

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.20 + phase: 0
(227 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
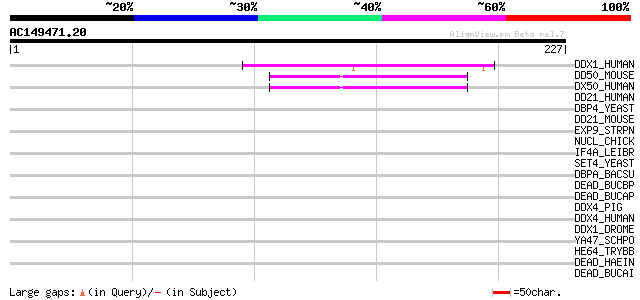
Score E
Sequences producing significant alignments: (bits) Value
DDX1_HUMAN (Q92499) ATP-dependent helicase DDX1 (DEAD-box protei... 51 3e-06
DD50_MOUSE (Q99MJ9) DEAD-box protein 50 (EC 3.6.1.-) (Nucleolar ... 49 1e-05
DX50_HUMAN (Q9BQ39) DEAD-box protein 50 (EC 3.6.1.-) (Nucleolar ... 48 2e-05
DD21_HUMAN (Q9NR30) Nucleolar RNA helicase II (Nucleolar RNA hel... 40 0.003
DBP4_YEAST (P20448) Probable ATP-dependent RNA helicase DBP4 (He... 40 0.006
DD21_MOUSE (Q9JIK5) Nucleolar RNA helicase II (Nucleolar RNA hel... 39 0.008
EXP9_STRPN (P35599) Probable RNA helicase exp9 (Exported protein 9) 37 0.050
NUCL_CHICK (P15771) Nucleolin (Protein C23) 36 0.086
IF4A_LEIBR (Q25225) Probable eukaryotic initiation factor 4A (eI... 35 0.11
SET4_YEAST (P42948) SET domain protein 4 35 0.19
DBPA_BACSU (P42305) ATP-dependent RNA helicase dbpA 34 0.25
DEAD_BUCBP (Q89AF9) Cold-shock DEAD-box protein A homolog (ATP-d... 34 0.33
DEAD_BUCAP (Q8K9H6) Cold-shock DEAD-box protein A homolog (ATP-d... 34 0.33
DDX4_PIG (Q6GWX0) DEAD-box protein 4 (VASA homolog) (VASA-like p... 33 0.42
DDX4_HUMAN (Q9NQI0) DEAD-box protein 4 (VASA homolog) 33 0.42
DDX1_DROME (Q9VNV3) ATP-dependent helicase DDX1 (DEAD box protei... 33 0.42
YA47_SCHPO (Q09719) Putative ATP-dependent RNA helicase C31A2.07c 33 0.55
HE64_TRYBB (Q26696) Putative DEAD-box RNA helicase HEL64 33 0.55
DEAD_HAEIN (P44586) Cold-shock DEAD-box protein A homolog (ATP-d... 33 0.55
DEAD_BUCAI (P57453) Cold-shock DEAD-box protein A homolog (ATP-d... 32 0.95
>DDX1_HUMAN (Q92499) ATP-dependent helicase DDX1 (DEAD-box protein
1) (DEAD-box protein-retinoblastoma) (DBP-RB)
Length = 740
Score = 50.8 bits (120), Expect = 3e-06
Identities = 35/109 (32%), Positives = 55/109 (50%), Gaps = 6/109 (5%)
Query: 96 SEVVEGKIPPGSPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKL----FSKHIKLQEQIS 151
++V + K P +P LIV PS + L + K KL + ++Q+S
Sbjct: 275 AQVTQTKFLPNAPKALIVEPSRELAEQTLNNIKQFKKYIDNPKLRELLIIGGVAARDQLS 334
Query: 152 LLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLDMHPDV--KGYSLF 198
+L+N V+I GTP R+ L+ L LS+++ LVLD + +GYS F
Sbjct: 335 VLENGVDIVVGTPGRLDDLVSTGKLNLSQVRFLVLDEADGLLSQGYSDF 383
>DD50_MOUSE (Q99MJ9) DEAD-box protein 50 (EC 3.6.1.-) (Nucleolar
protein Gu2) (Gu-beta)
Length = 734
Score = 48.5 bits (114), Expect = 1e-05
Identities = 29/81 (35%), Positives = 48/81 (58%), Gaps = 1/81 (1%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
SP VL+++P+ + + K F+ +T++ S V F Q QI+ ++N ++I GTP R
Sbjct: 208 SPKVLVLAPTRELANQVAKDFKDITRKLS-VACFYGGTSYQSQINQIRNGIDILVGTPGR 266
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
IK + L LS+L+ +VLD
Sbjct: 267 IKDHLQSGRLDLSKLRHVVLD 287
>DX50_HUMAN (Q9BQ39) DEAD-box protein 50 (EC 3.6.1.-) (Nucleolar
protein Gu2) (Gu-beta)
Length = 737
Score = 48.1 bits (113), Expect = 2e-05
Identities = 29/81 (35%), Positives = 48/81 (58%), Gaps = 1/81 (1%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
SP VL+++P+ + + K F+ +T++ S V F Q QI+ ++N ++I GTP R
Sbjct: 211 SPKVLVLAPTRELANQVAKDFKDITRKLS-VACFYGGTSYQSQINHIRNGIDILVGTPGR 269
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
IK + L LS+L+ +VLD
Sbjct: 270 IKDHLQSGRLDLSKLRHVVLD 290
>DD21_HUMAN (Q9NR30) Nucleolar RNA helicase II (Nucleolar RNA
helicase Gu) (RH II/Gu) (DEAD-box protein 21)
Length = 783
Score = 40.4 bits (93), Expect = 0.003
Identities = 27/81 (33%), Positives = 44/81 (53%), Gaps = 1/81 (1%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
+P VL+++P+ + + K F +TK+ S V F Q ++N ++I GTP R
Sbjct: 260 APQVLVLAPTRELANQVSKDFSDITKKLS-VACFYGGTPYGGQFERMRNGIDILVGTPGR 318
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
IK I L L++L+ +VLD
Sbjct: 319 IKDHIQNGKLDLTKLKHVVLD 339
>DBP4_YEAST (P20448) Probable ATP-dependent RNA helicase DBP4
(Helicase CA4) (Helicase UF1)
Length = 770
Score = 39.7 bits (91), Expect = 0.006
Identities = 56/203 (27%), Positives = 94/203 (45%), Gaps = 30/203 (14%)
Query: 4 RKGRENNDDRRKKKKKKTETTGSEQLRFFVNEFQSA-------NDVQLSSIELESLKADS 56
+K R N R+ ++K+ E E L+ ++E+ D+ +S L+ L+ +S
Sbjct: 3 KKNRLNTTQRKTLRQKEDEYI--ENLKTKIDEYDPKITKAKFFKDLPISDPTLKGLR-ES 59
Query: 57 SILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPP------GSPSV 110
S ++L+ Q + V L G D+ A + L V K+
Sbjct: 60 SFIKLTEIQAD---SIPVSLQGHDVLAAAKTGSGKTLAFLVPVIEKLYREKWTEFDGLGA 116
Query: 111 LIVSPS---ALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQ-EQISLLKNRVNIASGTPSR 166
LI+SP+ A++ +L T + + + K +K + E+IS R+NI GTP R
Sbjct: 117 LIISPTRELAMQIYEVLTKIGSHTSFSAGLVIGGKDVKFELERIS----RINILIGTPGR 172
Query: 167 IKKLIDVEALGL--SRLQVLVLD 187
I + +D +A+GL S LQ+LVLD
Sbjct: 173 ILQHLD-QAVGLNTSNLQMLVLD 194
>DD21_MOUSE (Q9JIK5) Nucleolar RNA helicase II (Nucleolar RNA
helicase Gu) (RH II/Gu) (DEAD-box protein 21)
Length = 851
Score = 39.3 bits (90), Expect = 0.008
Identities = 26/81 (32%), Positives = 45/81 (55%), Gaps = 1/81 (1%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
+P VL+++P+ + + K F +TK+ S V F QI +++ ++I GTP R
Sbjct: 332 APQVLVLAPTRELANQVSKDFSDITKKLS-VACFYGGTPYGGQIERMRSGIDILVGTPGR 390
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
IK + L L++L+ +VLD
Sbjct: 391 IKDHLQNGKLDLTKLKHVVLD 411
>EXP9_STRPN (P35599) Probable RNA helicase exp9 (Exported protein 9)
Length = 524
Score = 36.6 bits (83), Expect = 0.050
Identities = 21/61 (34%), Positives = 33/61 (53%)
Query: 127 FRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVL 186
FRF + V+ +++QI LK+ +I GTP R+ LI +AL L ++ L+L
Sbjct: 90 FRFGRSKGVKVRSVYGGSSIEKQIKALKSGAHIVVGTPGRLLDLIKRKALKLQDIETLIL 149
Query: 187 D 187
D
Sbjct: 150 D 150
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 35.8 bits (81), Expect = 0.086
Identities = 30/97 (30%), Positives = 43/97 (43%), Gaps = 16/97 (16%)
Query: 2 GKRKGRENNDDRRKKKKKKTETTGS---------------EQLRFFVNEFQSANDVQLSS 46
GKRK N + KKKKTET S E+LR + EF ++Q+S
Sbjct: 255 GKRKKEMANKSAPEAKKKKTETPASAFSLFVKNLTPTKDYEELRTAIKEFFGKKNLQVSE 314
Query: 47 IELESLKADSSILELSNSQNSRDTDLD-VKLLGGDIK 82
+ + S K + LS + L+ KL+G +IK
Sbjct: 315 VRIGSSKRFGYVDFLSAEDMDKALQLNGKKLMGLEIK 351
>IF4A_LEIBR (Q25225) Probable eukaryotic initiation factor 4A
(eIF4A) (eIF-4A)
Length = 403
Score = 35.4 bits (80), Expect = 0.11
Identities = 24/81 (29%), Positives = 43/81 (52%), Gaps = 4/81 (4%)
Query: 111 LIVSPS---ALRSIHLLKGF-RFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
L++SP+ AL++ ++ F++ + F ++Q+ + L+ V +A GTP R
Sbjct: 98 LVLSPTRELALQTAEVISRIGEFLSNSAKFCETFVGGTRVQDDLRKLQAGVVVAVGTPGR 157
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
+ +I AL L+VLVLD
Sbjct: 158 VSDVIKRGALRTESLRVLVLD 178
>SET4_YEAST (P42948) SET domain protein 4
Length = 560
Score = 34.7 bits (78), Expect = 0.19
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 11/79 (13%)
Query: 8 ENNDDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLK-----------ADS 56
EN++ RKK +++ S L+ +NE S+ND + +I + K D
Sbjct: 273 ENSNSIRKKLRQEKLVVSSHFLKPLLNEVSSSNDTEFKAITISEYKDKYVKMFIDNHYDD 332
Query: 57 SILELSNSQNSRDTDLDVK 75
+ SN ++SR D++V+
Sbjct: 333 DWVVCSNWESSRSADIEVR 351
>DBPA_BACSU (P42305) ATP-dependent RNA helicase dbpA
Length = 479
Score = 34.3 bits (77), Expect = 0.25
Identities = 26/84 (30%), Positives = 42/84 (49%), Gaps = 7/84 (8%)
Query: 108 PSVLIVSPSALRSIHLLKGF----RFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGT 163
P LI++P+ ++ + + RF K+ A +F K +Q + LK + +I GT
Sbjct: 71 PQALILTPTRELAVQVKEDITNIGRF--KRIKATAVFGKS-SFDKQKAELKQKSHIVVGT 127
Query: 164 PSRIKKLIDVEALGLSRLQVLVLD 187
P R+ I+ L L RL LV+D
Sbjct: 128 PGRVLDHIEKGTLPLDRLSYLVID 151
>DEAD_BUCBP (Q89AF9) Cold-shock DEAD-box protein A homolog
(ATP-dependent RNA helicase deaD homolog)
Length = 602
Score = 33.9 bits (76), Expect = 0.33
Identities = 21/82 (25%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Query: 108 PSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKH--IKLQEQISLLKNRVNIASGTPS 165
P +L+++P+ ++ + + F +K+ V + + + + Q+ L+ I GTP
Sbjct: 75 PQILVLTPTRELAVQVAEAFSNFSKKLIGVHVLALYGGQRYDLQLKSLRKGPQIIVGTPG 134
Query: 166 RIKKLIDVEALGLSRLQVLVLD 187
R+ + L LS L LVLD
Sbjct: 135 RLLDHLKRRTLSLSNLHSLVLD 156
>DEAD_BUCAP (Q8K9H6) Cold-shock DEAD-box protein A homolog
(ATP-dependent RNA helicase deaD homolog)
Length = 601
Score = 33.9 bits (76), Expect = 0.33
Identities = 20/83 (24%), Positives = 40/83 (48%), Gaps = 2/83 (2%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKH--IKLQEQISLLKNRVNIASGTP 164
+P +L+++P+ ++ + + F +K + + + + + Q+ L+ I GTP
Sbjct: 74 APQILVLAPTRELAVQVAEAFSDFSKYIMGIHVLPLYGGQRYEVQLRALRQGPQIVVGTP 133
Query: 165 SRIKKLIDVEALGLSRLQVLVLD 187
R+ + L LS L LVLD
Sbjct: 134 GRLLDHLKRGTLNLSNLYALVLD 156
>DDX4_PIG (Q6GWX0) DEAD-box protein 4 (VASA homolog) (VASA-like
protein)
Length = 722
Score = 33.5 bits (75), Expect = 0.42
Identities = 23/81 (28%), Positives = 38/81 (46%), Gaps = 1/81 (1%)
Query: 108 PSVLIVSPSA-LRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
P +IV+P+ L + L+ +F C + +L I + NI TP R
Sbjct: 364 PECIIVAPTRELVNQIYLEARKFSFGTCVRAVVIYGGTQLGHSIRQIVQGCNILCATPGR 423
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
+ +I E +GL +++ LVLD
Sbjct: 424 LMDIIGKEKIGLKQIKYLVLD 444
>DDX4_HUMAN (Q9NQI0) DEAD-box protein 4 (VASA homolog)
Length = 724
Score = 33.5 bits (75), Expect = 0.42
Identities = 23/81 (28%), Positives = 38/81 (46%), Gaps = 1/81 (1%)
Query: 108 PSVLIVSPSA-LRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
P +IV+P+ L + L+ +F C + +L I + NI TP R
Sbjct: 366 PECIIVAPTRELVNQIYLEARKFSFGTCVRAVVIYGGTQLGHSIRQIVQGCNILCATPGR 425
Query: 167 IKKLIDVEALGLSRLQVLVLD 187
+ +I E +GL +++ LVLD
Sbjct: 426 LMDIIGKEKIGLKQIKYLVLD 446
>DDX1_DROME (Q9VNV3) ATP-dependent helicase DDX1 (DEAD box protein
1)
Length = 727
Score = 33.5 bits (75), Expect = 0.42
Identities = 23/90 (25%), Positives = 46/90 (50%), Gaps = 4/90 (4%)
Query: 102 KIPPGSPSVLIVSPS---ALRSIHLLKGFRFMTKQCSAVKLFS-KHIKLQEQISLLKNRV 157
K P +P +I+ PS A ++ + ++ F++ L ++L+EQ + L
Sbjct: 281 KPAPNAPQAIIMEPSRELAEQTYNQIEKFKYHLSNPEVRSLLLIGGVRLEEQKAQLMQGT 340
Query: 158 NIASGTPSRIKKLIDVEALGLSRLQVLVLD 187
+I GTP R++++I+ + L+ + VLD
Sbjct: 341 HIVVGTPGRLEEMINSGLVLLTHCRFFVLD 370
>YA47_SCHPO (Q09719) Putative ATP-dependent RNA helicase C31A2.07c
Length = 848
Score = 33.1 bits (74), Expect = 0.55
Identities = 27/94 (28%), Positives = 46/94 (48%), Gaps = 5/94 (5%)
Query: 97 EVVEGKIPPGSPSVLIVSPS---ALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLL 153
E ++ + + LI+SP+ AL+++ ++K F T S + + L+EQ SLL
Sbjct: 129 EHLKSTLANSNTRALILSPNRELALQTVKVVKDFSKGTDLRSVAIVGG--VSLEEQFSLL 186
Query: 154 KNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLD 187
+ +I TP R L L LS ++ +V D
Sbjct: 187 SGKPDIVVATPGRFLHLKVEMKLELSSIEYVVFD 220
>HE64_TRYBB (Q26696) Putative DEAD-box RNA helicase HEL64
Length = 568
Score = 33.1 bits (74), Expect = 0.55
Identities = 15/39 (38%), Positives = 24/39 (61%)
Query: 149 QISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLD 187
Q+ LL+ V+I TP R+ +D++ + L R+ LVLD
Sbjct: 217 QLGLLRRGVHILVATPGRLIDFLDIKRINLHRVTYLVLD 255
>DEAD_HAEIN (P44586) Cold-shock DEAD-box protein A homolog
(ATP-dependent RNA helicase deaD homolog)
Length = 613
Score = 33.1 bits (74), Expect = 0.55
Identities = 21/82 (25%), Positives = 37/82 (44%), Gaps = 2/82 (2%)
Query: 108 PSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKH--IKLQEQISLLKNRVNIASGTPS 165
P +L+++P+ +I + K ++ + + + Q+ LK + GTP
Sbjct: 74 PQMLVMAPTRELAIQVADACELFVKYAQGTRIVTLYGGQRYDIQLRALKQGAQVVVGTPG 133
Query: 166 RIKKLIDVEALGLSRLQVLVLD 187
RI I L LS L+ +VLD
Sbjct: 134 RILDHIRRGTLNLSELRFIVLD 155
>DEAD_BUCAI (P57453) Cold-shock DEAD-box protein A homolog
(ATP-dependent RNA helicase deaD homolog)
Length = 601
Score = 32.3 bits (72), Expect = 0.95
Identities = 20/83 (24%), Positives = 40/83 (48%), Gaps = 2/83 (2%)
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKH--IKLQEQISLLKNRVNIASGTP 164
+P +L+++P+ ++ + + F +K + + + + + Q+ L+ I GTP
Sbjct: 74 APQILVLAPTRELAVQVAEAFSDFSKYMIGIHVLPLYGGQRYELQLRALRQGPQIVVGTP 133
Query: 165 SRIKKLIDVEALGLSRLQVLVLD 187
R+ + L LS L LVLD
Sbjct: 134 GRLLDHLKRGTLNLSNLHGLVLD 156
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,860,982
Number of Sequences: 164201
Number of extensions: 969050
Number of successful extensions: 3229
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 3189
Number of HSP's gapped (non-prelim): 65
length of query: 227
length of database: 59,974,054
effective HSP length: 106
effective length of query: 121
effective length of database: 42,568,748
effective search space: 5150818508
effective search space used: 5150818508
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC149471.20


