

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
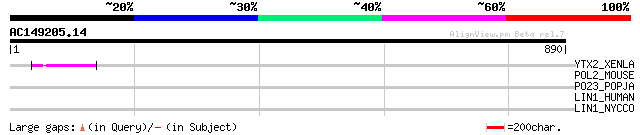
Query= AC149205.14 - phase: 0 /pseudo
(890 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 55 6e-07
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 42 0.009
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 37 0.30
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 37 0.30
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 35 0.88
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 55.5 bits (132), Expect = 6e-07
Identities = 33/105 (31%), Positives = 52/105 (49%), Gaps = 3/105 (2%)
Query: 35 PDRSILNNAMVAIEVIHFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQK 94
P R+I +N + +++HF + R L LD KA+DR+D YL + F +
Sbjct: 563 PGRTIFDNVFLIRDLLHFAR---RTGLSLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQ 619
Query: 95 WIQWMVVSTESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHL 139
++ ++ S + V +N P+ GRG+RQG PLS Y L
Sbjct: 620 FVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSL 664
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 41.6 bits (96), Expect = 0.009
Identities = 25/99 (25%), Positives = 51/99 (51%), Gaps = 4/99 (4%)
Query: 42 NAMVAIEVIHFMKTKTRGQDK-YVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWMV 100
N +I VIH++ + +DK ++ + LD KA+D++ ++ +++ + G ++ +
Sbjct: 601 NIRKSINVIHYIN---KLKDKNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIK 657
Query: 101 VSTESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHL 139
N+ VN E + + L G RQG PLS +++
Sbjct: 658 AIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNI 696
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 36.6 bits (83), Expect = 0.30
Identities = 19/89 (21%), Positives = 39/89 (43%)
Query: 51 HFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWMVVSTESVDYNV 110
H++K + Y + LDI KA+D + + M G +++ + N+
Sbjct: 11 HYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNI 70
Query: 111 LVNAEHVGPVILGRGLRQGGPLSLSFYHL 139
+V + + G++QG PLS +++
Sbjct: 71 VVGGRTTNKIYIRNGVKQGDPLSPVLFNI 99
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 36.6 bits (83), Expect = 0.30
Identities = 22/93 (23%), Positives = 47/93 (49%), Gaps = 4/93 (4%)
Query: 42 NAMVAIEVI-HFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWMV 100
N +I +I H +TK ++ + +D KA+D++ ++ + + K+G +++ +
Sbjct: 574 NIRKSINIIQHINRTK---DTNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIR 630
Query: 101 VSTESVDYNVLVNAEHVGPVILGRGLRQGGPLS 133
+ N+++N + + L G RQG PLS
Sbjct: 631 AIYDKPTANIILNGQKLEAPPLKTGTRQGCPLS 663
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 35.0 bits (79), Expect = 0.88
Identities = 24/99 (24%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 42 NAMVAIEVI-HFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWMV 100
N +I VI H K K + ++ L +D KA+D + ++ + K+G +++ +
Sbjct: 574 NIRKSINVIQHINKLKNKD---HMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIE 630
Query: 101 VSTESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHL 139
N+++N + L G RQG PLS +++
Sbjct: 631 AIYSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNI 669
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.368 0.164 0.641
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 81,634,134
Number of Sequences: 164201
Number of extensions: 2792639
Number of successful extensions: 11146
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 11139
Number of HSP's gapped (non-prelim): 6
length of query: 890
length of database: 59,974,054
effective HSP length: 119
effective length of query: 771
effective length of database: 40,434,135
effective search space: 31174718085
effective search space used: 31174718085
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149205.14


