

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.21 - phase: 0
(281 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
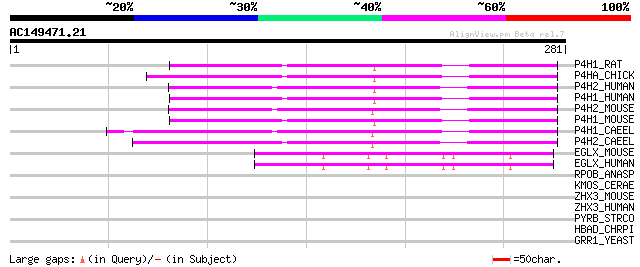
Score E
Sequences producing significant alignments: (bits) Value
P4H1_RAT (P54001) Prolyl 4-hydroxylase alpha-1 subunit precursor... 111 2e-24
P4HA_CHICK (P16924) Prolyl 4-hydroxylase alpha subunit (EC 1.14.... 110 5e-24
P4H2_HUMAN (O15460) Prolyl 4-hydroxylase alpha-2 subunit precurs... 105 2e-22
P4H1_HUMAN (P13674) Prolyl 4-hydroxylase alpha-1 subunit precurs... 105 2e-22
P4H2_MOUSE (Q60716) Prolyl 4-hydroxylase alpha-2 subunit precurs... 104 3e-22
P4H1_MOUSE (Q60715) Prolyl 4-hydroxylase alpha-1 subunit precurs... 104 3e-22
P4H1_CAEEL (Q10576) Prolyl 4-hydroxylase alpha-1 subunit precurs... 103 6e-22
P4H2_CAEEL (Q20065) Prolyl 4-hydroxylase alpha-2 subunit precurs... 102 1e-21
EGLX_MOUSE (Q8BG58) Putative HIF-prolyl hydroxylase PH-4 (EC 1.1... 60 6e-09
EGLX_HUMAN (Q9NXG6) Putative HIF-prolyl hydroxylase PH-4 (EC 1.1... 60 8e-09
RPOB_ANASP (P22703) DNA-directed RNA polymerase beta chain (EC 2... 37 0.041
KMOS_CERAE (P10650) Proto-oncogene serine/threonine-protein kina... 32 2.3
ZHX3_MOUSE (Q8C0Q2) Zinc fingers and homeoboxes protein 3 (Tripl... 30 5.0
ZHX3_HUMAN (Q9H4I2) Zinc fingers and homeoboxes protein 3 (Tripl... 30 6.6
PYRB_STRCO (Q9KXR2) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 30 8.6
HBAD_CHRPI (P02005) Hemoglobin alpha-D chain 30 8.6
GRR1_YEAST (P24814) Ubiquitin ligase complex F-box protein GRR1 30 8.6
>P4H1_RAT (P54001) Prolyl 4-hydroxylase alpha-1 subunit precursor
(EC 1.14.11.2) (4-PH alpha-1)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-1 subunit)
Length = 534
Score = 111 bits (278), Expect = 2e-24
Identities = 68/200 (34%), Positives = 95/200 (47%), Gaps = 19/200 (9%)
Query: 82 PRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKYP 141
PRII H+ +S E + ++ +A PRL +TV D TGK + R S +LS E P
Sbjct: 335 PRIIRFHDIISDAEIEIVKDLAKPRLSRATVHDPETGKLTTAQYRVSKSAWLSGYED--P 392
Query: 142 MIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFS----DTFNLKRGGQRIAT 197
++ I RI + + + E +QV Y Y PH D+ D F G RIAT
Sbjct: 393 VVSRINMRIQDLTGLDVSTAEELQVANYGVGGQYEPHFDFARKDEPDAFRELGTGNRIAT 452
Query: 198 MLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSV 257
L Y+ D GG T FP G+ V P KG AV ++++ G+ D +
Sbjct: 453 WLFYMSDVSAGGATVFPEVGAS-------------VWPKKGTAVFWYNLFASGEGDYSTR 499
Query: 258 HGGCPVLAGEKWSATKWMRQ 277
H CPVL G KW + KW+ +
Sbjct: 500 HAACPVLVGNKWVSNKWLHE 519
>P4HA_CHICK (P16924) Prolyl 4-hydroxylase alpha subunit (EC
1.14.11.2) (4-PH alpha)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha
subunit)
Length = 516
Score = 110 bits (274), Expect = 5e-24
Identities = 72/212 (33%), Positives = 100/212 (46%), Gaps = 19/212 (8%)
Query: 70 LGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSS 129
LG VK E PRI+ + +S EE + ++ +A PRL +TV D TGK + R S
Sbjct: 305 LGPVKQEDEWDKPRIVRFLDIISDEEIETVKELAKPRLSRATVHDPETGKLTTAHYRVSK 364
Query: 130 GMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFS----DT 185
+LS E P++ I RI + + + E +QV Y Y PH D+ D
Sbjct: 365 SAWLSGYES--PVVSRINTRIQDLTGLDVSTAEELQVANYGVGGQYEPHFDFGRKDEPDA 422
Query: 186 FNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWS 245
F G RIAT L Y+ D GG T FP G+ V P KG AV +++
Sbjct: 423 FKELGTGNRIATWLFYMSDVSAGGATVFPEVGAS-------------VWPKKGTAVFWYN 469
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQ 277
+ G+ D + H CPVL G KW + KW+ +
Sbjct: 470 LFPSGEGDYSTRHAACPVLVGNKWVSNKWLHE 501
>P4H2_HUMAN (O15460) Prolyl 4-hydroxylase alpha-2 subunit precursor
(EC 1.14.11.2) (4-PH alpha-2)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-2 subunit) (UNQ290/PRO330)
Length = 535
Score = 105 bits (261), Expect = 2e-22
Identities = 63/201 (31%), Positives = 98/201 (48%), Gaps = 19/201 (9%)
Query: 81 SPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKY 140
SP I+ ++ +S EE + ++ +A P+L +TV D TG + R S +L EE
Sbjct: 335 SPHIVRYYDVMSDEEIERIKEIAKPKLARATVRDPKTGVLTVASYRVSKSSWL--EEDDD 392
Query: 141 PMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFS----DTFNLKRGGQRIA 196
P++ + +R+ + + ++ EL+QV Y Y PH D+ DTF G R+A
Sbjct: 393 PVVARVNRRMQHITGLTVKTAELLQVANYGVGGQYEPHFDFSRNDERDTFKHLGTGNRVA 452
Query: 197 TMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDS 256
T L Y+ D GG T FP G+ + P KG AV ++++ G+ D +
Sbjct: 453 TFLNYMSDVEAGGATVFPDLGA-------------AIWPKKGTAVFWYNLLRSGEGDYRT 499
Query: 257 VHGGCPVLAGEKWSATKWMRQ 277
H CPVL G KW + KW +
Sbjct: 500 RHAACPVLVGCKWVSNKWFHE 520
>P4H1_HUMAN (P13674) Prolyl 4-hydroxylase alpha-1 subunit precursor
(EC 1.14.11.2) (4-PH alpha-1)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-1 subunit)
Length = 534
Score = 105 bits (261), Expect = 2e-22
Identities = 65/200 (32%), Positives = 94/200 (46%), Gaps = 19/200 (9%)
Query: 82 PRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKYP 141
PRII H+ +S E + ++ +A PRL+ +T+ + TG R S +LS E P
Sbjct: 335 PRIIRFHDIISDAEIEIVKDLAKPRLRRATISNPITGDLETVHYRISKSAWLSGYEN--P 392
Query: 142 MIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFS----DTFNLKRGGQRIAT 197
++ I RI + + + E +QV Y Y PH D+ D F G RIAT
Sbjct: 393 VVSRINMRIQDLTGLDVSTAEELQVANYGVGGQYEPHFDFARKDEPDAFKELGTGNRIAT 452
Query: 198 MLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSV 257
L Y+ D GG T FP G+ V P KG AV ++++ G+ D +
Sbjct: 453 WLFYMSDVSAGGATVFPEVGAS-------------VWPKKGTAVFWYNLFASGEGDYSTR 499
Query: 258 HGGCPVLAGEKWSATKWMRQ 277
H CPVL G KW + KW+ +
Sbjct: 500 HAACPVLVGNKWVSNKWLHE 519
>P4H2_MOUSE (Q60716) Prolyl 4-hydroxylase alpha-2 subunit precursor
(EC 1.14.11.2) (4-PH alpha-2)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-2 subunit)
Length = 537
Score = 104 bits (259), Expect = 3e-22
Identities = 62/201 (30%), Positives = 97/201 (47%), Gaps = 19/201 (9%)
Query: 81 SPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKY 140
SP I+ ++ +S EE + ++ +A P+L +TV D TG + R S +L EE
Sbjct: 337 SPHIVRYYDVMSDEEIERIKEIAKPKLARATVRDPKTGVLTVASYRVSKSSWL--EEDDD 394
Query: 141 PMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYF----SDTFNLKRGGQRIA 196
P++ + +R+ + + ++ EL+QV Y Y PH D+ D F G R+A
Sbjct: 395 PVVARVNRRMQHITGLTVKTAELLQVANYGMGGQYEPHFDFSRSDDEDAFKRLGTGNRVA 454
Query: 197 TMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDS 256
T L Y+ D GG T FP G+ + P KG AV ++++ G+ D +
Sbjct: 455 TFLNYMSDVEAGGATVFPDLGA-------------AIWPKKGTAVFWYNLLRSGEGDYRT 501
Query: 257 VHGGCPVLAGEKWSATKWMRQ 277
H CPVL G KW + KW +
Sbjct: 502 RHAACPVLVGCKWVSNKWFHE 522
>P4H1_MOUSE (Q60715) Prolyl 4-hydroxylase alpha-1 subunit precursor
(EC 1.14.11.2) (4-PH alpha-1)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-1 subunit)
Length = 534
Score = 104 bits (259), Expect = 3e-22
Identities = 65/200 (32%), Positives = 94/200 (46%), Gaps = 19/200 (9%)
Query: 82 PRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKYP 141
PRII H+ +S E + ++ +A PRL+ +T+ + TG R S +LS E P
Sbjct: 335 PRIIRFHDIISDAEIEIVKDLAKPRLRRATISNPVTGALETVHYRISKSAWLSGYED--P 392
Query: 142 MIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFS----DTFNLKRGGQRIAT 197
++ I RI + + + E +QV Y Y PH D+ D F G RIAT
Sbjct: 393 VVSRINMRIQDLTGLDVSTAEELQVANYGVGGQYEPHFDFARKDEPDAFRELGTGNRIAT 452
Query: 198 MLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSV 257
L Y+ D GG T FP G+ V P KG AV ++++ G+ D +
Sbjct: 453 WLFYMSDVSAGGATVFPEVGAS-------------VWPKKGTAVFWYNLFASGEGDYSTR 499
Query: 258 HGGCPVLAGEKWSATKWMRQ 277
H CPVL G KW + KW+ +
Sbjct: 500 HAACPVLVGNKWVSNKWLHE 519
>P4H1_CAEEL (Q10576) Prolyl 4-hydroxylase alpha-1 subunit precursor
(EC 1.14.11.2) (4-PH alpha-1)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-1 subunit)
Length = 559
Score = 103 bits (256), Expect = 6e-22
Identities = 65/232 (28%), Positives = 113/232 (48%), Gaps = 23/232 (9%)
Query: 50 LPRGYTYWNNNNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKI 109
+ R Y Y+ ++ L +K E+ ++P +L + +S +E ++ +A P+L
Sbjct: 300 ISRLYCYYK----RDRPFLVYAPIKVEIKRFNPLAVLFKDVISDDEVAAIQELAKPKLAR 355
Query: 110 STVVDANTGKGIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRY 169
+TV D+ TGK + + R S +L +E + ++ + KRI + + +E E +Q+ Y
Sbjct: 356 ATVHDSVTGKLVTATYRISKSAWL--KEWEGDVVETVNKRIGYMTNLEMETAEELQIANY 413
Query: 170 EKNQYYRPHHDYF----SDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGG 225
+Y PH D+ S +F G RIAT+L Y+ GG T F A S
Sbjct: 414 GIGGHYDPHFDHAKKEESKSFESLGTGNRIATVLFYMSQPSHGGGTVFTEAKST------ 467
Query: 226 KLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQ 277
+ P K +A+ ++++ G +PD+ H CPVL G KW + KW+ +
Sbjct: 468 -------ILPTKNDALFWYNLYKQGDGNPDTRHAACPVLVGIKWVSNKWIHE 512
>P4H2_CAEEL (Q20065) Prolyl 4-hydroxylase alpha-2 subunit precursor
(EC 1.14.11.2) (4-PH alpha-2)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-2 subunit)
Length = 539
Score = 102 bits (254), Expect = 1e-21
Identities = 62/219 (28%), Positives = 106/219 (48%), Gaps = 19/219 (8%)
Query: 63 KEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIK 122
++ L+L +K E+L + P +L N + E + ++ +A P+LK +TV ++ TG+
Sbjct: 306 RDKPFLKLAPIKVEILRFDPLAVLFKNVIHDSEIEVIKELASPKLKRATVQNSKTGELEH 365
Query: 123 SDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYF 182
+ R S +L + P+I + +RI ++ + E +QV Y +Y PH D+
Sbjct: 366 ATYRISKSAWLKGDLD--PVIDRVNRRIEDFTNLNQATSEELQVANYGLGGHYDPHFDFA 423
Query: 183 ----SDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKG 238
+ F G RIAT+L Y+ GG T F G+ V P K
Sbjct: 424 RKEEKNAFKTLNTGNRIATVLFYMSQPERGGATVFNHLGT-------------AVFPSKN 470
Query: 239 NAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQ 277
+A+ ++++ DG+ D + H CPVL G KW + KW+ +
Sbjct: 471 DALFWYNLRRDGEGDLRTRHAACPVLLGVKWVSNKWIHE 509
>EGLX_MOUSE (Q8BG58) Putative HIF-prolyl hydroxylase PH-4 (EC
1.14.11.-) (Hypoxia-inducible factor prolyl
4-hydroxylase)
Length = 503
Score = 60.1 bits (144), Expect = 6e-09
Identities = 57/187 (30%), Positives = 82/187 (43%), Gaps = 36/187 (19%)
Query: 125 VRTSSGMFLSHEERKYPMIHAIEKRISVYSQIP---IENGELMQVLRYEKNQYYRPHHD- 180
VR S +L E + ++ AI +R+ +++ +E E +QV+RY + +Y H D
Sbjct: 273 VRNSHHTWLHQGEGAHHVMRAIRQRVLRLTRLSPEIVEFSEPLQVVRYGEGGHYHAHVDS 332
Query: 181 --YFSDTFNLK-----------RGGQRIATMLMYLGDNVEGGETHFPSAGS---DECSC- 223
+ +T R T+L YL + GGET FP A + DE S
Sbjct: 333 GPVYPETICSHTKLVANESVPFETSCRYMTVLFYLNNVTGGGETVFPVADNRTYDEMSLI 392
Query: 224 ---------GGKLTKG-LCVKPVKGNAVLFWSMGLDGQS-----DPDSVHGGCPVLAGEK 268
KG L VKP +G AV +++ DGQ D S+HGGC V G K
Sbjct: 393 QDDVDLRDTRRHCDKGNLRVKPQQGTAVFWYNYLPDGQGWVGEVDDYSLHGGCLVTRGTK 452
Query: 269 WSATKWM 275
W A W+
Sbjct: 453 WIANNWI 459
>EGLX_HUMAN (Q9NXG6) Putative HIF-prolyl hydroxylase PH-4 (EC
1.14.11.-) (Hypoxia-inducible factor prolyl
4-hydroxylase)
Length = 502
Score = 59.7 bits (143), Expect = 8e-09
Identities = 57/187 (30%), Positives = 82/187 (43%), Gaps = 36/187 (19%)
Query: 125 VRTSSGMFLSHEERKYPMIHAIEKRISVYSQIP---IENGELMQVLRYEKNQYYRPHHD- 180
VR S +L E + ++ AI +R+ +++ +E E +QV+RY + +Y H D
Sbjct: 272 VRNSHHTWLYQGEGAHHIMRAIRQRVLRLTRLSPEIVELSEPLQVVRYGEGGHYHAHVDS 331
Query: 181 --YFSDTFNLK-----------RGGQRIATMLMYLGDNVEGGETHFPSAGS---DECSC- 223
+ +T R T+L YL + GGET FP A + DE S
Sbjct: 332 GPVYPETICSHTKLVANESVPFETSCRYMTVLFYLNNVTGGGETVFPVADNRTYDEMSLI 391
Query: 224 ---------GGKLTKG-LCVKPVKGNAVLFWSMGLDGQS-----DPDSVHGGCPVLAGEK 268
KG L VKP +G AV +++ DGQ D S+HGGC V G K
Sbjct: 392 QDDVDLRDTRRHCDKGNLRVKPQQGTAVFWYNYLPDGQGWVGDVDDYSLHGGCLVTRGTK 451
Query: 269 WSATKWM 275
W A W+
Sbjct: 452 WIANNWI 458
>RPOB_ANASP (P22703) DNA-directed RNA polymerase beta chain (EC
2.7.7.6) (RNAP beta subunit) (Transcriptase beta chain)
(RNA polymerase beta subunit)
Length = 1131
Score = 37.4 bits (85), Expect = 0.041
Identities = 31/105 (29%), Positives = 51/105 (48%), Gaps = 12/105 (11%)
Query: 118 GKGIKSDVRTSSGMFL-SHEERKYPMIHAIEKRISVYSQIPI------ENGELM----QV 166
G G+++ SGM + S + + A E R+ V Q+P +NG+L Q
Sbjct: 575 GTGLEAQGARDSGMVIVSRTDGDVVYVDATEIRVRVSGQLPTASGKSTDNGQLTSQKGQE 634
Query: 167 LRYEKNQYYRPHHDYFSDTFNLKRGGQR-IATMLMYLGDNVEGGE 210
+RY ++Y R + D + L R G+R +A ++ G + EGGE
Sbjct: 635 IRYTVSKYQRSNQDTCLNQKPLVRIGERVVAGQVLADGSSTEGGE 679
>KMOS_CERAE (P10650) Proto-oncogene serine/threonine-protein kinase
mos (EC 2.7.1.37) (c-mos) (Oocyte maturation factor mos)
Length = 346
Score = 31.6 bits (70), Expect = 2.3
Identities = 19/51 (37%), Positives = 24/51 (46%), Gaps = 7/51 (13%)
Query: 191 GGQRIATMLMYLGDNVE------GGETHFPS-AGSDECSCGGKLTKGLCVK 234
G + T++M G NV G +H AG CS GG LT G C+K
Sbjct: 130 GSNSLGTIIMEFGGNVTLHQVIYGAASHPEGDAGEPHCSTGGPLTLGKCLK 180
>ZHX3_MOUSE (Q8C0Q2) Zinc fingers and homeoboxes protein 3 (Triple
homeobox 1 protein)
Length = 951
Score = 30.4 bits (67), Expect = 5.0
Identities = 23/74 (31%), Positives = 36/74 (48%), Gaps = 5/74 (6%)
Query: 40 FTTTTTTRSLLPRGY--TYWNNNNDKEAQILRLGYVKPEVLSW--SPRIILLHNFLSYEE 95
+ T R LL + + T W +N D ++ + + G +PEV+ W R L + L + E
Sbjct: 764 YKKTAQQRHLLRQLFVQTQWPSNQDYDSIMAQTGLPRPEVVRWFGDSRYALKNGQLKWYE 823
Query: 96 CDYLRGVALPRLKI 109
DY RG P L +
Sbjct: 824 -DYKRGNFPPGLLV 836
>ZHX3_HUMAN (Q9H4I2) Zinc fingers and homeoboxes protein 3 (Triple
homeobox 1 protein)
Length = 956
Score = 30.0 bits (66), Expect = 6.6
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Query: 55 TYWNNNNDKEAQILRLGYVKPEVLSW--SPRIILLHNFLSYEECDYLRGVALPRLKI 109
T W +N D ++ + + G +PEV+ W R L + L + E DY RG P L +
Sbjct: 786 TQWPSNQDYDSIMAQTGLPRPEVVRWFGDSRYALKNGQLKWYE-DYKRGNFPPGLLV 841
>PYRB_STRCO (Q9KXR2) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 326
Score = 29.6 bits (65), Expect = 8.6
Identities = 26/90 (28%), Positives = 42/90 (45%), Gaps = 11/90 (12%)
Query: 163 LMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATM-----LMYLGDNVEGGETHFPSAG 217
L++V R N + P +S + L G R+A M +M+ G V G E A
Sbjct: 228 LLRVQRERMNAAFFPTEREYSRRYGLD--GDRMAKMPDHAIVMHPGPMVRGMEITAEVAD 285
Query: 218 SDECSCGGKLTKGLCVKPVKGNAVLFWSMG 247
SD C+ ++T G+ ++ AVL+ +G
Sbjct: 286 SDRCTVVEQVTNGVSIR----MAVLYLLLG 311
>HBAD_CHRPI (P02005) Hemoglobin alpha-D chain
Length = 141
Score = 29.6 bits (65), Expect = 8.6
Identities = 19/67 (28%), Positives = 34/67 (50%), Gaps = 2/67 (2%)
Query: 133 LSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGG 192
L+H+E++ + HA EK + E E M + Y + + Y PH D D+ ++ G
Sbjct: 2 LNHDEKQL-IKHAWEKVLGHQEDFGAEALERMFAV-YPQTKTYFPHFDLHHDSEQIRHHG 59
Query: 193 QRIATML 199
+++ T L
Sbjct: 60 KKVVTAL 66
>GRR1_YEAST (P24814) Ubiquitin ligase complex F-box protein GRR1
Length = 1151
Score = 29.6 bits (65), Expect = 8.6
Identities = 18/59 (30%), Positives = 28/59 (46%), Gaps = 1/59 (1%)
Query: 8 IVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLL-PRGYTYWNNNNDKEA 65
+++ LL M++ + RL L++ T T R LL + WNNNND +A
Sbjct: 994 LIWRLLFIDKFIMVVQKYKLSTVVLRLYLKDNITLLTRQRELLIAHQRSAWNNNNDNDA 1052
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,630,727
Number of Sequences: 164201
Number of extensions: 1572609
Number of successful extensions: 3044
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3004
Number of HSP's gapped (non-prelim): 17
length of query: 281
length of database: 59,974,054
effective HSP length: 109
effective length of query: 172
effective length of database: 42,076,145
effective search space: 7237096940
effective search space used: 7237096940
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC149471.21


