

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
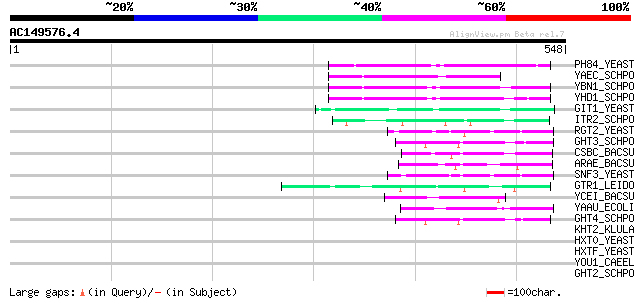
Score E
Sequences producing significant alignments: (bits) Value
PH84_YEAST (P25297) Inorganic phosphate transporter PHO84 140 9e-33
YAEC_SCHPO (Q09852) Putative inorganic phosphate transporter C23... 125 4e-28
YBN1_SCHPO (O42885) Putative inorganic phosphate transporter C8E... 120 7e-27
YHD1_SCHPO (Q9P6J9) Putative inorganic phosphate transporter C16... 119 2e-26
GIT1_YEAST (P25346) Probable metabolite transport protein GIT1 65 3e-10
ITR2_SCHPO (P87110) Myo-inositol transporter 2 52 4e-06
RGT2_YEAST (Q12300) High-affinity glucose transporter RGT2 49 4e-05
GHT3_SCHPO (Q92339) High-affinity gluconate transporter ght3 (He... 48 6e-05
CSBC_BACSU (P46333) Probable metabolite transport protein csbC 47 1e-04
ARAE_BACSU (P96710) Arabinose-proton symporter (Arabinose transp... 47 1e-04
SNF3_YEAST (P10870) High-affinity glucose transporter SNF3 46 2e-04
GTR1_LEIDO (Q01440) Membrane transporter D1 46 2e-04
YCEI_BACSU (O34691) Hypothetical metabolite transport protein yceI 45 4e-04
YAAU_ECOLI (P31679) Hypothetical metabolite transport protein yaaU 45 5e-04
GHT4_SCHPO (O59932) High-affinity hexose transporter ght4 (Hexos... 45 5e-04
KHT2_KLULA (P53387) Hexose transporter 2 44 0.001
HXT0_YEAST (P43581) Hexose transporter HXT10 44 0.001
HXTF_YEAST (P47185) Hexose transporter HXT16 43 0.002
YOU1_CAEEL (P30638) Hypothetical protein ZK637.1 in chromosome III 43 0.002
GHT2_SCHPO (O74969) High-affinity glucose transporter ght2 (Hexo... 43 0.002
>PH84_YEAST (P25297) Inorganic phosphate transporter PHO84
Length = 587
Score = 140 bits (353), Expect = 9e-33
Identities = 80/221 (36%), Positives = 118/221 (53%), Gaps = 10/221 (4%)
Query: 315 WRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAV 374
W++G+ L + +WF LD+ FY L S + + + ++VY++ + A IL
Sbjct: 341 WKYGKILLGTAGSWFTLDVAFYGLSL-NSAVILQTIGYAGSKNVYKKLYDTAVGNLILIC 399
Query: 375 CSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY 434
++PGY+ +V+ +D +GR IQ+ GF + F +GF Y+ K +H G + +Y
Sbjct: 400 AGSLPGYWVSVFTVDIIGRKPIQLAGFIILTALFCVIGFAYH----KLGDH---GLLALY 452
Query: 435 GLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGF-LWASHKEKE 493
+ FF NFGPNTTTFIVP E FP R+RST HGIS A GKVGAII H
Sbjct: 453 VICQFFQNFGPNTTTFIVPGECFPTRYRSTAHGISAASGKVGAIIAQTALGTLIDHNCAR 512
Query: 494 EGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
+G P + + I ++G+F T ET ++LEE
Sbjct: 513 DGKPTNCWLPHVMEIFALFMLLGIFTT-LLIPETKRKTLEE 552
>YAEC_SCHPO (Q09852) Putative inorganic phosphate transporter
C23D3.12
Length = 559
Score = 125 bits (313), Expect = 4e-28
Identities = 68/170 (40%), Positives = 96/170 (56%), Gaps = 8/170 (4%)
Query: 315 WRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAV 374
W H + L A + +WFLLDI FY L QS I K + ++ Y A I+A+
Sbjct: 331 WHHFKHLLATAVSWFLLDIAFYGVNLNQSVILKA-IGFSSGKNEYHTLMRGAIGNLIIAI 389
Query: 375 CSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY 434
+PGY+FTV+ ++++GR IQ+ G F + F L + + T G +
Sbjct: 390 AGYVPGYWFTVFLVEKLGRKWIQLQGLFITGLMFAILAGSWDTISTGGR-------FACF 442
Query: 435 GLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGF 484
+A FF+NFGPN TTF+ PAE+FPAR R T HG+S A+GK GAI+ S+ F
Sbjct: 443 VIAQFFSNFGPNATTFLYPAEVFPARVRGTAHGLSAALGKCGAILASLLF 492
>YBN1_SCHPO (O42885) Putative inorganic phosphate transporter
C8E4.01c
Length = 572
Score = 120 bits (302), Expect = 7e-27
Identities = 77/220 (35%), Positives = 109/220 (49%), Gaps = 18/220 (8%)
Query: 315 WRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAV 374
WRH + L S WFLLDI FY L QS I K + + Y+ A I+AV
Sbjct: 342 WRHFKHLLGTSVCWFLLDIAFYGVNLNQSVILKN-IGFSTGTNEYRTLMKNAIGNLIIAV 400
Query: 375 CSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY 434
+PGY+F V+ ++ +GR IQ+ GF + F L W + G +
Sbjct: 401 AGYVPGYWFNVFLVEILGRKWIQLQGFVITGLMFAILA----GRWNE---ISTGGRFACF 453
Query: 435 GLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEE 494
+A F+NFGPN+TTFI PAE+FPAR R T HG+S A+GK GAI+ S+ F
Sbjct: 454 VIAQLFSNFGPNSTTFIYPAEVFPARVRGTAHGVSAALGKCGAILASLLF---------- 503
Query: 495 GYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
+ G+ +++ + C+ G + ET GR +E
Sbjct: 504 NFLTGVIGYGNVMWIFCGCMWGGILFTLLLPETKGRDADE 543
>YHD1_SCHPO (Q9P6J9) Putative inorganic phosphate transporter
C1683.01
Length = 573
Score = 119 bits (299), Expect = 2e-26
Identities = 80/220 (36%), Positives = 108/220 (48%), Gaps = 18/220 (8%)
Query: 315 WRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAV 374
W H + L S WFLLDI FY L QS I K + + Y+ A I+AV
Sbjct: 343 WHHFKHLLGTSVCWFLLDIAFYGVNLNQSVILKN-IGFSSGTNEYRTLMKNAIGNLIIAV 401
Query: 375 CSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY 434
+PGY+F V+ ++ +GR IQ+ GF + F L W + G +
Sbjct: 402 AGYVPGYWFNVFLVEILGRKWIQLQGFVITGLMFAILA----GRWNE---ISTGGRFACF 454
Query: 435 GLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEE 494
+A F+NFGPN+TTFI PAE+FPAR R T HGIS A+GK GAI+ S+ F + +
Sbjct: 455 VIAQLFSNFGPNSTTFIYPAEVFPARVRGTAHGISAALGKCGAILASLLFNFLT------ 508
Query: 495 GYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
IG + I G C+ G + ET GR +E
Sbjct: 509 ---SIIGYGNVMWIFCG-CMWGGILFTLLLPETKGRDADE 544
>GIT1_YEAST (P25346) Probable metabolite transport protein GIT1
Length = 518
Score = 65.5 bits (158), Expect = 3e-10
Identities = 53/236 (22%), Positives = 92/236 (38%), Gaps = 25/236 (10%)
Query: 303 NVAYPLLSREFLWRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEA 362
N+ Y L+ +F W+ L WF+ D V + +F S I + +++D E
Sbjct: 255 NIPY-FLALKFYWKR---LLGTCGTWFMYDFVTFPNGIFSSTIISSVIKDQNDLVKVAEW 310
Query: 363 FHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKG 422
L + A+L V Y DR+GR M GF + +G Y
Sbjct: 311 NLLLGVLAVLGVP-------IGAYLSDRIGRKYTLMFGFSGYIIFGLIIGCAYDQL---- 359
Query: 423 ENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSV 482
F++ Y N GP ++ +E R +G+S GK+G+++G
Sbjct: 360 -KKITPLFIIFYAFMNMLGNAGPGDMLGVISSEASATAVRGVFYGLSAVTGKIGSVVGVE 418
Query: 483 GFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDE 538
F + +G + + II ++G+ +TYFF ++ L + + E
Sbjct: 419 CF---------QPIRDNLGARWTFIIAAICGLIGIIITYFFVPHSLESDLMKQDVE 465
>ITR2_SCHPO (P87110) Myo-inositol transporter 2
Length = 557
Score = 52.0 bits (123), Expect = 4e-06
Identities = 55/229 (24%), Positives = 88/229 (38%), Gaps = 45/229 (19%)
Query: 319 RDLF-ACSANWFLL-----DIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAIL 372
R LF C WF I ++S ++FQS +K ++ +
Sbjct: 331 RSLFIGCFLQWFQQFSGTNAIQYFSAIIFQSVGFKNSIS--------------------V 370
Query: 373 AVCSTIPGYFFTVY---FIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKG 429
++ + FT+ FIDR+GR +I + M Y+ + N G
Sbjct: 371 SIVVGATNFVFTIVAFMFIDRIGRRRILLCTSAVMIAGLALCAIAYHFLPADTTQNTNSG 430
Query: 430 --FMVIYGLAFFFANFGPNTTTFIVP---AELFPARFRSTCHGISGAVGKVGAIIGSVGF 484
++V+ + F A++ +P AELFP R+ G S A+ VG +I S F
Sbjct: 431 WQYVVLASIIIFLASYASGIGN--IPWQQAELFPMEVRALGAGFSTAINWVGNLIISASF 488
Query: 485 LWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE 533
L + I + + G C VG+ +YF E G S+E
Sbjct: 489 LTMM---------ESITPTGTFALFAGFCFVGLVTSYFTYPELAGMSIE 528
>RGT2_YEAST (Q12300) High-affinity glucose transporter RGT2
Length = 763
Score = 48.5 bits (114), Expect = 4e-05
Identities = 44/166 (26%), Positives = 72/166 (42%), Gaps = 14/166 (8%)
Query: 374 VCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVI 433
V +IPG +Y +DR+GR + + G MA++ + S +G+ M+
Sbjct: 404 VAFSIPG----MYLVDRIGRRPVLLAGGVIMAIANLVIAIVGVS---EGKTVVASKIMIA 456
Query: 434 YGLAFFFANFGPNT--TTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE 491
+ + F A F ++V AEL+P RS C I A ++ L +
Sbjct: 457 F-ICLFIAAFSATWGGVVWVVSAELYPLGVRSKCTAICAAANW---LVNFTCALITPYIV 512
Query: 492 KEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENED 537
+ +G K I GG+ +V + V YF ET G +LEE ++
Sbjct: 513 DVGSHTSSMGPKI-FFIWGGLNVVAVIVVYFAVYETRGLTLEEIDE 557
>GHT3_SCHPO (Q92339) High-affinity gluconate transporter ght3
(Hexose transporter 3)
Length = 555
Score = 48.1 bits (113), Expect = 6e-05
Identities = 41/162 (25%), Positives = 72/162 (44%), Gaps = 18/162 (11%)
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFF---ALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
F +Y ID +GR + G F ++ FF A+G + +H M+++ F
Sbjct: 316 FGALYTIDNLGRRNPLIFGAAFQSICFFIYAAVGDRKLIYKNGTSDHRAGSVMIVFSCLF 375
Query: 439 FFA---NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEG 495
F+ ++GP +++ E FP R+RS C ++ + +G + S + ++
Sbjct: 376 LFSYCCSWGP--MGWVIVGETFPIRYRSKCASVATSGNWLGNFMISFFTPFINN------ 427
Query: 496 YPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENED 537
IG K I + + F+ +F KET G +LEE D
Sbjct: 428 ---AIGFKLGYIY-ACINLFSSFMIFFLAKETKGLTLEEVND 465
>CSBC_BACSU (P46333) Probable metabolite transport protein csbC
Length = 461
Score = 47.0 bits (110), Expect = 1e-04
Identities = 41/153 (26%), Positives = 69/153 (44%), Gaps = 19/153 (12%)
Query: 388 IDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIY---GLAFFFANFG 444
IDRVGR K+ + G + +S AL T G + V++ + F+ A +G
Sbjct: 301 IDRVGRKKLLIWGSVGITLSLAALSGVLL---TLGLSASTAWMTVVFLGVYIVFYQATWG 357
Query: 445 PNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGF-LWASHKEKEEGYPKGIGMK 503
P +++ ELFP++ R G + V +I S+ F L S +G+
Sbjct: 358 P--VVWVLMPELFPSKARGAATGFTTLVLSAANLIVSLVFPLMLS----------AMGIA 405
Query: 504 ASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
++ +C++ F ++ ET G+SLEE E
Sbjct: 406 WVFMVFSVICLLSFFFAFYMVPETKGKSLEEIE 438
>ARAE_BACSU (P96710) Arabinose-proton symporter (Arabinose
transporter)
Length = 464
Score = 47.0 bits (110), Expect = 1e-04
Identities = 44/158 (27%), Positives = 67/158 (41%), Gaps = 26/158 (16%)
Query: 385 VYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF---FFA 441
V ID+VGR K+ +G FMA+ +G +Y T G M++ L F F
Sbjct: 322 VLLIDKVGRKKLMSIGSAFMAIFMILIGTSFYFELTSGI------MMIVLILGFVAAFCV 375
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGI- 500
+ GP T+I+ +E+FP R+ GI+ FLW ++ + P I
Sbjct: 376 SVGP--ITWIMISEIFPNHLRARAAGIATI------------FLWGANWAIGQFVPMMID 421
Query: 501 --GMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G+ + I + I+ ET +SLEE E
Sbjct: 422 SFGLAYTFWIFAVINILCFLFVVTICPETKNKSLEEIE 459
>SNF3_YEAST (P10870) High-affinity glucose transporter SNF3
Length = 818
Score = 46.2 bits (108), Expect = 2e-04
Identities = 42/164 (25%), Positives = 71/164 (42%), Gaps = 11/164 (6%)
Query: 374 VCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVI 433
V +PG FF +F GR K+ ++G M ++ F + S T F+ +
Sbjct: 336 VVFNVPGLFFVEFF----GRRKVLVVGGVIMTIANFIVAIVGCSLKTVAAAKVMIAFICL 391
Query: 434 YGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKE 493
+ +A F A +G +++ AEL+P RS C I A ++ + L +
Sbjct: 392 F-IAAFSATWGG--VVWVISAELYPLGVRSKCTAICAAANW---LVNFICALITPYIVDT 445
Query: 494 EGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENED 537
+ +G K I G + +G+ V Y ET G +LEE ++
Sbjct: 446 GSHTSSLGAKI-FFIWGSLNAMGVIVVYLTVYETKGLTLEEIDE 488
>GTR1_LEIDO (Q01440) Membrane transporter D1
Length = 547
Score = 46.2 bits (108), Expect = 2e-04
Identities = 51/274 (18%), Positives = 107/274 (38%), Gaps = 36/274 (13%)
Query: 269 LVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACSANW 328
L+ + AK + +V + + E LP PL++R+ +R + S
Sbjct: 192 LLSKGHADRAKAVADKFEVDLCEFQEGDELPSVRIDYRPLMARDMRFR----VVLSSGLQ 247
Query: 329 FLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFT---V 385
+ + +++ S + +Y F A + +L++ FT +
Sbjct: 248 IIQQFSGINTIMYYSSVI-----------LYDAGFRDAIMPVVLSIPLAFMNALFTAVAI 296
Query: 386 YFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGP 445
+ +DR GR ++ ++ F V + + T+ ++ G + + LA F A + P
Sbjct: 297 FTVDRFGRRRMLLISVFGCLVLLVVIAIIGFFIGTR-ISYSVGGGLFLALLAVFLALYAP 355
Query: 446 NT--TTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYP---KGI 500
+++ E+FP R++ ++ W ++ + +P I
Sbjct: 356 GIGCIPWVIMGEIFPTHLRTSAASVATMAN------------WGANVLVSQVFPILMGAI 403
Query: 501 GMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
G+ + I+ G+ +G YFF ET G +LE+
Sbjct: 404 GVGGTFTIISGLMALGCIFVYFFAVETKGLTLEQ 437
>YCEI_BACSU (O34691) Hypothetical metabolite transport protein yceI
Length = 400
Score = 45.4 bits (106), Expect = 4e-04
Identities = 32/121 (26%), Positives = 51/121 (41%), Gaps = 13/121 (10%)
Query: 371 ILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGF 430
+L + +PGYF + I++ GR I ++ A S + G D+
Sbjct: 257 LLMTLAQLPGYFSAAWLIEKAGRKWILVVYLIGTAGSAYFFG-----------TADSLSL 305
Query: 431 MVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGS--VGFLWAS 488
++ G+ F N G + E +P R+T G + A G++G I G VG L A
Sbjct: 306 LLTAGVLLSFFNLGAWGVLYAYTPEQYPTAIRATGSGTTAAFGRIGGIFGPLLVGTLAAR 365
Query: 489 H 489
H
Sbjct: 366 H 366
>YAAU_ECOLI (P31679) Hypothetical metabolite transport protein yaaU
Length = 443
Score = 45.1 bits (105), Expect = 5e-04
Identities = 33/153 (21%), Positives = 68/153 (43%), Gaps = 22/153 (14%)
Query: 387 FIDRVGRVKIQMMGFFFMAVSFFALGF-PYYSHWTKGENHDNKGFMVIYGLAFF-FANFG 444
+++ GR + + F M ++ LG P W +V+ A + F + G
Sbjct: 302 WLNTAGRRPLLIGSFAMMTLALAVLGLIPDMGIW-----------LVVMAFAVYAFFSGG 350
Query: 445 PNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKA 504
P ++ P ELFP R++ G+ ++ ++G I+ + WA + G+
Sbjct: 351 PGNLQWLYPNELFPTDIRASAVGVIMSLSRIGTIVST----WAL-----PIFINNYGISN 401
Query: 505 SLIILGGVCIVGMFVTYFFTKETMGRSLEENED 537
++++ G+ + G+ ++ F ET G SL + +
Sbjct: 402 TMLMGAGISLFGLLISVAFAPETRGMSLAQTSN 434
>GHT4_SCHPO (O59932) High-affinity hexose transporter ght4 (Hexose
transporter 4)
Length = 557
Score = 45.1 bits (105), Expect = 5e-04
Identities = 40/159 (25%), Positives = 69/159 (43%), Gaps = 18/159 (11%)
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFF---ALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
F +Y ID +GR + G F ++ FF A+G + +H M+++ F
Sbjct: 316 FGALYTIDNLGRRNPLIFGAAFQSICFFIYAAVGDRKLIYKNGTSDHRAGAVMIVFSCLF 375
Query: 439 FFA---NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEG 495
F+ ++GP +++ E FP R+RS C ++ + +G + S + S+
Sbjct: 376 LFSYCCSWGP--MGWVIVGETFPIRYRSKCAAVATSGNWLGNFMVSFFTPFISN------ 427
Query: 496 YPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
IG K I + + F + KET G +LEE
Sbjct: 428 ---SIGFKLG-YIYACINMTSAFQIFLMAKETKGLTLEE 462
>KHT2_KLULA (P53387) Hexose transporter 2
Length = 566
Score = 43.9 bits (102), Expect = 0.001
Identities = 48/190 (25%), Positives = 81/190 (42%), Gaps = 32/190 (16%)
Query: 361 EAFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMA---VSFFALGFPYYS 417
++F + + I+ ST F +Y +D+ GR K + G M V F ++G
Sbjct: 355 DSFETSIVLGIVNFAST----FVAIYVVDKFGRRKCLLWGAAAMTACMVVFASVGVTRL- 409
Query: 418 HWTKGENHD---NKGF---MVIYGLAFFFA---NFGPNTTTFIVPAELFPARFRSTCHGI 468
W G NH +KG M+++ + F ++ P ++V AE +P R ++ C I
Sbjct: 410 -WPDGANHPETASKGAGNCMIVFACFYIFCFATSWAP--IAYVVVAESYPLRVKAKCMAI 466
Query: 469 SGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFV-TYFFTKET 527
+ A + + GF I + +G C+V MF +FF ET
Sbjct: 467 ATASNWIWGFLN--GFFTPF-------ITSAIHFYYGYVFMG--CLVAMFFYVFFFVPET 515
Query: 528 MGRSLEENED 537
G +LEE ++
Sbjct: 516 KGLTLEEVQE 525
>HXT0_YEAST (P43581) Hexose transporter HXT10
Length = 546
Score = 43.9 bits (102), Expect = 0.001
Identities = 44/182 (24%), Positives = 76/182 (41%), Gaps = 22/182 (12%)
Query: 360 QEAFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSH- 418
Q++F + + + ST F +Y +D+ GR K + G MA+ F +
Sbjct: 341 QDSFETSIVLGAVNFAST----FVALYIVDKFGRRKCLLWGSASMAICFVIFATVGVTRL 396
Query: 419 WTKGENH---DNKGFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGIS-GAV 472
W +G++ + G ++I FF +F +++ AE +P R ++ I+ GA
Sbjct: 397 WPQGKDQPSSQSAGNVMIVFTCFFIFSFAITWAPIAYVIVAETYPLRVKNRAMAIAVGAN 456
Query: 473 GKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSL 532
G +IG + + G+ G + G I F +FF ET G +L
Sbjct: 457 WMWGFLIG----FFTPFITRSIGFSYG-------YVFMGCLIFSYFYVFFFVCETKGLTL 505
Query: 533 EE 534
EE
Sbjct: 506 EE 507
>HXTF_YEAST (P47185) Hexose transporter HXT16
Length = 567
Score = 43.1 bits (100), Expect = 0.002
Identities = 44/159 (27%), Positives = 67/159 (41%), Gaps = 18/159 (11%)
Query: 385 VYFIDRVGRVKIQMMGFFFMA---VSFFALGFP-YYSHWTKGENHDNKGFMVIYGLAFFF 440
V +D++GR K + G M V F ++G Y H G + G +I F+
Sbjct: 374 VMVVDKIGRRKCLLFGAASMMACMVIFASIGVKCLYPHGQDGPSSKGAGNAMIVFTCFYI 433
Query: 441 ANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPK 498
F +IV AE FP++ +S IS A + + +GF
Sbjct: 434 FCFATTWAPVAYIVVAESFPSKVKSKAMSISTAFNWLWQFL--IGFFTPF-------ITG 484
Query: 499 GIGMKASLIILGGVCIVGMFV-TYFFTKETMGRSLEENE 536
I + +G C+V MF+ +FF ET+G SLEE +
Sbjct: 485 SIHFYYGYVFVG--CLVAMFLYVFFFLPETIGLSLEETQ 521
>YOU1_CAEEL (P30638) Hypothetical protein ZK637.1 in chromosome III
Length = 520
Score = 42.7 bits (99), Expect = 0.002
Identities = 47/205 (22%), Positives = 78/205 (37%), Gaps = 23/205 (11%)
Query: 333 IVFYSQVLFQS--EIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYFIDR 390
+V ++ VLFQS E + + +V Q + + PG TV I+
Sbjct: 336 MVLFTTVLFQSHDECHGGLFSNGTQMEVCQPLTRSDYFDLLSTTLAEFPGLIITVLIIEW 395
Query: 391 VGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPNTTTF 450
GR K + + A+ F L F D V+ +A F + G +
Sbjct: 396 FGRKKTMALEYAVFAIFTFLLYFCL----------DRFTVTVLIFVARAFIS-GAFQCAY 444
Query: 451 IVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILG 510
+ E++P R+ G A+ ++GAI+ F+ EK P G I G
Sbjct: 445 VYTPEVYPTTLRAVGLGTCSAMARIGAIV--TPFIAQVASEKSLSLPIG--------IYG 494
Query: 511 GVCIVGMFVTYFFTKETMGRSLEEN 535
I+G+ + ET GR + ++
Sbjct: 495 TAAILGLIASLSLPIETKGRQMMDS 519
>GHT2_SCHPO (O74969) High-affinity glucose transporter ght2 (Hexose
transporter 2)
Length = 531
Score = 42.7 bits (99), Expect = 0.002
Identities = 40/160 (25%), Positives = 67/160 (41%), Gaps = 20/160 (12%)
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFF---ALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
F +Y ++R GR ++G + ++ FF A+G H G ++ G ++I
Sbjct: 320 FGGMYVLERFGRRNPLIIGGIWQSICFFIYSAVGSRALYH-KNGTSNTRAGAVMIVMACL 378
Query: 439 FFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGS--VGFLWASHKEKEE 494
F F +++ E +P R+RS C ++ A + + S F+ AS
Sbjct: 379 FIFGFAQTWAPAAYVIVGESYPVRYRSKCAAVATASNWLWNFLISFFTPFIQAS------ 432
Query: 495 GYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
IG K + + G V + F KET G +LEE
Sbjct: 433 -----IGFKYGYVF-ASCNLTGAIVIFLFAKETKGLTLEE 466
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 58,931,623
Number of Sequences: 164201
Number of extensions: 2336020
Number of successful extensions: 7431
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 7354
Number of HSP's gapped (non-prelim): 93
length of query: 548
length of database: 59,974,054
effective HSP length: 115
effective length of query: 433
effective length of database: 41,090,939
effective search space: 17792376587
effective search space used: 17792376587
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC149576.4


