

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
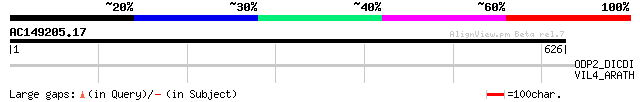
Score E
Sequences producing significant alignments: (bits) Value
ODP2_DICDI (P36413) Dihydrolipoyllysine-residue acetyltransferas... 32 3.8
VIL4_ARATH (O65570) Villin 4 32 5.0
>ODP2_DICDI (P36413) Dihydrolipoyllysine-residue acetyltransferase
component of pyruvate dehydrogenase complex,
mitochondrial precursor (EC 2.3.1.12) (E2)
(Dihydrolipoamide acetyltransferase component of
pyruvate dehydrogenase complex) (PDC-E2) (Fragment)
Length = 592
Score = 32.3 bits (72), Expect = 3.8
Identities = 36/141 (25%), Positives = 59/141 (41%), Gaps = 9/141 (6%)
Query: 470 PRKRSASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTS----QGDKIVDDDAPS 525
P + A K E K +P+HK+VG + S + G S +GD+I DA
Sbjct: 140 PVQEEAPKPKQEAPKKSTKTYPAHKVVGMP-ALSPSMETGGIASWTKKEGDQIKAGDA-I 197
Query: 526 GSIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTPLIF--VDDCKHVLEDGNES 583
+ TM F+ +G LA+ V GG + N P +I +DC + E
Sbjct: 198 AEVETDKATMDFQYEDGNGYLAKILVPGGTS-GIQINQPVCIIVKNKEDCDKFADYSVEE 256
Query: 584 EEARLSTDTICQSENQTESYS 604
+ + S+ + + + + S S
Sbjct: 257 QSSSSSSSSQESTPSSSSSSS 277
>VIL4_ARATH (O65570) Villin 4
Length = 974
Score = 32.0 bits (71), Expect = 5.0
Identities = 20/79 (25%), Positives = 43/79 (54%), Gaps = 1/79 (1%)
Query: 472 KRSASSSKCEPSAKIPPEFPS-HKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPK 530
K SA +S+ KIPP+ PS K V + +S SN+ + ++ ++D GS+
Sbjct: 832 KSSAIASRSALFEKIPPQEPSIPKPVKASPKTPESPAPESNSKEQEEKKENDKEEGSMSS 891
Query: 531 LLKTMSFEKSVEDGLLAEK 549
+++++ ++ ++G+ E+
Sbjct: 892 RIESLTIQEDAKEGVEDEE 910
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 65,174,563
Number of Sequences: 164201
Number of extensions: 2489958
Number of successful extensions: 9923
Number of sequences better than 10.0: 2
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 9923
Number of HSP's gapped (non-prelim): 2
length of query: 626
length of database: 59,974,054
effective HSP length: 116
effective length of query: 510
effective length of database: 40,926,738
effective search space: 20872636380
effective search space used: 20872636380
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149205.17


