct: 897 VDSTTTESTSQSPSGIFSESGVSVETESSTVTTAQTNPSVPTTES--EVVFTTKGNNENG 954
Query: 503 PVEEPSTIVKPSL-RPCEFHIRNEIETVPNVEHQFIRT--SFAGKSPVHQSFLKEDMQFK 559
P E PST VK S+ EF T ++E++ I T S SP+ S E
Sbjct: 955 PYESPSTNVKSSMDENSEFTTSTAASTSTDIENETIATTGSVEASSPIISSSADETTTVT 1014
Query: 560 SLSE 563
+ +E
Sbjct: 1015 TTAE 1018
>YKV5_YEAST (P28273) Hypothetical 140.4 kDa protein in URA1-DOA1
intergenic region
Length = 1286
Score = 31.6 bits (70), Expect = 6.2
Identities = 21/68 (30%), Positives = 36/68 (52%), Gaps = 2/68 (2%)
Query: 510 IVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAG-KSPVHQSFLKEDMQFKSLSELRNEP 568
+++ + PC F I E ET+ V+ +F+ S K+ + QSF +ED+ + LR E
Sbjct: 532 VIEENQEPCSF-ILGEPETILKVKKRFLELSKNSIKNLLSQSFSREDIVLERYLNLRYEG 590
Query: 569 DEDNFVSL 576
E + + L
Sbjct: 591 TETSLMIL 598
>RLA0_LEICH (P39096) 60S acidic ribosomal protein P0
Length = 322
Score = 31.6 bits (70), Expect = 6.2
Identities = 25/90 (27%), Positives = 42/90 (45%), Gaps = 10/90 (11%)
Query: 414 GTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQE----QMNGSTA 469
G +N + L G P + + + +LL + + T+ E + +G+E +NG A
Sbjct: 224 GLSNVAAMALGAGIPTSSTIGPMLVDAFKNLLAVSVATSYEFEEHNGKELREAAINGLLA 283
Query: 470 GIGSTVGE------SSPTVPIKEKLEESFE 493
G GS E ++P+ KE+ EES E
Sbjct: 284 GSGSAAAEPAAAAPAAPSAAAKEEPEESDE 313
>IBP2_RAT (P12843) Insulin-like growth factor binding protein 2
precursor (IGFBP-2) (IBP-2) (IGF-binding protein 2)
(BRL-BP)
Length = 304
Score = 31.6 bits (70), Expect = 6.2
Identities = 28/109 (25%), Positives = 41/109 (36%), Gaps = 9/109 (8%)
Query: 422 KLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQMNGSTAGIGSTVGESSPT 481
K RVG P+ D D E LV E D + GS+AG
Sbjct: 117 KRRVGATPQQVADSED-----DHSEGGLV---ENHVDGTMNMLGGSSAGRKPPKSGMKEL 168
Query: 482 VPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLR-PCEFHIRNEIETV 529
+EK+ E +H+ +EEP + P R PC+ + +E +
Sbjct: 169 AVFREKVNEQHRQMGKGAKHLSLEEPKKLRPPPARTPCQQELDQVLERI 217
>WRK4_ARATH (Q9XI90) Probable WRKY transcription factor 4 (WRKY
DNA-binding protein 4)
Length = 514
Score = 31.2 bits (69), Expect = 8.1
Identities = 17/76 (22%), Positives = 31/76 (40%)
Query: 191 YTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQ 250
+++L G S + + + A+ Y LG S +G+ + F NR + +
Sbjct: 61 FSQLLAGAMSSPATAAAAAAAATASDYQRLGEGTNSSSGDVDPRFKQNRPTGLMISQSQS 120
Query: 251 KMFHPLPPSLGPGVYL 266
+PP L P + L
Sbjct: 121 PSMFTVPPGLSPAMLL 136
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,860,531
Number of Sequences: 164201
Number of extensions: 3480987
Number of successful extensions: 7200
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 7199
Number of HSP's gapped (non-prelim): 9
length of query: 597
length of database: 59,974,054
effective HSP length: 116
effective length of query: 481
effective length of database: 40,926,738
effective search space: 19685760978
effective search space used: 19685760978
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149576.6


