

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.4 - phase: 0 /partial
(542 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
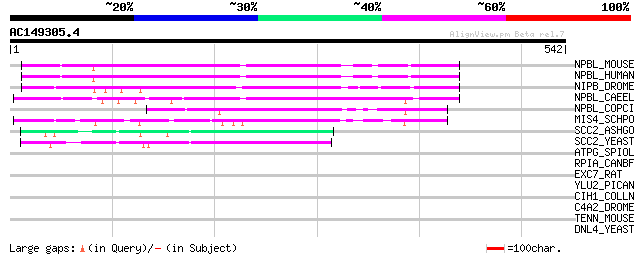
Sequences producing significant alignments: (bits) Value
NPBL_MOUSE (Q6KCD5) Nipped-B-like protein (Delangin homolog) (SC... 164 4e-40
NPBL_HUMAN (Q6KC79) Nipped-B-like protein (Delangin) (SCC2 homolog) 164 5e-40
NIPB_DROME (Q7PLI2) Nipped-B protein (SCC2 homolog) 142 3e-33
NPBL_CAEEL (Q95XZ5) Nipped-B-like protein pqn-85 (Prion-like-(Q/... 104 5e-22
NPBL_COPCI (Q00333) Rad9 protein (SCC2 homolog) 80 1e-14
MIS4_SCHPO (Q09725) Sister chromatid cohesion protein mis4 (SCC2... 69 2e-11
SCC2_ASHGO (Q750S2) Sister chromatid cohesion protein 2 52 4e-06
SCC2_YEAST (Q04002) Sister chromatid cohesion protein 2 46 3e-04
ATPG_SPIOL (P05435) ATP synthase gamma chain, chloroplast precur... 32 4.2
RPIA_CANBF (Q7VRG1) Ribose-5-phosphate isomerase A (EC 5.3.1.6) ... 32 5.5
EXC7_RAT (O54922) Exocyst complex component 7 (Exocyst complex c... 32 5.5
YLU2_PICAN (P34735) Hypothetical protein in LEU2 3'region (Fragm... 31 7.2
CIH1_COLLN (Q9P403) Intracellular hyphae protein 1 precursor 31 7.2
C4A2_DROME (Q9VMS8) Probable cytochrome P450 4ac2 (EC 1.14.-.-) ... 31 7.2
TENN_MOUSE (Q80Z71) Tenascin N precursor 31 9.4
DNL4_YEAST (Q08387) DNA ligase II (EC 6.5.1.1) (Polydeoxyribonuc... 31 9.4
>NPBL_MOUSE (Q6KCD5) Nipped-B-like protein (Delangin homolog) (SCC2
homolog)
Length = 2798
Score = 164 bits (416), Expect = 4e-40
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2044 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2102
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2103 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2162
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2163 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNSILSDKNSSVN 2222
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2223 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2277
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2278 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2337
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2338 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2385
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2386 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVTMLLYIADNLACFPYQTQEEP 2440
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2441 LFIMHHIDITLSVSGSNLLQSFK 2463
>NPBL_HUMAN (Q6KC79) Nipped-B-like protein (Delangin) (SCC2 homolog)
Length = 2804
Score = 164 bits (415), Expect = 5e-40
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2050 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2108
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2109 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2168
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2169 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 2228
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2229 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2283
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2284 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2343
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2344 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2391
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2392 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 2446
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2447 LFIMHHIDITLSVSGSNLLQSFK 2469
>NIPB_DROME (Q7PLI2) Nipped-B protein (SCC2 homolog)
Length = 2077
Score = 142 bits (357), Expect = 3e-33
Identities = 116/446 (26%), Positives = 216/446 (48%), Gaps = 36/446 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ S + LE+ L +++ + A V +C+ CL ++ +I
Sbjct: 1414 ILEKVVPLVNNASESFLASLEEHLMLLVVSRN-QAEVTSCVSCLGALVNKITHNFKLIRD 1472
Query: 72 LIQVFFKCLD---TEAVVNKQLVG--------RSLFCLGLLIRYGNC---LLASSGNKLV 117
Q F++ LD ++ + V RSLF +G+L+RY + + N +
Sbjct: 1473 CFQKFYRVLDVSRSQVIQGNNSVDNIYTPSFRRSLFTIGILMRYFDFKSPIALGETNDGL 1532
Query: 118 DVKRSLNLF---MKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIAD 174
V ++F M + + ++ ++L +LG + Y+ +++ + LSSIA+
Sbjct: 1533 PVSICEDVFHCLMFFCRCTNQEIRKQALISLGSFCVLNDGYLTRSELKNLYCEILSSIAN 1592
Query: 175 DR-LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQ 233
D KI ++N++ YL ++E M +E + + + V++G + IIQ
Sbjct: 1593 DAGFKIICMRNIWIYLTESEMFMHNKEKEWEKQSKHEDLKEMNDVSSG-----MASRIIQ 1647
Query: 234 LYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHH 293
LY IL ++ + VR A+K++++VLRQGLVHP+ VPYLI L TD ++ A
Sbjct: 1648 LYLEEILECFLNRDDTVRLWAVKVIQIVLRQGLVHPVRMVPYLICLSTDHRIESAHRADA 1707
Query: 294 LLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSL 353
LL ++ + Y F + GLQ+ F + + +N++ + +I + G D+
Sbjct: 1708 LLKDIDKTYSGFVNMKVQFGLQLCFKLQKIL------QINNRGKLEI-IRGYASRGPDNT 1760
Query: 354 AQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAP 413
+ +Y L+R + R + ++ ++FD+ K + + + Y + LA P+V
Sbjct: 1761 TTALNDF--LYTLLRTTKPQRRALVQTVTKQFDDQKTS---LQQMLYIADNLAYFPYVVQ 1815
Query: 414 DEPLYLIYTINRVVQVRAGPLEANFK 439
DEPLYLI+ I+ ++ + L A FK
Sbjct: 1816 DEPLYLIHQIDLLISMAGTHLLATFK 1841
>NPBL_CAEEL (Q95XZ5) Nipped-B-like protein pqn-85
(Prion-like-(Q/n-rich)-domain-bearing protein 85) (SCC2
homolog)
Length = 2203
Score = 104 bits (260), Expect = 5e-22
Identities = 104/470 (22%), Positives = 210/470 (44%), Gaps = 52/470 (11%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
Q+ + +I +++ V+PL+ +++ ++++L ++I+ + VV A + C+ S+ +
Sbjct: 1587 QVTKEMIGMLERVIPLVPFPSNIVLDSIDENLCKVIMFNGMALVVSA-VSCVASIYKKFK 1645
Query: 64 KGAAVIEHLIQVFFKCLDTEAVVNKQ---------------LVGRSLFCLGLLIRY---G 105
+GA + + K L+ V+ + ++ RS+F LG+L RY
Sbjct: 1646 RGATKTIDVFSTYLKHLE---VIKRNFDSNPRYDLDPKLFPILSRSIFTLGVLSRYFQFE 1702
Query: 106 NCLLASSGNKLVDVKR-----SLNLFMKYLAGEDYALKARSLQALGYVLIARPEYM---- 156
+ + V+ + +L F +Y G L+ ++L A+G+ Y+
Sbjct: 1703 EFVKEDPTEEKVEALKEKVFITLEFFSRYHKG---GLRQKALTAMGHFCAQHSTYLTKRQ 1759
Query: 157 LENDIGKILEGTLSSIADDRLKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSV 216
L N +IL +S + +I LQN+ +L E K+ + +
Sbjct: 1760 LTNTYLEILNAA-NSPQQQQQRILVLQNLEMFLQCEEQKLAASHDKWDENKEAQNLKEME 1818
Query: 217 PVAAGAGDTNICGGIIQLYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYL 276
+G G + +IQ YW +L VD + Q+R++A+++V + L QGLV P +P L
Sbjct: 1819 LSGSGLGSS-----VIQKYWKAVLESYVDADIQLRRAAVQVVWLTLNQGLVTPGASIPTL 1873
Query: 277 IALETDPLESNSKMAHHLLMNMHEKYPAFFESCLGDGLQMSF-MFMQSIFVSPDENVNHK 335
IA+ TDP++ LL + KY +S G+++S+ + ++ + ++ V
Sbjct: 1874 IAMTTDPVDVIRNRIDILLKEIDSKYSGMVQSKAMQGVRLSYKLHLKLRMLQQEKFVRGF 1933
Query: 336 SQSKIAVSGKGKPEADSLAQSRVGVSRIYKLIRGNRISRNKFMSSIVRKF------DNPK 389
++ + +S +Y+ +R NR R F+ S+V+ F D P+
Sbjct: 1934 RFCDFHLNTLPNALPEKTHDGMAVLSGLYQSLRTNRQQRRSFLQSMVKLFSEEFSHDKPQ 1993
Query: 390 WNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYTINRVVQVRAGPLEANFK 439
+++ + + LA+ P+ DEPLY++ I++ + L +K
Sbjct: 1994 LMEYI-----FIADNLAMFPYQMIDEPLYVMRQIDQNIAQTGQSLLVQYK 2038
>NPBL_COPCI (Q00333) Rad9 protein (SCC2 homolog)
Length = 2157
Score = 80.1 bits (196), Expect = 1e-14
Identities = 70/302 (23%), Positives = 135/302 (44%), Gaps = 27/302 (8%)
Query: 134 DYA-LKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQALQNMFEYLLDA 192
D+A ++ R LQ LG++ A+P M + + I++ +S ++ + + L+ M ++L+
Sbjct: 1636 DFADIRPRILQCLGFLFRAQPTLMTKEESAAIMDAIFAS-EEEEGRARLLKIMQDFLISE 1694
Query: 193 ESKMETEEVDG----KVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNILGRCVDFNT 248
K +E + V + V G D+ + I+Q Y ++IL + N+
Sbjct: 1695 SEKHSAKEKESAKNKNKANTDVNMEELVGNTDGFADSGVSSAIVQRYLSHILDAALSQNS 1754
Query: 249 QVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHEKYPAFFES 308
Q++ +A+ ++ ++QGL H + P +IALET P S A L ++ K+ + +
Sbjct: 1755 QIQMAAIDVLTFTIKQGLAHTLQSFPVIIALETSPHAVLSARAIALHSILNSKHASLLNT 1814
Query: 309 CLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGVSRIYKLIR 368
+ SF + + I + V H ++ G P A + R Y L+R
Sbjct: 1815 RYSISARKSFDYQKKIV----DGVVHGFRT------NGHPTA--------LLQRWYTLVR 1856
Query: 369 GNRISRNKFMSSIVRKF---DNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYTINR 425
R +R F+ S+V+ F D+ + + + F Y E A + +E +I +
Sbjct: 1857 EKRATRQDFLKSLVKVFSENDSYQATQDDVDFTRYMAENFASFEYKTQEEVFTVIKHLTT 1916
Query: 426 VV 427
V+
Sbjct: 1917 VL 1918
>MIS4_SCHPO (Q09725) Sister chromatid cohesion protein mis4 (SCC2
homolog)
Length = 1583
Score = 69.3 bits (168), Expect = 2e-11
Identities = 97/460 (21%), Positives = 188/460 (40%), Gaps = 72/460 (15%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
+ L ++ I VLP +++ S + LE L Q + + A + + CLCS+
Sbjct: 1056 RFLYYVVAIFRQVLPFQKEISESFLRSLESVLLQRLTKAG-TATLMEIVPCLCSLFTRLN 1114
Query: 64 KGAAVIEHLIQVFFKCLDT-----EAVVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVD 118
E L ++ CL + + N Q + R + +GL RYG+ + D
Sbjct: 1115 D----YERLKKIVVSCLKSLEEARHSENNFQKMVRLIDLIGLFSRYGDLNRIND-----D 1165
Query: 119 VKRSLNL---------------FMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGK 163
K SL+ F K L L+ + + + + I
Sbjct: 1166 WKHSLDFISPECDDAYVILLGYFQKLLKDAKGQLRIHIIDNMSRICLRETSLF----ISP 1221
Query: 164 ILEGTLSSI-ADDRL-KIQALQNMFEYLLDAESKMETEEVDGKVP---GHSVRAGQSVP- 217
++ TL I A++ + ++ L F LL A+ + E D K+ +V++ +SV
Sbjct: 1222 LMLSTLDMIIAENNVNEVSVLFKSFLELLAADEDL-IFEADQKLSLKGKQNVQSNKSVDR 1280
Query: 218 -VAAGAGDTN----ICGGIIQLYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITC 272
+ G D + ++Q + IL C N + ++I++ ++ QGLV+P C
Sbjct: 1281 DMLKGTKDKQWIEGVSASLMQHFLPCILDSCFSKNLRYSMLGIEILKCIIHQGLVNPRMC 1340
Query: 273 VPYLIALETDPLESNSKMAHHLLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENV 332
+IALE++ ++ ++A L +H ++ + + + F ++ +E
Sbjct: 1341 FSTIIALESNAIKETREVAILLHTELHRRHESLIDGLYAQSADLIFSLQKT-----EEYQ 1395
Query: 333 NHKSQSKIAVSGKGKPEADSLAQSRVGVSRIYKLIRGNRISRNKFMSSIVRK-----FDN 387
K G+ A + V + K SR K + I++ D
Sbjct: 1396 TFK---------LGEFSPFQSAYTIVSADKSSK-------SRKKLIMQILKPLKLDGIDL 1439
Query: 388 PKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYTINRVV 427
P + + ++F+++C LA +P+V+ +EPL +I T++ V+
Sbjct: 1440 PSFTEEKVSFVSFCCVCLAGIPYVSIEEPLMIISTVDSVL 1479
>SCC2_ASHGO (Q750S2) Sister chromatid cohesion protein 2
Length = 1479
Score = 52.0 bits (123), Expect = 4e-06
Identities = 63/320 (19%), Positives = 127/320 (39%), Gaps = 30/320 (9%)
Query: 11 FIIDSVLPLLRKLPPSIVEELEQ--DLKQMILRH--SFLAVVHACIKCLCSMSELAGKGA 66
F+ D +L +LP V EL++ L + RH + AC CL +S
Sbjct: 975 FLYDLETTILSRLPRMNVRELDEAIPLSWSLSRHRKDDTRICKACASCLGQLSPYIANAT 1034
Query: 67 AVIEHLIQVFFKCLDTEAVVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSLNLF 126
D AV + R L+ R+ C ++ K ++K N+F
Sbjct: 1035 T-------------DPSAVRPDGKLQRLLYLATGFARF--CSFENTEGKFPNLKTRENIF 1079
Query: 127 ------MKYLAGEDYALKARSLQALGYVLIAR--PEYMLENDIGKILEGTLSSIADDRLK 178
+ E+ R + V IA P+ + +L+ D ++
Sbjct: 1080 EYVTKCLLMFTKENIHHVIRRIATKNLVKIASRYPKLFNSRHVLTVLDTQFEKGILD-IQ 1138
Query: 179 IQALQNMFEYLLDAESKMETEE-VDGKVPGHS-VRAGQSVPVAAGAGDTNICGGIIQLYW 236
+ L++++++ L E + + VDG + ++ +R + + + + IC ++ Y
Sbjct: 1139 LVVLESLYDFFLAEERRSLIQVGVDGTISSNNELRKVVANHTKSDSINDGICSALVSRYL 1198
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+ IL C+ + A++ ++ +L G +P CVP +IAL P ++ +
Sbjct: 1199 DKILKICLIADLNNAMVAIRFLKQILTYGYTNPSLCVPTVIALTASPNSYMRDLSSEMFS 1258
Query: 297 NMHEKYPAFFESCLGDGLQM 316
+ + Y + +CL G+++
Sbjct: 1259 ELLQGYESMAFNCLNQGIRL 1278
>SCC2_YEAST (Q04002) Sister chromatid cohesion protein 2
Length = 1493
Score = 45.8 bits (107), Expect = 3e-04
Identities = 61/319 (19%), Positives = 133/319 (41%), Gaps = 32/319 (10%)
Query: 11 FIIDSVLPLLRKLPPSIVEELEQDLKQM----ILRHSFLAVVHACIKCLCSMSELAGKGA 66
F+ D LL +LP V E+++ + + RH V AC CL
Sbjct: 990 FLYDLETTLLSRLPKMNVREIDEAMPLIWSVATHRHDTARVAKACSSCL----------- 1038
Query: 67 AVIEHLIQVFFKCLDTEA-VVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSLNL 125
HL K + EA +V + R ++ R+ C S +K+ ++ L
Sbjct: 1039 ---SHLHPYINKANNEEAAIVVDGKLQRLIYLSTGFARF--CFPKPSNDKIAFLQEGETL 1093
Query: 126 FMKY-----LAGED---YALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRL 177
+ + +D + ++ +++ L + P+ + +L+ S D +
Sbjct: 1094 YEHITKCLLVLSKDKITHVIRRVAVKNLTKLCGNHPKLFNSRHVLHLLDKEFQSDQLD-I 1152
Query: 178 KIQALQNMFE-YLLDAESKMETEEVDGKVPGHSVRAGQSVPV-AAGAGDTNICGGIIQLY 235
K+ L+++++ +LL+ + V+ + +S+ + + + +C + +
Sbjct: 1153 KLVILESLYDLFLLEERKSVRNTGVNSTLSSNSILKKKLLKTNRVEFANDGVCSALATRF 1212
Query: 236 WNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLL 295
+NIL C+ + + A+++++++L+ G +P +P +IAL + +A+ LL
Sbjct: 1213 LDNILQLCLLRDLKNSLVAIRLLKLILKFGYTNPSHSIPTVIALFASTSQYIRHVAYELL 1272
Query: 296 MNMHEKYPAFFESCLGDGL 314
++ EKY S L G+
Sbjct: 1273 EDLFEKYETLVFSSLSRGV 1291
>ATPG_SPIOL (P05435) ATP synthase gamma chain, chloroplast precursor
(EC 3.6.3.14)
Length = 364
Score = 32.0 bits (71), Expect = 4.2
Identities = 18/58 (31%), Positives = 31/58 (53%), Gaps = 7/58 (12%)
Query: 183 QNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNIL 240
+ + E L + +++TE+VD VP +R + V + GD +CGG +NN+L
Sbjct: 87 ETLVEVLYNMNEQLQTEDVD--VPLTKIRTVKKVALMVVTGDRGLCGG-----FNNML 137
>RPIA_CANBF (Q7VRG1) Ribose-5-phosphate isomerase A (EC 5.3.1.6)
(Phosphoriboisomerase A) (PRI)
Length = 222
Score = 31.6 bits (70), Expect = 5.5
Identities = 13/32 (40%), Positives = 21/32 (65%)
Query: 88 KQLVGRSLFCLGLLIRYGNCLLASSGNKLVDV 119
+ +V +SL CLG L Y N ++ +GN ++DV
Sbjct: 142 RSVVSKSLICLGGLPEYRNGVITDNGNSILDV 173
>EXC7_RAT (O54922) Exocyst complex component 7 (Exocyst complex
component Exo70) (rExo70)
Length = 653
Score = 31.6 bits (70), Expect = 5.5
Identities = 46/185 (24%), Positives = 78/185 (41%), Gaps = 17/185 (9%)
Query: 7 ESIIFIIDSV--LPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGK 64
E++ FI DS+ L K SI+ E L M L +S + V H + L + E K
Sbjct: 23 ETLSFIRDSLEKSDQLTKNMVSILSSFESRL--MKLENSIIPV-HKQTENLQRLQENVEK 79
Query: 65 GAAVIEHLIQVFFKCLDTEAVVNKQLVGRSLFCLGLLIRYGNCLLASSGN-----KLVDV 119
+ ++H+I + DTE ++ + GR LG + + + N +L V
Sbjct: 80 TLSCLDHVISYYHVASDTEKIIREGPTGRLEEYLGSMAKIQKAVEYFQDNSPDSPELNKV 139
Query: 120 KRSLNLFMKYLAGEDYALKARSLQALGYVLI-----ARPEYMLENDIGKILEGTLSSIAD 174
K + L E +L R + + VL+ A E ++ D+ +LE S+
Sbjct: 140 KLLFERGKESLESEFRSLMTRHSKVISPVLVLDLISADDELEVQEDV--VLEHLPESVLQ 197
Query: 175 DRLKI 179
D ++I
Sbjct: 198 DVIRI 202
>YLU2_PICAN (P34735) Hypothetical protein in LEU2 3'region
(Fragment)
Length = 373
Score = 31.2 bits (69), Expect = 7.2
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 1/74 (1%)
Query: 313 GLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSL-AQSRVGVSRIYKLIRGNR 371
GL + F+F + +N++ K S I S +P A S A + +G +RI +RG +
Sbjct: 273 GLLLLFLFWRRRQRDDRDNLSEKRASSILASSSRQPPAGSRGAAAGIGANRIRIHVRGRQ 332
Query: 372 ISRNKFMSSIVRKF 385
+ ++ R+F
Sbjct: 333 TGHAGHVETVQRRF 346
>CIH1_COLLN (Q9P403) Intracellular hyphae protein 1 precursor
Length = 230
Score = 31.2 bits (69), Expect = 7.2
Identities = 13/28 (46%), Positives = 18/28 (63%)
Query: 201 VDGKVPGHSVRAGQSVPVAAGAGDTNIC 228
VDGK+ H V++G+S+ A DT IC
Sbjct: 103 VDGKIKTHKVKSGESLTTIAEKYDTGIC 130
>C4A2_DROME (Q9VMS8) Probable cytochrome P450 4ac2 (EC 1.14.-.-)
(CYPIVAC2)
Length = 497
Score = 31.2 bits (69), Expect = 7.2
Identities = 23/87 (26%), Positives = 43/87 (48%), Gaps = 4/87 (4%)
Query: 226 NICGGIIQLYWNNILG-RCVDFNTQVR-QSALKIVEVVLRQGLVHPI--TCVPYLIALET 281
N+CG I + LG + D + +R + ++ +E V++Q L +P V + + +
Sbjct: 173 NVCGNAINVIAETALGVKLDDLSEGIRYRQSIHAIEEVMQQRLCNPFFYNIVYFFLFGDY 232
Query: 282 DPLESNSKMAHHLLMNMHEKYPAFFES 308
+N K+AH N+ EK + F+S
Sbjct: 233 RKQVNNLKIAHEFSSNIIEKRRSLFKS 259
>TENN_MOUSE (Q80Z71) Tenascin N precursor
Length = 1560
Score = 30.8 bits (68), Expect = 9.4
Identities = 23/89 (25%), Positives = 35/89 (38%)
Query: 442 SSSLLQSEGQGTPHGNGMYQRATDETIHSTQGQSMDLNGPFQQNVDVQPYVDDMTSVDLN 501
+S LL + TP G + T+ T + + Q D Q DD TS+
Sbjct: 16 ASVLLVASAPATPESPGCSNKEQQVTVSHTYKIDVPKSALVQVETDPQSLSDDGTSLLAP 75
Query: 502 GTNHQLPDYPLSHNGRLKVKPQAAGFADS 530
G + + + HN RL+ + ADS
Sbjct: 76 GEDGEEQNIIFRHNIRLQTPQKNCDLADS 104
>DNL4_YEAST (Q08387) DNA ligase II (EC 6.5.1.1)
(Polydeoxyribonucleotide synthase [ATP]) (DNA ligase IV
homolog)
Length = 944
Score = 30.8 bits (68), Expect = 9.4
Identities = 41/159 (25%), Positives = 66/159 (40%), Gaps = 25/159 (15%)
Query: 78 KCLDTEAVVNKQLVGRSLFCLGLLIRYGNCL--LASSGNKLVDVKRS-LNLFMKYL---- 130
+C E++ V SL G++++Y N +AS N + VK L F + L
Sbjct: 422 RCYGVESIKKSLEVAISLGSEGVVLKYYNSSYNVASRNNNWIKVKPEYLEEFGENLDLIV 481
Query: 131 AGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQALQNM----- 185
G D K + LG +++ EY K +G S I D + + +QN
Sbjct: 482 IGRDSGKKDSFM--LGLLVLDEEEY-------KKHQGDSSEIVDHSSQEKHIQNSRRRVK 532
Query: 186 ----FEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAA 220
F + + S+ E +E+D K GH R + P A+
Sbjct: 533 KILSFCSIANGISQEEFKEIDRKTRGHWKRTSEVAPPAS 571
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 62,093,788
Number of Sequences: 164201
Number of extensions: 2609021
Number of successful extensions: 6109
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6073
Number of HSP's gapped (non-prelim): 21
length of query: 542
length of database: 59,974,054
effective HSP length: 115
effective length of query: 427
effective length of database: 41,090,939
effective search space: 17545830953
effective search space used: 17545830953
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC149305.4


