

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149209.11 - phase: 0
(607 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
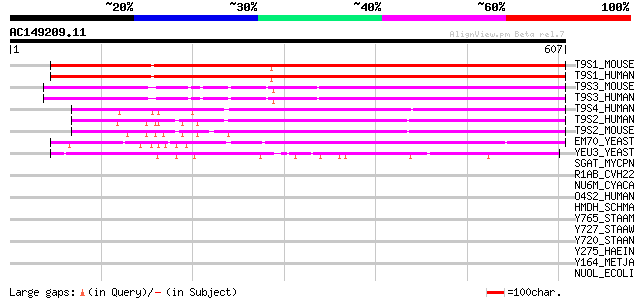
Score E
Sequences producing significant alignments: (bits) Value
T9S1_MOUSE (Q9DBU0) Transmembrane 9 superfamily protein member 1... 518 e-146
T9S1_HUMAN (O15321) Transmembrane 9 superfamily protein member 1... 511 e-144
T9S3_MOUSE (Q9ET30) Transmembrane 9 superfamily protein member 3... 417 e-116
T9S3_HUMAN (Q9HD45) Transmembrane 9 superfamily protein member 3... 416 e-116
T9S4_HUMAN (Q92544) Transmembrane 9 superfamily protein member 4 367 e-101
T9S2_HUMAN (Q99805) Transmembrane 9 superfamily protein member 2... 323 8e-88
T9S2_MOUSE (P58021) Transmembrane 9 superfamily protein member 2... 320 9e-87
EM70_YEAST (P32802) Endosomal P24A protein precursor (70 kDa end... 289 1e-77
YEU3_YEAST (P40071) Hypothetical 81.5 kDa protein in USS1-BEB1 i... 131 6e-30
SGAT_MYCPN (P75291) Putative transport protein sgaT homolog 36 0.26
R1AB_CVH22 (Q05002) Replicase polyprotein 1ab (pp1ab) (ORF1ab po... 35 0.74
NU6M_CYACA (P48925) NADH-ubiquinone oxidoreductase chain 6 (EC 1... 34 0.97
O4S2_HUMAN (Q8NH73) Olfactory receptor 4S2 34 1.3
HMDH_SCHMA (P16237) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 34 1.3
Y765_STAAM (P67108) Hypothetical UPF0042 protein SAV0765 33 2.8
Y727_STAAW (P67110) Hypothetical UPF0042 protein MW0727 33 2.8
Y720_STAAN (P67109) Hypothetical UPF0042 protein SA0720 33 2.8
Y275_HAEIN (P43975) Hypothetical protein HI0275 32 3.7
Y164_METJA (Q57628) Hypothetical protein MJ0164 32 6.3
NUOL_ECOLI (P33607) NADH-quinone oxidoreductase chain L (EC 1.6.... 32 6.3
>T9S1_MOUSE (Q9DBU0) Transmembrane 9 superfamily protein member 1
precursor
Length = 606
Score = 518 bits (1334), Expect = e-146
Identities = 269/573 (46%), Positives = 363/573 (62%), Gaps = 12/573 (2%)
Query: 45 YKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPYKL 104
YK GD V Y NKVGP+HNP ETY Y+ LP C PE + K LGEVL+GDR+ + Y++
Sbjct: 36 YKPGDPVILYVNKVGPYHNPQETYHYYQLPVCCPEKIRHKSLSLGEVLDGDRMAESLYEI 95
Query: 105 EFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFETDEKDVDTN 164
F + + +C L+ +V Q R A+ + Y+++ DDLPI GF+G E E +
Sbjct: 96 RFRENVEKRILCHMQLSSAQVEQLRQAIEELYYFEFVVDDLPIRGFVGYME--ESGFLPH 153
Query: 165 EATVYLFRNVHFEILYNNDRII--DVFVKN-DPNAVVDLTEDREVNVDFTYSVKWIETDI 221
+ L+ ++ F + ++ DRII +V V++ P+++ L D + + TYSV+W ET +
Sbjct: 154 SHKIGLWTHLDFHLEFHGDRIIFANVSVRDVKPHSLDGLRSDELLGLTHTYSVRWSETSV 213
Query: 222 PFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVLLLTGFLAMILMRVLKNDFVKFTPDEE 281
+ LEIHW SIINS V V LL GF+A+ILMRVL+ND ++ DEE
Sbjct: 214 EHRSDRRRGDDGGFFPRTLEIHWLSIINSMVLVFLLVGFVAVILMRVLRNDLARYNLDEE 273
Query: 282 ALD-------DQEETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVIFIFMLALVGVF 334
DQ + GWK IH DVFR+P ++ L A LG G Q L I ++AL+G+F
Sbjct: 274 TSSGGSSDDFDQGDNGWKIIHTDVFRFPPYRGLLCAVLGVGAQFLALGTGIIVMALLGMF 333
Query: 335 YPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSLFSGPLFFTFSF 394
+ GA+ +A +++YALT I+ Y ++ FY I G+ WV ++LT SLFS P F T+S
Sbjct: 334 NVHRHGAINSAAILLYALTCCISGYVSSHFYRQIGGERWVWNIILTSSLFSVPFFLTWSV 393
Query: 395 LNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNKYPRE 454
+N+V A ST ALP TI+++ +W LV PL V+GGI GKN+ S F APCRT RE
Sbjct: 394 VNSVHWANGSTQALPATTILLLLTVWLLVGFPLTVIGGIFGKNNASPFDAPCRTKNIARE 453
Query: 455 IPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQIYTIYSILFIVFIILLIVTA 514
IP PWY+ T+ M + GFLPFSAI +ELYYIFA+VWG + YT+Y ILF VF ILL V A
Sbjct: 454 IPPQPWYKSTVIHMTVGGFLPFSAISVELYYIFATVWGREQYTLYGILFFVFAILLSVGA 513
Query: 515 FGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSGFMQTSFFFGYM 574
++ALTYFQL+ ED+ WWWRS L GSTGLFI+ Y +F+Y RS+MSG +QT FFGY
Sbjct: 514 CISIALTYFQLSGEDYRWWWRSVLSVGSTGLFIFLYSVFYYARRSNMSGAVQTVEFFGYS 573
Query: 575 ACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
Y FFLMLGT+SF +SL F+R+IY +LK +
Sbjct: 574 LLTGYVFFLMLGTISFFSSLKFIRYIYVNLKMD 606
>T9S1_HUMAN (O15321) Transmembrane 9 superfamily protein member 1
precursor (hMP70)
Length = 606
Score = 511 bits (1316), Expect = e-144
Identities = 268/573 (46%), Positives = 361/573 (62%), Gaps = 12/573 (2%)
Query: 45 YKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPYKL 104
YK GD V Y NKVGP+HNP ETY Y+ LP C PE + K LGEVL+GDR+ + Y++
Sbjct: 36 YKAGDPVILYVNKVGPYHNPQETYHYYQLPVCCPEKIRHKSLSLGEVLDGDRMAESLYEI 95
Query: 105 EFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFETDEKDVDTN 164
F + + +C L+ +V Q R A+ + Y+++ DDLPI GF+G E E +
Sbjct: 96 RFRENVEKRILCHMQLSSAQVEQLRQAIEELYYFEFVVDDLPIRGFVGYME--ESGFLPH 153
Query: 165 EATVYLFRNVHFEILYNNDRII--DVFVKN-DPNAVVDLTEDREVNVDFTYSVKWIETDI 221
+ L+ ++ F + ++ DRII +V V++ P+++ L D + + TYSV+W ET +
Sbjct: 154 SHKIGLWTHLDFHLEFHGDRIIFANVSVRDVKPHSLDGLRPDEFLGLTHTYSVRWSETSV 213
Query: 222 PFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVLLLTGFLAMILMRVLKNDFVKFTPDEE 281
+ LEIHW SIINS V V LL GF+A+ILMRVL+ND ++ DEE
Sbjct: 214 ERRSDRRRGDDGGFFPRTLEIHWLSIINSMVLVFLLVGFVAVILMRVLRNDLARYNLDEE 273
Query: 282 ALD-------DQEETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVIFIFMLALVGVF 334
DQ + GWK IH DVFR+P ++ L A LG G Q L I ++AL+G+F
Sbjct: 274 TTSAGSGDDFDQGDNGWKIIHTDVFRFPPYRGLLCAVLGVGAQFLALGTGIIVMALLGMF 333
Query: 335 YPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSLFSGPLFFTFSF 394
+ GA+ +A +++YALT I+ Y ++ FY I G+ WV ++LT SLFS P F T+S
Sbjct: 334 NVHRHGAINSAAILLYALTCCISGYVSSHFYRQIGGERWVWNIILTTSLFSVPFFLTWSV 393
Query: 395 LNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNKYPRE 454
+N+V A ST ALP TI+++ +W LV PL V+GGI GKN+ S F APCRT RE
Sbjct: 394 VNSVHWANGSTQALPATTILLLLTVWLLVGFPLTVIGGIFGKNNASPFDAPCRTKNIARE 453
Query: 455 IPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQIYTIYSILFIVFIILLIVTA 514
I PWY+ T M + GFLPFSAI +ELYYIFA+VWG + YT+Y ILF VF ILL V A
Sbjct: 454 INPQPWYKSTDIHMTVGGFLPFSAISVELYYIFATVWGREQYTLYGILFFVFAILLSVGA 513
Query: 515 FGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSGFMQTSFFFGYM 574
++ALTYFQL+ ED+ WWWRS L GSTGLFI+ Y +F+Y RS+MSG +QT FFGY
Sbjct: 514 SISIALTYFQLSGEDYRWWWRSVLSVGSTGLFIFLYSVFYYARRSNMSGAVQTVEFFGYS 573
Query: 575 ACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
Y FFLMLGT+SF +SL F+R+IY +LK +
Sbjct: 574 LLTGYVFFLMLGTISFFSSLKFIRYIYVNLKMD 606
>T9S3_MOUSE (Q9ET30) Transmembrane 9 superfamily protein member 3
precursor
Length = 587
Score = 417 bits (1072), Expect = e-116
Identities = 212/577 (36%), Positives = 332/577 (56%), Gaps = 22/577 (3%)
Query: 38 SDASDHRYKEGDFVPFYANKVGPFHNPSETYRYFDLPFC--SPENVEEKKEDLGEVLNGD 95
SD +H Y++ + V + N VGP+HN ETY+YF LPFC S +++ E LGE L G
Sbjct: 26 SDEHEHTYQDKEEVVLWMNTVGPYHNRQETYKYFSLPFCVGSKKSISHYHETLGEALQGV 85
Query: 96 RLVVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFE 155
L + ++F D P + C+ L +++ F +A+ Y+YQMY DDLPIWG +G
Sbjct: 86 ELEFSGLDIKFKDDVMPGTYCEIDLDKEKRDAFVYAIKNHYWYQMYIDDLPIWGIVG--- 142
Query: 156 TDEKDVDTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVVDLTEDREVNVDFTYSVK 215
+ D N YL+ EI +N +RI+DV + ++ V L + ++ + +YSVK
Sbjct: 143 ----EADENGEDYYLWTYKKLEIGFNGNRIVDVNLTSEGK--VKLVPNTKIQM--SYSVK 194
Query: 216 WIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVLLLTGFLAMILMRVLKNDFVK 275
W ++D+ FE R +KY S H IHWFSI NS + V+ L G ++MILMR L+ D+ +
Sbjct: 195 WKKSDVKFEDRFDKYLDPSFFQHR--IHWFSIFNSFMMVIFLVGLVSMILMRTLRKDYAR 252
Query: 276 FTPDEEALDDQE-----ETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVIFIFMLAL 330
++ +EE +DD + E GWK +HGDVFR +F++ +G+G Q+F + + + ++A+
Sbjct: 253 YSKEEE-MDDMDRDLGDEYGWKQVHGDVFRPSSHPLIFSSLIGSGCQIFAVSLIVIIVAM 311
Query: 331 VGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSLFSGPLFF 390
+ Y RG++ + + +YA TS + Y S Y G+ W+K + + L +
Sbjct: 312 IEDLYT-ERGSMLSTAIFVYAATSPVNGYFGGSLYARQGGRRWIKQMFIGAFLIPAMVCG 370
Query: 391 TFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNK 450
T F+N +A+ Y+++ A+PFGT+V + I V PL ++G I G+N + PCR N
Sbjct: 371 TAFFINFIAIYYHASRAIPFGTMVAVCCICFFVILPLNLVGTILGRNLSGQPNFPCRVNA 430
Query: 451 YPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQIYTIYSILFIVFIILL 510
PR IP+ W+ + + + G LPF +IFIE+Y+IF S W ++IY +Y + +V +IL
Sbjct: 431 VPRPIPEKKWFMEPAVIVCLGGILPFGSIFIEMYFIFTSFWAYKIYYVYGFMMLVLVILC 490
Query: 511 IVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSGFMQTSFF 570
IVT + TYF L AED+ W W SFL ST +++Y Y ++Y ++ M G QTSF+
Sbjct: 491 IVTVCVTIVCTYFLLNAEDYRWQWTSFLSAASTAIYVYMYSFYYYFFKTKMYGLFQTSFY 550
Query: 571 FGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
FGYMA +M G + + + FVR IY ++K +
Sbjct: 551 FGYMAVFSTALGIMCGAIGYMGTSAFVRKIYTNVKID 587
>T9S3_HUMAN (Q9HD45) Transmembrane 9 superfamily protein member 3
precursor (SM-11044 binding protein) (EP70-P-iso)
Length = 589
Score = 416 bits (1070), Expect = e-116
Identities = 211/577 (36%), Positives = 332/577 (56%), Gaps = 22/577 (3%)
Query: 38 SDASDHRYKEGDFVPFYANKVGPFHNPSETYRYFDLPFC--SPENVEEKKEDLGEVLNGD 95
+D +H Y++ + V + N VGP+HN ETY+YF LPFC S +++ E LGE L G
Sbjct: 28 ADEHEHTYQDKEEVVLWMNTVGPYHNRQETYKYFSLPFCVGSKKSISHYHETLGEALQGV 87
Query: 96 RLVVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFE 155
L + ++F D P + C+ L +++ F +A+ Y+YQMY DDLPIWG +G
Sbjct: 88 ELEFSGLDIKFKDDVMPATYCEIDLDKEKRDAFVYAIKNHYWYQMYIDDLPIWGIVG--- 144
Query: 156 TDEKDVDTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVVDLTEDREVNVDFTYSVK 215
+ D N YL+ EI +N +RI+DV + ++ V L + ++ + +YSVK
Sbjct: 145 ----EADENGEDYYLWTYKKLEIGFNGNRIVDVNLTSEGK--VKLVPNTKIQM--SYSVK 196
Query: 216 WIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVLLLTGFLAMILMRVLKNDFVK 275
W ++D+ FE R +KY S H IHWFSI NS + V+ L G ++MILMR L+ D+ +
Sbjct: 197 WKKSDVKFEDRFDKYLDPSFFQHR--IHWFSIFNSFMMVIFLVGLVSMILMRTLRKDYAR 254
Query: 276 FTPDEEALDDQE-----ETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVIFIFMLAL 330
++ +EE +DD + E GWK +HGDVFR +F++ +G+G Q+F + + + ++A+
Sbjct: 255 YSKEEE-MDDMDRDLGDEYGWKQVHGDVFRPSSHPLIFSSLIGSGCQIFAVSLIVIIVAM 313
Query: 331 VGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSLFSGPLFF 390
+ Y RG++ + + +YA TS + Y S Y G+ W+K + + L +
Sbjct: 314 IEDLYT-ERGSMLSTAIFVYAATSPVNGYFGGSLYARQGGRRWIKQMFIGAFLIPAMVCG 372
Query: 391 TFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNK 450
T F+N +A+ Y+++ A+PFGT+V + I V PL ++G I G+N + PCR N
Sbjct: 373 TAFFINFIAIYYHASRAIPFGTMVAVCCICFFVILPLNLVGTILGRNLSGQPNFPCRVNA 432
Query: 451 YPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQIYTIYSILFIVFIILL 510
PR IP+ W+ + + + G LPF +IFIE+Y+IF S W ++IY +Y + +V +IL
Sbjct: 433 VPRPIPEKKWFMEPAVIVCLGGILPFGSIFIEMYFIFTSFWAYKIYYVYGFMMLVLVILC 492
Query: 511 IVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSGFMQTSFF 570
IVT + TYF L AED+ W W SFL ST +++Y Y ++Y ++ M G QTSF+
Sbjct: 493 IVTVCVTIVCTYFLLNAEDYRWQWTSFLSAASTAIYVYMYSFYYYFFKTKMYGLFQTSFY 552
Query: 571 FGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
FGYMA +M G + + + FVR IY ++K +
Sbjct: 553 FGYMAVFSTALGIMCGAIGYMGTSAFVRKIYTNVKID 589
>T9S4_HUMAN (Q92544) Transmembrane 9 superfamily protein member 4
Length = 625
Score = 367 bits (942), Expect = e-101
Identities = 194/592 (32%), Positives = 318/592 (52%), Gaps = 57/592 (9%)
Query: 68 YRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPYKLEFLIDKKPESICQK-----MLTR 122
Y Y+ LPFC P + K E+LGEVL GDR+V P+++ +KK E +C + LT
Sbjct: 39 YEYYSLPFCQPSKITYKAENLGEVLRGDRIVNTPFQVLMNSEKKCEVLCSQSNKPVTLTV 98
Query: 123 KEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRF-------ETDEKDV-------------- 161
++ + +DY+ + D+LP+ L + + EKDV
Sbjct: 99 EQSRLVAERITEDYYVHLIADNLPVATRLELYSNRDSDDKKKEKDVQFEHGYRLGFTDVN 158
Query: 162 -----------------DTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVV------ 198
D E + +R V FE++ + R+ D+ + +
Sbjct: 159 KIYLHNHLSFILYYHREDMEEDQEHTYRVVRFEVIPQSIRLEDLKADEKSSCTLPEGTNS 218
Query: 199 ---DLTEDREVNVDFTYSVKWIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVL 255
++ +E + FTYSV W E+DI + R + Y S ++IHWFSIINS V V
Sbjct: 219 SPQEIDPTKENQLYFTYSVHWEESDIKWASRWDTYLTMSD----VQIHWFSIINSVVVVF 274
Query: 256 LLTGFLAMILMRVLKNDFVKFTPDEEALDDQEETGWKYIHGDVFRYPRFKSLFAAALGTG 315
L+G L+MI++R L+ D + +++ D EE+GWK +HGDVFR P++ + ++ LG+G
Sbjct: 275 FLSGILSMIIIRTLRKDIANYNKEDDIEDTMEESGWKLVHGDVFRPPQYPMILSSLLGSG 334
Query: 316 TQLFTLVIFIFMLALVGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVK 375
QLF +++ + +A++G+ P +RGAL T ++ +SA Y ++G W K
Sbjct: 335 IQLFCMILIVIFVAMLGMLSPSSRGALMTTACFLFMFMGVFGGFSAGRLYRTLKGHRWKK 394
Query: 376 ILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAG 435
T +L+ G +F LN +S+ A+PF T+V + +W ++ PL+ LG G
Sbjct: 395 GAFCTATLYPGVVFGICFVLNCFIWGKHSSGAVPFPTMVALLCMWFGISLPLVYLGYYFG 454
Query: 436 KNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQI 495
Q + P RTN+ PR+IP+ WY + MAG LPF A+FIEL++IF+++W +Q
Sbjct: 455 FRKQ-PYDNPVRTNQIPRQIPEQRWYMNRFVGILMAGILPFGAMFIELFFIFSAIWENQF 513
Query: 496 YTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFY 555
Y ++ LF+VFIIL++ + ++ + YFQL AED+ WWWR+FL G + ++ Y +F++
Sbjct: 514 YYLFGFLFLVFIILVVSCSQISIVMVYFQLCAEDYRWWWRNFLVSGGSAFYVLVYAIFYF 573
Query: 556 HARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
+ D+ F+ + +FGY A + F+L+ GT+ F A+ +FVR IY ++K +
Sbjct: 574 VNKLDIVEFIPSLLYFGYTALMVLSFWLLTGTIGFYAAYMFVRKIYAAVKID 625
>T9S2_HUMAN (Q99805) Transmembrane 9 superfamily protein member 2
precursor (p76)
Length = 663
Score = 323 bits (828), Expect = 8e-88
Identities = 197/595 (33%), Positives = 305/595 (51%), Gaps = 62/595 (10%)
Query: 68 YRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPYKLEFLIDKKPESIC------QKMLT 121
Y Y FC + E+LG+VL G+R+ +PYK F + + +C +K
Sbjct: 76 YEYTAFDFCQASEGKRPSENLGQVLFGERIEPSPYKFTFNKKETCKLVCTKTYHTEKAED 135
Query: 122 RKEVAQFRHAVLKDYFYQMYYDDLPI-W-------------GF-LGRFETDE---KDV-- 161
++++ + ++L +Y + D++P+ W GF +G + TD+ KD
Sbjct: 136 KQKLEFLKKSMLLNYQHHWIVDNMPVTWCYDVEDGQRFCNPGFPIGCYITDKGHAKDACV 195
Query: 162 ---DTNEA-TVYLFRNVHFEILYNNDRIID--------VFVKNDPNAVVDLTEDR----- 204
D +E T Y+F +V +I Y+ +++ V K +P + D+
Sbjct: 196 ISSDFHERDTFYIFNHVDIKIYYH---VVETGSMGARLVAAKLEPKSFKHTHIDKPDCSG 252
Query: 205 -----------EVNVDFTYSVKWIETD-IPFEKRLEKYSQTSSLSHHLEIHWFSIINSCV 252
E+ + +TYSV + E D I + R + ++ H I WFSI+NS V
Sbjct: 253 PPMDISNKASGEIKIAYTYSVSFEEDDKIRWASRWDYILESMP---HTHIQWFSIMNSLV 309
Query: 253 TVLLLTGFLAMILMRVLKNDFVKFTPDEEALDDQEETGWKYIHGDVFRYPRFKSLFAAAL 312
VL L+G +AMI++R L D ++ + D QEE GWK +HGD+FR PR L + L
Sbjct: 310 IVLFLSGMVAMIMLRTLHKDIARYNQMDSTEDAQEEFGWKLVHGDIFRPPRKGMLLSVFL 369
Query: 313 GTGTQLFTLVIFIFMLALVGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKN 372
G+GTQ+ + A +G P NRGAL T VV++ L A Y AA FY G+
Sbjct: 370 GSGTQILIMTFVTLFFACLGFLSPANRGALMTCAVVLWVLLGTPAGYVAARFYKSFGGEK 429
Query: 373 WVKILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGG 432
W ++LT L G +F F +N + S+AA+PFGT+V I +W ++ PL +G
Sbjct: 430 WKTNVLLTSFLCPGIVFADFFIMNLILWGEGSSAAIPFGTLVAILALWFCISVPLTFIGA 489
Query: 433 IAGKNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWG 492
G ++ + P RTN+ PR+IP+ +Y K L + M G LPF IFI+L++I S+W
Sbjct: 490 YFG-FKKNAIEHPVRTNQIPRQIPEQSFYTKPLPGIIMGGILPFGCIFIQLFFILNSIWS 548
Query: 493 HQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCM 552
HQ+Y ++ LF+VFIIL+I + + L YF L AED+ W WRSFL G T ++ Y +
Sbjct: 549 HQMYYMFGFLFLVFIILVITCSEATILLCYFHLCAEDYHWQWRSFLTSGFTAVYFLIYAV 608
Query: 553 FFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
++ ++ ++G T +FGY + FFL GT+ F A FV IY +K +
Sbjct: 609 HYFFSKLQITGTASTILYFGYTMIMVLIFFLFTGTIGFFACFWFVTKIYSVVKVD 663
>T9S2_MOUSE (P58021) Transmembrane 9 superfamily protein member 2
precursor
Length = 662
Score = 320 bits (819), Expect = 9e-87
Identities = 194/597 (32%), Positives = 299/597 (49%), Gaps = 66/597 (11%)
Query: 68 YRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPYKLEFLIDKKPESICQKMLTRKEVAQ 127
Y Y FC + E+LG+VL G+R+ +PYK F + + +C K ++
Sbjct: 75 YEYTAFDFCQASEGKRPSENLGQVLFGERIEPSPYKFTFNKKETCKLVCTKTYNTEKAED 134
Query: 128 ------FRHAVLKDYFYQMYYDDLPI-W-------------GF-LGRFETDE---KDVDT 163
+ ++L +Y + D++P+ W GF +G + TD+ KD
Sbjct: 135 KQKLDFLKKSMLLNYQHHWIVDNMPVTWCYEVEDSQKFCNPGFPIGCYITDKGHAKDACV 194
Query: 164 NEA------TVYLFRNVHFEILYNNDRIID--------VFVKNDPNAVVDLTEDR----- 204
+ T Y+F +V +I Y+ +++ V K +P + D+
Sbjct: 195 ISSEFHERDTFYIFNHVDIKIYYH---VVETGSMGARLVAAKLEPKSFKHTHIDKPDCSG 251
Query: 205 -----------EVNVDFTYSVKWIETDIPFEKRLEKYSQTSSLSH---HLEIHWFSIINS 250
E+ + +TYS+ + E EK + S+ + H I WFSI+NS
Sbjct: 252 PAMDISNKASGEIKIAYTYSISFEE-----EKNIRWASRWDYILESMPHTHIQWFSIMNS 306
Query: 251 CVTVLLLTGFLAMILMRVLKNDFVKFTPDEEALDDQEETGWKYIHGDVFRYPRFKSLFAA 310
V VL L+G +AMI++R L D ++ + D QEE GWK +HGD+FR PR L +
Sbjct: 307 LVIVLFLSGMVAMIMLRTLHKDIARYNQMDSTEDAQEEFGWKLVHGDIFRPPRKGMLLSV 366
Query: 311 ALGTGTQLFTLVIFIFMLALVGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEG 370
LG+GTQ+ + A +G P NRGAL T VV++ L A Y AA FY G
Sbjct: 367 FLGSGTQILIMTFVTLFFACLGFLSPANRGALMTCAVVLWVLLGTPAGYVAARFYKSFGG 426
Query: 371 KNWVKILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVL 430
+ W ++LT L G +F F +N + S+AA+PFGT+V I +W ++ PL +
Sbjct: 427 EKWKTNVLLTSFLCPGIVFADFFIMNLILWGEGSSAAIPFGTLVAILALWFCISVPLTFI 486
Query: 431 GGIAGKNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASV 490
G G ++ + P RTN+ PR+IP+ +Y K L + M G LPF IFI+L++I S+
Sbjct: 487 GAYFG-FKKNAIEHPVRTNQIPRQIPEQSFYTKPLPGIIMGGILPFGCIFIQLFFILNSI 545
Query: 491 WGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGY 550
W HQ+Y ++ LF+VFIIL+I + + L YF L AED+ W WRSFL G T ++ Y
Sbjct: 546 WSHQMYYMFGFLFLVFIILVITCSEATILLCYFHLCAEDYHWQWRSFLTSGFTAVYFLIY 605
Query: 551 CMFFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
+ ++ ++ ++G T +FGY + FFL GT+ F A FV IY +K +
Sbjct: 606 AIHYFFSKLQITGTASTILYFGYTMIMVLIFFLFTGTIGFFACFWFVTKIYSVVKVD 662
>EM70_YEAST (P32802) Endosomal P24A protein precursor (70 kDa
endomembrane protein) (Pheromone alpha-factor
transporter) (Acidic 24 kDa late endocytic intermediate
component)
Length = 667
Score = 289 bits (740), Expect = 1e-77
Identities = 192/644 (29%), Positives = 307/644 (46%), Gaps = 90/644 (13%)
Query: 45 YKEGDFVPFYANKVGPFHNP-------------------SETYRYFDLPFCSPENVEEKK 85
Y+E D +P N + P N S Y Y FC PE VE++
Sbjct: 33 YRENDNIPLLVNHLTPSMNYQHKDEDGNNVSGDKENFLYSYDYYYNRFHFCQPEKVEKQP 92
Query: 86 EDLGEVLNGDRLVVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMY---- 141
E LG V+ GDR+ +P++L L +K+ ES+C+ ++ + A+F + ++K+ F+Q +
Sbjct: 93 ESLGSVIFGDRIYNSPFQLNMLQEKECESLCKTVIPGDD-AKFINKLIKNGFFQNWLIDG 151
Query: 142 -------YDDLPIWGFLGR---------------------FETDEKDV------DTNEAT 167
YD F G ET +KDV D N
Sbjct: 152 LPAAREVYDGRTKTSFYGAGFNLGFVQVTQGTDIEATPKGAETTDKDVELETRNDRNMVK 211
Query: 168 VY---LFRNVHFEILYN-------NDRIIDVFVK------------NDPNAVVDLTEDRE 205
Y F N HF+I+ N R++ V V+ + + L E +
Sbjct: 212 TYELPYFAN-HFDIMIEYHDRGEGNYRVVGVIVEPVSIKRSSPGTCETTGSPLMLDEGND 270
Query: 206 VNVDFTYSVKWIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTVLLLTGFLAMIL 265
V FTYSVK+ E+ + R +KY S I WFS+IN + V+LL+ + L
Sbjct: 271 NEVYFTYSVKFNESATSWATRWDKYLHVYDPS----IQWFSLINFSLVVVLLSSVVIHSL 326
Query: 266 MRVLKNDFVKFTPDEEALDD--QEETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVI 323
+R LK+DF ++ +E LDD QE++GWK HGDVFR P + +G+G QLF +V
Sbjct: 327 LRALKSDFARY--NELNLDDDFQEDSGWKLNHGDVFRSPSQSLTLSILVGSGVQLFLMVT 384
Query: 324 FIFMLALVGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSL 383
A +G P +RG+L T + ++YAL + SY++ Y G W L+LT L
Sbjct: 385 CSIFFAALGFLSPSSRGSLATVMFILYALFGFVGSYTSMGIYKFFNGPYWKANLILTPLL 444
Query: 384 FSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQ 443
G + LN + +S+ +P T+ + +W L + PL G + + +
Sbjct: 445 VPGAILLIIIALNFFLMFVHSSGVIPASTLFFMVFLWFLFSIPLSFAGSLIARKRCHWDE 504
Query: 444 APCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGHQIYTIYSILF 503
P +TN+ R+IP PWY KT+ +AG PF +I +ELY+I+ S+W ++I+ ++ LF
Sbjct: 505 HPTKTNQIARQIPFQPWYLKTIPATLIAGIFPFGSIAVELYFIYTSLWFNKIFYMFGFLF 564
Query: 504 IVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSG 563
F++L + ++ + +TY L E+ +W WR F+ GG+ G +Y + + + G
Sbjct: 565 FSFLLLTLTSSLVTILITYHSLCLENWKWQWRGFIIGGA-GCALYVFIHSILFTKFKLGG 623
Query: 564 FMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
F + GY + I L+ G++ F +S++FVR IY S+K +
Sbjct: 624 FTTIVLYVGYSSVISLLCCLVTGSIGFISSMLFVRKIYSSIKVD 667
>YEU3_YEAST (P40071) Hypothetical 81.5 kDa protein in USS1-BEB1
intergenic region precursor
Length = 706
Score = 131 bits (329), Expect = 6e-30
Identities = 151/656 (23%), Positives = 271/656 (41%), Gaps = 111/656 (16%)
Query: 45 YKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSPENVEEKKE--DLGEVLNGDRLVVAPY 102
Y++GD + NKV Y Y+DLPF P + +K L E++ GDR + Y
Sbjct: 57 YRKGDPLELIVNKVES-DLTQLPYAYYDLPFTCPPTMHKKPLHLSLNEIIRGDRKWESDY 115
Query: 103 KLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFETDEKD-- 160
KL+F D E++C + T++ + V + Y Q DD TD K
Sbjct: 116 KLKFGEDNPCETLCARKTTKEGMQTLDKLVREGYVVQWLIDDELPAATTFISTTDHKKYY 175
Query: 161 --------VDTNEATVYLFRNVHFEILYN---NDRIIDVFVKNDPNAVVDL--------- 200
+D + YL +V I ++ ND+ V + P +V D
Sbjct: 176 ASGFPLGFIDPDTDKTYLHNHVMLVIRFHASDNDKNTIVGFEVYPRSVSDYHCPGASKNY 235
Query: 201 ---------TEDREVNVDFTYSVKWIET-DIPFEKRLEKYSQTSSLSHH--LEIHWFSII 248
E+ + FTYSV W E ++ + R + + LS ++ HW S+
Sbjct: 236 EQYEIVIPEDENELTYLPFTYSVYWREEFEVDWNHRWDYFLNAGELSDEQSIQFHWMSLA 295
Query: 249 NSCVTVLLLTGFLAMILMRVLKND-----FVKFTPDEEALDDQEETGWKYIHGDVFRYPR 303
NS VL ++ +I +RV+ D K+ + E ++ +++ D +Y +
Sbjct: 296 NSVGIVLSISFITLIIYVRVMYTDKSNSKSPKYMINIEGIETEDDL-------DDDKYGK 348
Query: 304 FKSLFAAA---LGTGT-QLFTLVIFIFMLALVGVFYPYN-----------------RGAL 342
+ S++ A + G LF L + I +++ GV + + R ++
Sbjct: 349 Y-SVYTVAKDWIQNGRPNLFGLKVLILLVSF-GVQFLFTIIGSLTISCSMNKLHNVRNSV 406
Query: 343 FTALVVIYALTSGIASY-------------SAASFY----YMIEGKNWVKIL-VLTGSLF 384
T ++ + L + +AS+ + A++ Y+ + K + I +L GS
Sbjct: 407 LTMAILFFVLGAFMASFVGTRLSMVTKTKRTKANYLDDNRYLKDYKKFSPIFTILCGSSL 466
Query: 385 SGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVLGGIAGKN------- 437
G + + LN++ A++ST+ALPF TIV I+ +V PL + GGI N
Sbjct: 467 PGIVMVSTFLLNSIVWAHDSTSALPFKTIVFFMSIYFIVCIPLSLFGGIVANNIPLPQYW 526
Query: 438 ----SQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFASVWGH 493
++ E + P+ K + + + G P I++E+ Y++ S+W
Sbjct: 527 LSGITKDESNSDGNGLFVPKSRAK--FNPLVYCGIYLCGIFPLLVIYVEMQYVYKSLWLE 584
Query: 494 Q--IYTIYSILFIVFIILLIVTAFGNVALTY------FQLAAEDHEWWWRSFLCGGSTGL 545
+ Y Y LF+ I+L ++T ++ +Y F+ + W W+ F G S G+
Sbjct: 585 KTTFYYFYGFLFLSIILLCVLTMEISIIGSYLLMRFCFEDKVVRNNWRWKCFEMGFSGGV 644
Query: 546 FIYGYCMFFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIY 601
++ Y +++ A ++ GF Y L LG +S+ + F+ IY
Sbjct: 645 YMELYSLYYIFAVLNIHGFSSILISICYSLIFNVMCSLGLGALSYLTASWFINKIY 700
>SGAT_MYCPN (P75291) Putative transport protein sgaT homolog
Length = 660
Score = 36.2 bits (82), Expect = 0.26
Identities = 43/156 (27%), Positives = 74/156 (46%), Gaps = 12/156 (7%)
Query: 340 GALFTAL-VVIYALTSGIASYSAASFYYMIEGKNWVKILVLTG--SLFSGPLFFTFSFLN 396
GAL T + ++ ++ SG+ +A + I+G +++L SL S FFT SFLN
Sbjct: 99 GALKTVIGFILLSIGSGVLVSTARPVFDTIKGLGGTGVVLLDPYFSLASANDFFTNSFLN 158
Query: 397 TVAVAYNSTAALPFGTIVVIFL---IWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNKYPR 453
V+ + + L + +IF+ WT T+ ++V G + + + A T Y
Sbjct: 159 NDYVSLIAFSLLVGFIVNIIFVGLKRWT-NTNSIMVTGHVMLQQA-----AVVTTLFYIV 212
Query: 454 EIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFAS 489
++P +A A AG + S IF+ +Y+ AS
Sbjct: 213 LFRQIPLLGTGIAYGAQAGLVIISGIFLGVYWSTAS 248
>R1AB_CVH22 (Q05002) Replicase polyprotein 1ab (pp1ab) (ORF1ab
polyprotein) [Includes: Replicase polyprotein 1a (pp1a)
(ORF1a)] [Contains: p9; p87; p195 (EC 3.4.24.-)
(Papain-like proteinases 1/2) (PL1-PRO/PL2-PRO); Peptide
HD2; 3C-like proteinase (EC 3.4
Length = 6758
Score = 34.7 bits (78), Expect = 0.74
Identities = 22/69 (31%), Positives = 32/69 (45%), Gaps = 10/69 (14%)
Query: 128 FRHAVLKDYFYQMYYDDLPIWGFLGRFETDEKDVDTNEATVYLFRNVHF----EILYNND 183
F+HA+ DY Y Y D+ WG++G T+ + A + RN H I+
Sbjct: 5818 FKHALGCDYVYNPYVIDIQQWGYVGSLSTN------HHAICNVHRNEHVASGDAIMTRCL 5871
Query: 184 RIIDVFVKN 192
+ D FVKN
Sbjct: 5872 AVYDCFVKN 5880
>NU6M_CYACA (P48925) NADH-ubiquinone oxidoreductase chain 6 (EC
1.6.5.3)
Length = 201
Score = 34.3 bits (77), Expect = 0.97
Identities = 19/58 (32%), Positives = 33/58 (56%), Gaps = 2/58 (3%)
Query: 469 AMAGFLPFSAIFIELYYIFAS--VWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQ 524
A + FL I+++ Y + + + G+ +YT YS LFI+ +L+V G + LT +Q
Sbjct: 116 ASSYFLERKRIWVQELYSYTNLQILGNVLYTSYSYLFILSGFVLLVAILGAIILTLYQ 173
>O4S2_HUMAN (Q8NH73) Olfactory receptor 4S2
Length = 311
Score = 33.9 bits (76), Expect = 1.3
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 2/47 (4%)
Query: 363 SFYYMIE--GKNWVKILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAA 407
SF+Y+I G + + V +LF P++F SFL+ V + Y+S A
Sbjct: 30 SFFYIIILLGNLLIMLTVCLSNLFKSPMYFFLSFLSFVDICYSSVTA 76
>HMDH_SCHMA (P16237) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 948
Score = 33.9 bits (76), Expect = 1.3
Identities = 27/100 (27%), Positives = 44/100 (44%), Gaps = 9/100 (9%)
Query: 316 TQLFTLVIFIFMLALVG--VFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIE---- 369
T +TL++F F L + +F PY A+F ++ L S ++ + YY++E
Sbjct: 91 TLFYTLILFTFALCSLSSVLFVPYTSFAIFLLSTSVFLLFSDLSVFFIVLEYYLLEIELV 150
Query: 370 GKNWVKILVLTGSLFSGPLFFTF---SFLNTVAVAYNSTA 406
K L LFS LF FL T + ++T+
Sbjct: 151 NYEHAKRHCLLSHLFSNQLFVDHMLGMFLKTSLFSISTTS 190
>Y765_STAAM (P67108) Hypothetical UPF0042 protein SAV0765
Length = 303
Score = 32.7 bits (73), Expect = 2.8
Identities = 24/73 (32%), Positives = 38/73 (51%), Gaps = 4/73 (5%)
Query: 154 FETDEKDVDTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVVDLTEDREVNVD-FTY 212
+E +E + T T + F++ I + D + DV +P VVDL ++ D + Y
Sbjct: 167 YEDEEFETFTINVTSFGFKH---GIQMDADLVFDVRFLPNPYYVVDLRPLTGLDKDVYNY 223
Query: 213 SVKWIETDIPFEK 225
+KW ET+I FEK
Sbjct: 224 VMKWKETEIFFEK 236
>Y727_STAAW (P67110) Hypothetical UPF0042 protein MW0727
Length = 303
Score = 32.7 bits (73), Expect = 2.8
Identities = 24/73 (32%), Positives = 38/73 (51%), Gaps = 4/73 (5%)
Query: 154 FETDEKDVDTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVVDLTEDREVNVD-FTY 212
+E +E + T T + F++ I + D + DV +P VVDL ++ D + Y
Sbjct: 167 YEDEEFETFTINVTSFGFKH---GIQMDADLVFDVRFLPNPYYVVDLRPLTGLDKDVYNY 223
Query: 213 SVKWIETDIPFEK 225
+KW ET+I FEK
Sbjct: 224 VMKWKETEIFFEK 236
>Y720_STAAN (P67109) Hypothetical UPF0042 protein SA0720
Length = 303
Score = 32.7 bits (73), Expect = 2.8
Identities = 24/73 (32%), Positives = 38/73 (51%), Gaps = 4/73 (5%)
Query: 154 FETDEKDVDTNEATVYLFRNVHFEILYNNDRIIDVFVKNDPNAVVDLTEDREVNVD-FTY 212
+E +E + T T + F++ I + D + DV +P VVDL ++ D + Y
Sbjct: 167 YEDEEFETFTINVTSFGFKH---GIQMDADLVFDVRFLPNPYYVVDLRPLTGLDKDVYNY 223
Query: 213 SVKWIETDIPFEK 225
+KW ET+I FEK
Sbjct: 224 VMKWKETEIFFEK 236
>Y275_HAEIN (P43975) Hypothetical protein HI0275
Length = 551
Score = 32.3 bits (72), Expect = 3.7
Identities = 33/132 (25%), Positives = 57/132 (43%), Gaps = 22/132 (16%)
Query: 316 TQLFTLVIFIFMLA-LVGVFYPYNRG-----ALFTALVVIYALTSGIASYSA----ASFY 365
T F + +F F+ ++ V + R LF L++ + L I Y + F+
Sbjct: 26 TYFFAITLFTFLFGGMLMVSSQWQRALNFSSVLFVVLMLFHRLK--IHYYKQPLLISDFF 83
Query: 366 YMIEGKNWVKILVLTGSLFS--------GPLFFTFSFLNTVAVAYNSTAALPFGTIVVIF 417
+++ +NW ++ G+LF G F F+ + ++ V NS AL F IV
Sbjct: 84 LVVDWRNWETLIHYKGALFGVIGLLALLGYAIFGFNDVESLGVLGNSIGALLF--IVSFS 141
Query: 418 LIWTLVTSPLLV 429
L+W +P V
Sbjct: 142 LMWHYSKNPSAV 153
>Y164_METJA (Q57628) Hypothetical protein MJ0164
Length = 395
Score = 31.6 bits (70), Expect = 6.3
Identities = 28/113 (24%), Positives = 48/113 (41%), Gaps = 15/113 (13%)
Query: 151 LGRFETDEKDVDT--NEATVYLFRNVHFEIL----YNNDRII-------DVFVKNDPNAV 197
+G + TD + + N+ V N +++ YN D+I ++F D N V
Sbjct: 57 VGCYYTDSESTEVFGNKEKVIELLNYENKVIDIYKYNKDKINLMKWLYPEIFACKDTNKV 116
Query: 198 VDLTEDREVNVDFTYSVKWIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINS 250
+ ED D I+ DIP + +E + T +LE + + IIN+
Sbjct: 117 SEKNEDMSEKRDIVEKYLNIKLDIPLDNLIE--ANTKDFEKYLEDNKYIIINA 167
>NUOL_ECOLI (P33607) NADH-quinone oxidoreductase chain L (EC
1.6.99.5) (NADH dehydrogenase I, chain L) (NDH-1, chain
L) (NUO12)
Length = 613
Score = 31.6 bits (70), Expect = 6.3
Identities = 53/218 (24%), Positives = 79/218 (35%), Gaps = 59/218 (27%)
Query: 334 FYPYNRGALFTALVVIYALTS------------GIASYSAASFYYMI--EGKNWVKILVL 379
F+ Y LF A +V+ L G+ SY FYY G +K V+
Sbjct: 116 FFAYTN--LFIASMVVLVLADNLLLMYLGWEGVGLCSYLLIGFYYTDPKNGAAAMKAFVV 173
Query: 380 TGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVI----------FLIWTLVTSPLLV 429
T G +F F+ + YN L F +V + L+W + L++
Sbjct: 174 TRV---GDVFLAFALF----ILYNELGTLNFREMVELAPAHFADGNNMLMW----ATLML 222
Query: 430 LGGIAGKNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYIFAS 489
LGG GK++Q +P W AMAG P SA+ + A
Sbjct: 223 LGGAVGKSAQ---------------LPLQTWLAD-----AMAGPTPVSALIHAATMVTAG 262
Query: 490 VWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAA 527
V + I + + + +L +V G V L AA
Sbjct: 263 V--YLIARTHGLFLMTPEVLHLVGIVGAVTLLLAGFAA 298
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.329 0.143 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,225,156
Number of Sequences: 164201
Number of extensions: 3123543
Number of successful extensions: 10110
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 10061
Number of HSP's gapped (non-prelim): 34
length of query: 607
length of database: 59,974,054
effective HSP length: 116
effective length of query: 491
effective length of database: 40,926,738
effective search space: 20095028358
effective search space used: 20095028358
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149209.11


