

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149293.2 - phase: 2
(212 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
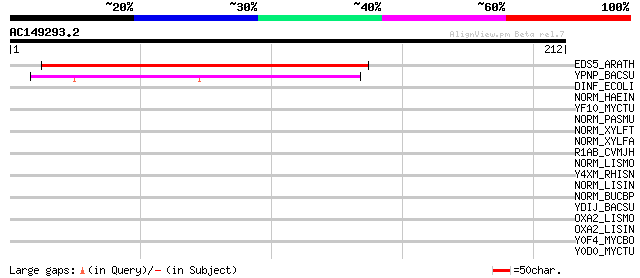
Score E
Sequences producing significant alignments: (bits) Value
EDS5_ARATH (Q945F0) Enhanced disease susceptibility 5 (Eds5) (Sa... 179 5e-45
YPNP_BACSU (P54181) Hypothetical protein ypnP 44 3e-04
DINF_ECOLI (P28303) DNA-damage-inducible protein F 37 0.026
NORM_HAEIN (P45272) Probable multidrug resistance protein norM (... 37 0.034
YF10_MYCTU (P71789) Hypothetical protein Rv1510/MT1558/MT1560 36 0.075
NORM_PASMU (Q9CMZ9) Probable multidrug resistance protein norM (... 35 0.13
NORM_XYLFT (Q879Z5) Probable multidrug resistance protein norM (... 32 1.4
NORM_XYLFA (Q9PA34) Probable multidrug resistance protein norM (... 31 1.8
R1AB_CVMJH (P19751) Replicase polyprotein 1ab (pp1ab) (ORF1ab po... 31 2.4
NORM_LISMO (Q8Y654) Probable multidrug resistance protein norM (... 31 2.4
Y4XM_RHISN (P55705) Hypothetical transport protein y4xM 30 3.1
NORM_LISIN (Q92AG2) Probable multidrug resistance protein norM (... 30 3.1
NORM_BUCBP (Q89AX2) Multidrug resistance protein norM (Na(+)/dru... 30 3.1
YDIJ_BACSU (O05523) Hypothetical protein ydiJ 29 7.0
OXA2_LISMO (Q8Y7A9) Membrane protein oxaA 2 precursor 29 7.0
OXA2_LISIN (Q92BX6) Membrane protein oxaA 2 precursor 29 7.0
Y0F4_MYCBO (P59985) Hypothetical protein Mb3654 29 9.2
Y0D0_MYCTU (O06377) Hypothetical protein Rv3630/MT3732 29 9.2
>EDS5_ARATH (Q945F0) Enhanced disease susceptibility 5 (Eds5)
(Salicylic acid induction deficient 1) (Sid1)
Length = 543
Score = 179 bits (453), Expect = 5e-45
Identities = 86/125 (68%), Positives = 106/125 (84%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L G VAQSASLGMK+SWGPLKALAAA++ING+GD +LC +LG GIAGAAWAT ASQ+V+A
Sbjct: 235 LVGLVAQSASLGMKNSWGPLKALAAATIINGLGDTILCLFLGQGIAGAAWATTASQIVSA 294
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM +LN +GYNA++ +IPS +E I LAAPVF+++ SK+AFYS +IY ATSMGTH
Sbjct: 295 YMMMDSLNKEGYNAYSFAIPSPQELWKISALAAPVFISIFSKIAFYSFIIYCATSMGTHV 354
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 355 LAAHQ 359
>YPNP_BACSU (P54181) Hypothetical protein ypnP
Length = 445
Score = 43.9 bits (102), Expect = 3e-04
Identities = 38/128 (29%), Positives = 59/128 (45%), Gaps = 2/128 (1%)
Query: 9 GLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
G+L+L G+ S L + DS PL+ +A A V+N V + + GIAGAA++T+ S
Sbjct: 142 GILFLFGYNFISTVLRALGDSKTPLRFIAFAVVLNTVLAPLFISVFRMGIAGAAYSTILS 201
Query: 68 QVVA-AYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFAT 126
Q +A Y + + K +P E IL L P + MM ++
Sbjct: 202 QGIAFLYGLFYVIKHKLVPFSIPRMPKWEESALILKLGIPAGLQMMVITGGMMAIMSVVN 261
Query: 127 SMGTHTMA 134
S G H ++
Sbjct: 262 SYGDHVVS 269
>DINF_ECOLI (P28303) DNA-damage-inducible protein F
Length = 459
Score = 37.4 bits (85), Expect = 0.026
Identities = 32/138 (23%), Positives = 61/138 (44%), Gaps = 11/138 (7%)
Query: 5 SIPAGL--LYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAW 62
S PA L L L GW+ LG++ + P+ L +++N V D+ L L + GAA
Sbjct: 156 SAPASLANLVLLGWL-----LGVQYARAPVILLVVGNILNIVLDVWLVMGLHMNVQGAAL 210
Query: 63 ATM----ASQVVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFY 118
AT+ A+ ++ M+ + L ++G + L F +L L + + + +
Sbjct: 211 ATVIAEYATLLIGLLMVRKILKLRGISGEMLKTAWRGNFRRLLALNRDIMLRSLLLQLCF 270
Query: 119 SLLIYFATSMGTHTMAAH 136
+ +G+ +A +
Sbjct: 271 GAITVLGARLGSDIIAVN 288
>NORM_HAEIN (P45272) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 464
Score = 37.0 bits (84), Expect = 0.034
Identities = 23/93 (24%), Positives = 46/93 (48%), Gaps = 3/93 (3%)
Query: 53 LGYGIAGAA--WATMASQVVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMT 110
+G GIA A WA + +Y + ++K ++ + +P+ + +L L P+ +
Sbjct: 195 VGCGIATAIVNWAMCLMMIFYSYTNTQERSLKVFSQL-IEMPNPKTLKKLLRLGLPIAIA 253
Query: 111 MMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLS 143
+ +VA Y+L + +G +A+HQ +N S
Sbjct: 254 ICCEVALYALTSLMLSPLGATIVASHQITLNTS 286
>YF10_MYCTU (P71789) Hypothetical protein Rv1510/MT1558/MT1560
Length = 432
Score = 35.8 bits (81), Expect = 0.075
Identities = 27/99 (27%), Positives = 44/99 (44%), Gaps = 7/99 (7%)
Query: 1 VDSRSIPAGLLYL--CGWVAQSASLGMK---DSWGPLKALAAASVINGVGDIVLCTYLGY 55
V+ R + GLL + G+ AQ+ LG D W +L + + +G+
Sbjct: 137 VEGRWLSVGLLSVGVAGFCAQATLLGALAGVDRWTQYGSLMVTDAVIRLAVAAAAVVIGW 196
Query: 56 GIAGAAWATMASQVVAAYMMMRTLNMKGYNAFALSIPSG 94
G+AG WA A V A+++M + +A +L P G
Sbjct: 197 GLAGYLWAATAGAV--AWLLMLMASPTARSAASLLTPGG 233
>NORM_PASMU (Q9CMZ9) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 464
Score = 35.0 bits (79), Expect = 0.13
Identities = 25/100 (25%), Positives = 45/100 (45%), Gaps = 9/100 (9%)
Query: 53 LGYGIAGAAWATMASQVVAAYMMMRTL-------NMKGYNAFA--LSIPSGREFITILGL 103
LG GA +A+ +V +M + + N + FA L P+ +L L
Sbjct: 187 LGIPAFGAVGCGIATAIVNWFMCILMIAYCKNARNQRDLKVFAKILEKPNFTTLKKLLNL 246
Query: 104 AAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLS 143
P+ + + +VA ++L + +GT +A+HQ +N S
Sbjct: 247 GFPIAVALFCEVALFALTALLLSPLGTDIVASHQIALNTS 286
>NORM_XYLFT (Q879Z5) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 469
Score = 31.6 bits (70), Expect = 1.4
Identities = 24/114 (21%), Positives = 49/114 (42%), Gaps = 13/114 (11%)
Query: 55 YGIAGAAWATMA----SQVVAAYMMMRTLNMKGYNAFA-LSIPSGREFITILGLAAPVFM 109
YG+ G AT+ VV A + R+ FA L +P +L + P+ +
Sbjct: 209 YGVEGLGIATVTVMWLQAVVFALYLWRSRRFAHLQLFAHLELPCWARIRDLLNIGLPIGI 268
Query: 110 TMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCYMELIGICQ 163
+++ + + + GT +AAHQ +++++ C+M +G+ +
Sbjct: 269 SILMEGGLFIVTTLLIGRFGTDEIAAHQIALSVAQL--------CFMIPMGVAE 314
>NORM_XYLFA (Q9PA34) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 469
Score = 31.2 bits (69), Expect = 1.8
Identities = 24/114 (21%), Positives = 49/114 (42%), Gaps = 13/114 (11%)
Query: 55 YGIAGAAWATMA----SQVVAAYMMMRTLNMKGYNAFA-LSIPSGREFITILGLAAPVFM 109
YG+ G AT+ VV A + R+ FA L +P +L + P+ +
Sbjct: 209 YGVEGLGIATVMVMWLQAVVFALYLWRSRRFAHLQLFAHLELPCWARIRDLLNIGLPIGI 268
Query: 110 TMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCYMELIGICQ 163
+++ + + + GT +AAHQ +++++ C+M +G+ +
Sbjct: 269 SILMEGGLFIVTTLLIGRFGTDEIAAHQIALSVAQL--------CFMIPMGVAE 314
>R1AB_CVMJH (P19751) Replicase polyprotein 1ab (pp1ab) (ORF1ab
polyprotein) [Includes: Replicase polyprotein 1a (pp1a)
(ORF1a)] [Contains: p28; p65; p210 (EC 3.4.24.-)
(Papain-like proteinases 1/2) (PL1-PRO/PL2-PRO); Peptide
HD2 (p44); 3C-like proteinase
Length = 7180
Score = 30.8 bits (68), Expect = 2.4
Identities = 13/44 (29%), Positives = 25/44 (56%)
Query: 118 YSLLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCYMELIGI 161
Y +IY T+ H++ +++ V ++R LC+ M+LIG+
Sbjct: 5928 YDFVIYSQTAQTAHSVNVNRFNVAITRAKKGILCVMSSMQLIGV 5971
>NORM_LISMO (Q8Y654) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 456
Score = 30.8 bits (68), Expect = 2.4
Identities = 20/87 (22%), Positives = 43/87 (48%), Gaps = 5/87 (5%)
Query: 56 GIAGAAWATMASQ----VVAAYMMMRTLNMKGYNAF-ALSIPSGREFITILGLAAPVFMT 110
G AG+ +AT + +V+ ++ ++ + F L+ + I+G+ P +T
Sbjct: 192 GGAGSGYATGITYWLVVLVSVILIQTQTRLRKFGVFKGLTALRFSKIKEIIGIGVPNGLT 251
Query: 111 MMSKVAFYSLLIYFATSMGTHTMAAHQ 137
++ + + +S + + GT T+AAHQ
Sbjct: 252 ILFETSIFSAVTILMGAFGTETIAAHQ 278
>Y4XM_RHISN (P55705) Hypothetical transport protein y4xM
Length = 404
Score = 30.4 bits (67), Expect = 3.1
Identities = 26/92 (28%), Positives = 40/92 (43%), Gaps = 9/92 (9%)
Query: 19 QSASLGMKDSWGP-----LKALAAASVINGVGDI--VLCTYLGYGIAGAAWATMASQVVA 71
Q S+G + GP +KAL+ + + G+ I +C LG + A W M +
Sbjct: 22 QFISMGAMEMNGPFWPIQIKALSPSDSVFGLAGIGVYVCPMLGVSLTSAFWGRMGDRYGN 81
Query: 72 AYMMMRTLNMKGYNAFALSIPSGREFITILGL 103
MM+R L G L + ++ TIL L
Sbjct: 82 RLMMVRAL--LGLAITQLLVAFAQDVWTILAL 111
>NORM_LISIN (Q92AG2) Probable multidrug resistance protein norM
(Na(+)/drug antiporter) (Multidrug-efflux transporter)
Length = 456
Score = 30.4 bits (67), Expect = 3.1
Identities = 22/90 (24%), Positives = 44/90 (48%), Gaps = 11/90 (12%)
Query: 56 GIAGAAWATMASQ----VVAAYMMMRTLNMKGYNAF----ALSIPSGREFITILGLAAPV 107
G AG+ +AT + +V+ ++ ++ + F AL +E I+G+ P
Sbjct: 192 GGAGSGYATGITYWLVVLVSVILIQTQTRLRKFGIFKGLTALRFSKVKE---IIGIGVPN 248
Query: 108 FMTMMSKVAFYSLLIYFATSMGTHTMAAHQ 137
+T++ + + +S + + GT T+AAHQ
Sbjct: 249 GLTILFETSIFSAVTILMGAFGTETIAAHQ 278
>NORM_BUCBP (Q89AX2) Multidrug resistance protein norM (Na(+)/drug
antiporter) (Multidrug-efflux transporter)
Length = 451
Score = 30.4 bits (67), Expect = 3.1
Identities = 12/62 (19%), Positives = 31/62 (49%)
Query: 82 KGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVN 141
K + + +P+ + + + P+ +++ ++ ++L+ SM T + AHQ +N
Sbjct: 225 KNISNLEMYLPNYKIIWNLFKMGFPIALSLFCEITLFTLITLLIASMETFQIIAHQIALN 284
Query: 142 LS 143
+S
Sbjct: 285 IS 286
>YDIJ_BACSU (O05523) Hypothetical protein ydiJ
Length = 254
Score = 29.3 bits (64), Expect = 7.0
Identities = 11/29 (37%), Positives = 20/29 (68%)
Query: 97 FITILGLAAPVFMTMMSKVAFYSLLIYFA 125
F+T LG+ P+F+ + K A+++LL+ A
Sbjct: 177 FLTRLGIVTPMFLAKIRKYAYFTLLVIAA 205
>OXA2_LISMO (Q8Y7A9) Membrane protein oxaA 2 precursor
Length = 275
Score = 29.3 bits (64), Expect = 7.0
Identities = 21/85 (24%), Positives = 42/85 (48%), Gaps = 7/85 (8%)
Query: 41 INGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMKGYNAFALSIPSGREFITI 100
I G +I T+L + + G+ +A Y+ ++M GY+ P ++ + I
Sbjct: 153 IRGSSEIASHTFLWFNL-GSPDMVLAIIAGLVYLAQYFVSMIGYS------PEQKKQMKI 205
Query: 101 LGLAAPVFMTMMSKVAFYSLLIYFA 125
+GL +P+ + +S A +L +Y+A
Sbjct: 206 IGLMSPIMILFVSFTAPSALALYWA 230
>OXA2_LISIN (Q92BX6) Membrane protein oxaA 2 precursor
Length = 275
Score = 29.3 bits (64), Expect = 7.0
Identities = 21/85 (24%), Positives = 42/85 (48%), Gaps = 7/85 (8%)
Query: 41 INGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMKGYNAFALSIPSGREFITI 100
I G +I T+L + + G+ +A Y+ ++M GY+ P ++ + I
Sbjct: 153 IRGSSEIASHTFLWFNL-GSPDMVLAIIAGLVYLAQYFVSMIGYS------PEQKKQMKI 205
Query: 101 LGLAAPVFMTMMSKVAFYSLLIYFA 125
+GL +P+ + +S A +L +Y+A
Sbjct: 206 IGLMSPIMILFVSFTAPSALALYWA 230
>Y0F4_MYCBO (P59985) Hypothetical protein Mb3654
Length = 431
Score = 28.9 bits (63), Expect = 9.2
Identities = 19/67 (28%), Positives = 28/67 (41%), Gaps = 3/67 (4%)
Query: 13 LCGWVAQSASLGM---KDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQV 69
L G+ + LGM + W AL A + V +G+ + G WAT+A V
Sbjct: 150 LAGFCLHATLLGMLAGTNRWTQYGALMVADAVIRVVVAAATFVIGWQLVGFIWATVAGSV 209
Query: 70 VAAYMMM 76
M+M
Sbjct: 210 AWLIMLM 216
>Y0D0_MYCTU (O06377) Hypothetical protein Rv3630/MT3732
Length = 431
Score = 28.9 bits (63), Expect = 9.2
Identities = 19/67 (28%), Positives = 28/67 (41%), Gaps = 3/67 (4%)
Query: 13 LCGWVAQSASLGM---KDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQV 69
L G+ + LGM + W AL A + V +G+ + G WAT+A V
Sbjct: 150 LAGFCLHATLLGMLAGTNRWTQYGALMVADAVIRVVVAAATFVIGWQLVGFIWATVAGSV 209
Query: 70 VAAYMMM 76
M+M
Sbjct: 210 AWLIMLM 216
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,855,872
Number of Sequences: 164201
Number of extensions: 763682
Number of successful extensions: 2677
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2665
Number of HSP's gapped (non-prelim): 20
length of query: 212
length of database: 59,974,054
effective HSP length: 106
effective length of query: 106
effective length of database: 42,568,748
effective search space: 4512287288
effective search space used: 4512287288
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149293.2


