

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.5 + phase: 0 /pseudo
(738 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
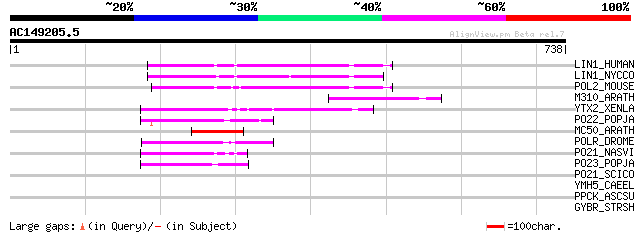
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 97 2e-19
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 96 3e-19
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 95 8e-19
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 86 5e-16
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 79 4e-14
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 65 6e-10
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 59 5e-08
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 56 4e-07
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 55 5e-07
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 50 3e-05
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 39 0.049
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 36 0.42
PPCK_ASCSU (Q05893) Phosphoenolpyruvate carboxykinase [GTP] (EC ... 33 3.5
GYBR_STRSH (P50074) DNA gyrase subunit B, novobiocin-resistant (... 32 4.6
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 97.1 bits (240), Expect = 2e-19
Identities = 81/330 (24%), Positives = 150/330 (44%), Gaps = 24/330 (7%)
Query: 184 VVDEAKKTKK-ELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKECISTTTASI 242
++ +TK ++ +D EKA+D + ++ + K+ + K I+ TA+I
Sbjct: 581 IIQHINRTKDTNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANI 640
Query: 243 LVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVV 302
++NG + L G RQG PLSP L I L VL +++ + KG +G V
Sbjct: 641 ILNGQKLEAPPLKTGTRQGCPLSPLLPNIV---LEVLARAIRQEKEIKGIQLGKEE---V 694
Query: 303 SHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAAS 362
FADD ++ E + + L +++ F+++SG K+N KS + S
Sbjct: 695 KLSLFADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMS 754
Query: 363 VLNCKVGKILFLYLGLSIGGDPRKL--AFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLKS 420
L + YLG+ + D + L ++P+++ IK + W++ S+ GR+ ++K
Sbjct: 755 ELPFTIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKM 814
Query: 421 VLTSLPVYALSF--FKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKEFGGL 478
+ +Y + K P + +E KF W N+K + IA S++ K + GG+
Sbjct: 815 AILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIW-----NQKRAHIAKSTLSQKNKAGGI 869
Query: 479 GVRLLREFNLALMSKWCWRLLIDRGGFWYR 508
+ + + A ++K W +WY+
Sbjct: 870 TLPDFKLYYKATVTKTAW--------YWYQ 891
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 95.9 bits (237), Expect = 3e-19
Identities = 84/319 (26%), Positives = 152/319 (47%), Gaps = 18/319 (5%)
Query: 184 VVDEAKKTK-KELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKECISTTTASI 242
V+ K K K+ ++ +D EKA+D++ ++ ++K+ + K I+ S TA+I
Sbjct: 581 VIQHINKLKNKDHMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANI 640
Query: 243 LVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVV 302
++NG F L G RQG PLSP LF I E VL ++ E KG +G+ +
Sbjct: 641 ILNGVKLKSFPLRSGTRQGCPLSPLLFNIVME---VLAIAIREEKAIKGIHIGSEE---I 694
Query: 303 SHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSL-LVGVNIGESWLLEAA 361
FADD ++ E + + L V+ ++ +SG K+N +KS+ + N ++
Sbjct: 695 KLSLFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKTVKD 754
Query: 362 SVLNCKVGKILFLYLGLSIGGDPRKL--AFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLK 419
S+ V K + YLG+ + D + L +E + I ++ W++ S+ GR+ ++K
Sbjct: 755 SIPFTVVPKKM-KYLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVK 813
Query: 420 SVLTSLPVYALSF--FKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKEFGG 477
+ +Y + K+P +E + F W N+K IA + + K + GG
Sbjct: 814 MSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIW-----NQKKPQIAKTLLSNKNKAGG 868
Query: 478 LGVRLLREFNLALMSKWCW 496
+ + LR + +++ K W
Sbjct: 869 ITLPDLRLYYKSIVIKTAW 887
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 94.7 bits (234), Expect = 8e-19
Identities = 85/324 (26%), Positives = 143/324 (43%), Gaps = 23/324 (7%)
Query: 189 KKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKECISTTTASILVNGSP 248
K K ++ +D EKA+D + ++ V+++ + IK S A+I VNG
Sbjct: 614 KLKDKNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEK 673
Query: 249 TDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVVSHLQFA 308
+ L G RQG PLSP+LF I L VL +++ + KG +G V +S L A
Sbjct: 674 LEAIPLKSGTRQGCPLSPYLFNIV---LEVLARAIRQQKEIKGIQIG-KEEVKISLL--A 727
Query: 309 DDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAASVLNCKV 368
DD ++ + R L ++ F E+ G K+N NKS+ + E +
Sbjct: 728 DDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKSMAFLYTKNKQAEKEIRETTPFSI 787
Query: 369 GKILFLYLGLSIGGDPRKL--AFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLKSVLTSLP 426
YLG+++ + + L ++ + IK L W+ S+ GR+ ++K +
Sbjct: 788 VTNNIKYLGVTLTKEVKDLYDKNFKSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILPKA 847
Query: 427 VYALSF--FKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKEFGGLGVRLLR 484
+Y + K P+ + +E KF W N K IA S + K+ GG+ + L+
Sbjct: 848 IYRFNAIPIKIPTQFFNELEGAICKFVW-----NNKKPRIAKSLLKDKRTSGGITMPDLK 902
Query: 485 EFNLALMSKWCWRLLIDRGGFWYR 508
+ A++ K W +WYR
Sbjct: 903 LYYRAIVIKTAW--------YWYR 918
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 85.5 bits (210), Expect = 5e-16
Identities = 50/153 (32%), Positives = 79/153 (50%), Gaps = 10/153 (6%)
Query: 424 SLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKE-FGGLGVRL 482
+LPVYA+S F+ + + S +F+W E+ RKISW+AW +C KE GGLG R
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 483 LREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVG-GRSVSSWWKEVAKIKDG 541
L FN AL++K +R++ R+L +RY + +E G S W+ + ++
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGREL 121
Query: 542 VGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGD 574
+ + + +GDG +T W DRW+ D
Sbjct: 122 LSRG--------LLRTIGDGIHTKVWLDRWIMD 146
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 79.0 bits (193), Expect = 4e-14
Identities = 80/315 (25%), Positives = 143/315 (45%), Gaps = 19/315 (6%)
Query: 174 ILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKE 233
I D + + +++ A++T L +D EKA+D VD YL +Q +F + ++K
Sbjct: 567 IFDNVFLIRDLLHFARRTGLSLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKT 626
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYV 293
++ + +N S T + RG+RQG PLS L+ +A E L++ + + K
Sbjct: 627 MYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLRKRLTGLVLK--- 683
Query: 294 VGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS--LLVGVN 351
VV+S +ADD +L+ + ++ + ++A S ++N++KS LL G +
Sbjct: 684 -EPDMRVVLS--AYADDVILVAQ-DLVDLERAQECQEVYAAASSARINWSKSSGLLEG-S 738
Query: 352 IGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQ--SCFL 409
+ +L A ++ + I +L + LS P F E + + TRL W+ + L
Sbjct: 739 LKVDFLPPAFRDISWESKIIKYLGVYLSAEEYPVSQNFIE-LEECVLTRLGKWKGFAKVL 797
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S GR +++ ++ S Y L I+ I+ F W G W++
Sbjct: 798 SMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFLWIGKH------WVSAGVS 851
Query: 470 CLKKEFGGLGVRLLR 484
L + GG GV +R
Sbjct: 852 SLPLKEGGQGVVCIR 866
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 65.1 bits (157), Expect = 6e-10
Identities = 50/181 (27%), Positives = 85/181 (46%), Gaps = 13/181 (7%)
Query: 175 LDGILIANEVVD----EAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
+DG L+ + ++D ++ +K + +D KA+D+V + +Q++ +
Sbjct: 113 IDGTLVNSLLLDTYISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNY 172
Query: 231 IKECISTTTASILVN-GSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLF 289
I +S +T +I V GS T + + RG++QGDPLSPFLF + L ++S
Sbjct: 173 ITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLCSLQSTPG---- 228
Query: 290 KGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVG 349
+ G + L FADD LLL + L V F + G+ +N KS+ +
Sbjct: 229 ---IGGTIGEEKIPVLAFADDLLLLEDNDVLLPTTLATVANFF-RLRGMSLNAKKSVSIS 284
Query: 350 V 350
V
Sbjct: 285 V 285
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 58.9 bits (141), Expect = 5e-08
Identities = 31/69 (44%), Positives = 44/69 (62%), Gaps = 1/69 (1%)
Query: 243 LVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVV 302
++NG+P + RGLRQGDPLSP+LF++ E L+ L + E G V ++NS +
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRV-SNNSPRI 71
Query: 303 SHLQFADDT 311
+HL FADDT
Sbjct: 72 NHLLFADDT 80
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 55.8 bits (133), Expect = 4e-07
Identities = 46/175 (26%), Positives = 79/175 (44%), Gaps = 10/175 (5%)
Query: 176 DGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKECI 235
D I + V+ + K + + +D KA+DS+ + ++ P + +++
Sbjct: 457 DNATIVDLVLRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQNTY 516
Query: 236 STTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVG 295
S+ +G ++EF RG++QGDPLSP LF + + L + S + G VG
Sbjct: 517 EGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPSEI------GAKVG 570
Query: 296 ASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGV 350
+ + + FADD +L E L L F + GLK+N +K VG+
Sbjct: 571 ---NAITNAAAFADDLVLFAETRMGLQVLLDKTLD-FLSIVGLKLNADKCFTVGI 621
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 55.5 bits (132), Expect = 5e-07
Identities = 40/143 (27%), Positives = 72/143 (49%), Gaps = 9/143 (6%)
Query: 174 ILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKE 233
+ + + + ++ EA+ K L + +D +KA+DSV+ + +++ PL R +I
Sbjct: 429 VAENTSLLSAMIKEARMKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMW 488
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYV 293
+ + V + RG+RQGDPLSP LF ++ +++ + EN G++
Sbjct: 489 VYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSPLLFNCV---MDAVLRRLPENT---GFL 542
Query: 294 VGASNSVVVSHLQFADDTLLLGE 316
+GA + L FADD +LL E
Sbjct: 543 MGAEK---IGALVFADDLVLLAE 562
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 49.7 bits (117), Expect = 3e-05
Identities = 35/143 (24%), Positives = 66/143 (45%), Gaps = 7/143 (4%)
Query: 175 LDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKEC 234
L I++ + + K + +D KA+D+V + M+ + +I
Sbjct: 3 LANIIMLEHYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMST 62
Query: 235 ISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVV 294
I+ +I+V G T++ + G++QGDPLSP LF N+++ +V +
Sbjct: 63 ITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLF-------NIVLDELVTRLNDEQPGA 115
Query: 295 GASNSVVVSHLQFADDTLLLGEK 317
+ + ++ L FADD LLL ++
Sbjct: 116 SMTPACKIASLAFADDLLLLEDR 138
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 38.9 bits (89), Expect = 0.049
Identities = 52/184 (28%), Positives = 81/184 (43%), Gaps = 23/184 (12%)
Query: 175 LDGILI----ANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
+DG+ I ++ EAK+ +KEL + +D KA++SV L + + P +
Sbjct: 267 IDGVCINVSMLTAIIAEAKRLRKELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDY 326
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAE-GLNVLMKSVVENNLF 289
I + + + G + S+ G+ QGDPLS LF +A E L L NN
Sbjct: 327 IADMYNNVITEMQFEGK-CELASILAGVYQGDPLSGPLFTLAYEKALRAL------NNEG 379
Query: 290 KGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAE---MSGLKVNFNKSL 346
+ + V V+ ++DD LLL V L+ L F E GL++N KS
Sbjct: 380 RFDIA----DVRVNASAYSDDGLLLA----MTVIGLQHNLDKFGETLAKIGLRINSRKSK 431
Query: 347 LVGV 350
V +
Sbjct: 432 TVSL 435
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 35.8 bits (81), Expect = 0.42
Identities = 25/101 (24%), Positives = 44/101 (42%), Gaps = 1/101 (0%)
Query: 190 KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKECISTTTASILVNG-SP 248
K +K L + DF KA+D V L + L W KE + T S+ +N
Sbjct: 743 KNEKSLDILFFDFAKAFDKVSHPILLKKLALFGLDKLTCSWFKEFLHLRTFSVKINKFVS 802
Query: 249 TDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLF 289
++ + + G+ QG P LF++ L + ++ + + F
Sbjct: 803 SNAYPISSGVPQGSVSGPLLFILFINDLLIDLEPNIHVSCF 843
>PPCK_ASCSU (Q05893) Phosphoenolpyruvate carboxykinase [GTP] (EC
4.1.1.32) (Phosphoenolpyruvate carboxylase) (PEPCK)
Length = 643
Score = 32.7 bits (73), Expect = 3.5
Identities = 33/141 (23%), Positives = 56/141 (39%), Gaps = 19/141 (13%)
Query: 345 SLLVGVNIG--ESWLLEAASVLNCKV--GKILFLYLGLSIGGDPRKLAFWEPVVSTIKTR 400
+L + +NIG E W+ E ++ G+ F+ LA EP + K R
Sbjct: 265 ALRIAMNIGYDEGWMAEHMLIMGVTSPKGEERFVAAAFPSACGKTNLAMLEPTIPGWKVR 324
Query: 401 LSGWQSCFLSFG--GRLVLLKSVLTSLPV----------YALSFFKSPSGIISLIESLFN 448
+ G ++ FG GRL + V A++ F+ + ++ E+
Sbjct: 325 VIGDDIAWMKFGADGRLYAINPEYGFFGVAPGTSHKTNPMAMASFQENTIFTNVAETADG 384
Query: 449 KFFWRGSE---DNRKISWIAW 466
++FW G E N K+ I W
Sbjct: 385 EYFWEGLEHEVKNPKVDMINW 405
>GYBR_STRSH (P50074) DNA gyrase subunit B, novobiocin-resistant (EC
5.99.1.3)
Length = 677
Score = 32.3 bits (72), Expect = 4.6
Identities = 29/121 (23%), Positives = 51/121 (41%), Gaps = 6/121 (4%)
Query: 239 TASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE---NNLFKGYVVG 295
T S +G TD F +H R+G+P P + IAAE L+ + + N + V
Sbjct: 248 TVSYRYDGGITD-FVVHLNARKGEPAHPSVITIAAEDTERLLSAEIALQWNGQYTDSVYS 306
Query: 296 ASNSVVVSHLQFADDTLLLGEKSWAN--VRALRAVLTLFAEMSGLKVNFNKSLLVGVNIG 353
+N++ ++ + N R R + A +SG + + ++ VN+G
Sbjct: 307 YANAIHTHEGGTHEEGFRTALTTVVNRYAREKRLLRDKDANLSGEDIREGLTAIISVNVG 366
Query: 354 E 354
E
Sbjct: 367 E 367
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.335 0.146 0.498
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,520,843
Number of Sequences: 164201
Number of extensions: 3535251
Number of successful extensions: 10997
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 10964
Number of HSP's gapped (non-prelim): 24
length of query: 738
length of database: 59,974,054
effective HSP length: 118
effective length of query: 620
effective length of database: 40,598,336
effective search space: 25170968320
effective search space used: 25170968320
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149205.5


