

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.10 + phase: 0 /pseudo
(55 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
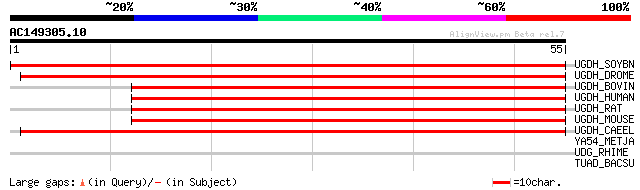
Score E
Sequences producing significant alignments: (bits) Value
UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 81 5e-16
UGDH_DROME (O02373) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 59 3e-09
UGDH_BOVIN (P12378) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 57 9e-09
UGDH_HUMAN (O60701) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 57 1e-08
UGDH_RAT (O70199) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (UDP... 55 3e-08
UGDH_MOUSE (O70475) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 55 3e-08
UGDH_CAEEL (Q19905) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 51 5e-07
YA54_METJA (Q58454) Hypothetical protein MJ1054 (EC 1.1.1.-) [Co... 38 0.006
UDG_RHIME (O54068) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (UD... 31 0.54
TUAD_BACSU (O32271) UDP-glucose 6-dehydrogenase (EC 1.1.1.22) (U... 29 2.7
>UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 480
Score = 81.3 bits (199), Expect = 5e-16
Identities = 38/55 (69%), Positives = 45/55 (81%)
Query: 1 MAAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
MA IALK S +V VVDI+KSRI AW +DQ+PIY+PG DG+VKQCR KNLFF T+
Sbjct: 17 MAVIALKCPSIEVAVVDISKSRIAAWNSDQLPIYEPGLDGVVKQCRGKNLFFSTD 71
>UGDH_DROME (O02373) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH) (Sugarless
protein)
Length = 476
Score = 58.5 bits (140), Expect = 3e-09
Identities = 26/54 (48%), Positives = 37/54 (68%)
Query: 2 AAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
A +ALK +T+VD + RI W +D++PIY+PG D +VK+CR NLFF T+
Sbjct: 17 AVMALKCPDIVITLVDKSSERIAQWNSDKLPIYEPGLDEVVKKCRNVNLFFSTD 70
>UGDH_BOVIN (P12378) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 494
Score = 57.0 bits (136), Expect = 9e-09
Identities = 26/43 (60%), Positives = 32/43 (73%)
Query: 13 VTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
VTVVDI +SRI AW + +PIY+PG +V+ CR KNLFF TN
Sbjct: 32 VTVVDINESRINAWNSPTLPIYEPGLKEVVESCRGKNLFFSTN 74
>UGDH_HUMAN (O60701) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 494
Score = 56.6 bits (135), Expect = 1e-08
Identities = 25/43 (58%), Positives = 32/43 (74%)
Query: 13 VTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
VTVVD+ +SRI AW + +PIY+PG +V+ CR KNLFF TN
Sbjct: 32 VTVVDVNESRINAWNSPTLPIYEPGLKEVVESCRGKNLFFSTN 74
>UGDH_RAT (O70199) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 493
Score = 55.5 bits (132), Expect = 3e-08
Identities = 24/43 (55%), Positives = 32/43 (73%)
Query: 13 VTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
VTVVD+ ++RI AW + +PIY+PG +V+ CR KNLFF TN
Sbjct: 32 VTVVDVNEARINAWNSPTLPIYEPGLKEVVESCRGKNLFFSTN 74
>UGDH_MOUSE (O70475) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 493
Score = 55.5 bits (132), Expect = 3e-08
Identities = 24/43 (55%), Positives = 32/43 (73%)
Query: 13 VTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
VTVVD+ ++RI AW + +PIY+PG +V+ CR KNLFF TN
Sbjct: 32 VTVVDVNEARINAWNSPTLPIYEPGLKEVVESCRGKNLFFSTN 74
>UGDH_CAEEL (Q19905) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH) (Squashed
vulva protein 4)
Length = 481
Score = 51.2 bits (121), Expect = 5e-07
Identities = 24/54 (44%), Positives = 35/54 (64%)
Query: 2 AAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
A IA K VTVVD+ ++I W +D++PIY+PG D +V R +NLFF ++
Sbjct: 26 AMIAHKCPHITVTVVDMNTAKIAEWNSDKLPIYEPGLDEIVFAARGRNLFFSSD 79
>YA54_METJA (Q58454) Hypothetical protein MJ1054 (EC 1.1.1.-)
[Contains: Mja UDPGD intein]
Length = 895
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/53 (35%), Positives = 31/53 (57%)
Query: 3 AIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
A+ L DV +DI +S++ A + P+Y+ G +GL+K+ KNL F T+
Sbjct: 16 AVGLAEFGFDVVGIDIDESKVKALNRGECPLYEEGLEGLLKKHVNKNLTFTTS 68
>UDG_RHIME (O54068) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
Length = 437
Score = 31.2 bits (69), Expect = 0.54
Identities = 15/31 (48%), Positives = 19/31 (60%)
Query: 12 DVTVVDITKSRIVAWYNDQIPIYQPGFDGLV 42
DV VD + +I A QIPI++PG D LV
Sbjct: 25 DVVCVDKDEGKISALKKGQIPIFEPGLDHLV 55
>TUAD_BACSU (O32271) UDP-glucose 6-dehydrogenase (EC 1.1.1.22)
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH)
(Teichuronic acid biosynthesis protein tuaD)
Length = 461
Score = 28.9 bits (63), Expect = 2.7
Identities = 14/32 (43%), Positives = 20/32 (61%)
Query: 13 VTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQ 44
V DI +S+I + N IPIY+PG LV++
Sbjct: 27 VVCCDIDESKIRSLKNGVIPIYEPGLADLVEK 58
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,162,429
Number of Sequences: 164201
Number of extensions: 174786
Number of successful extensions: 484
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 474
Number of HSP's gapped (non-prelim): 10
length of query: 55
length of database: 59,974,054
effective HSP length: 31
effective length of query: 24
effective length of database: 54,883,823
effective search space: 1317211752
effective search space used: 1317211752
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149305.10


