

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.4 - phase: 0 /pseudo
(640 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
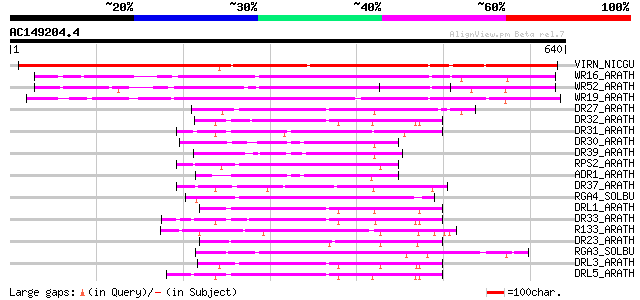
Score E
Sequences producing significant alignments: (bits) Value
VIRN_NICGU (Q40392) TMV resistance protein N 453 e-127
WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY ... 269 1e-71
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 248 3e-65
WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY ... 246 2e-64
DR27_ARATH (Q9T048) Putative disease resistance protein At4g27190 90 2e-17
DR32_ARATH (Q9FG91) Probable disease resistance protein At5g43730 87 1e-16
DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400 82 6e-15
DR30_ARATH (Q9LZ25) Probable disease resistance protein At5g04720 81 7e-15
DR39_ARATH (Q9LVT1) Putative disease resistance protein At5g47280 81 1e-14
RPS2_ARATH (Q42484) Disease resistance protein RPS2 (Resistance ... 80 2e-14
ADR1_ARATH (Q9FW44) Disease resistance protein ADR1 (Activated d... 79 3e-14
DR37_ARATH (Q9LVT4) Probable disease resistance protein At5g47250 79 4e-14
RGA4_SOLBU (Q7XA39) Putative disease resistance protein RGA4 (RG... 79 5e-14
DRL1_ARATH (P60838) Probable disease resistance protein At1g12280 78 8e-14
DR33_ARATH (Q9FG90) Probable disease resistance protein At5g43740 78 8e-14
R133_ARATH (Q9STE7) Putative disease resistance RPP13-like prote... 77 1e-13
DR23_ARATH (O82484) Putative disease resistance protein At4g10780 77 1e-13
RGA3_SOLBU (Q7XA40) Putative disease resistance protein RGA3 (RG... 76 2e-13
DRL3_ARATH (Q9LMP6) Probable disease resistance protein At1g15890 75 4e-13
DRL5_ARATH (Q9C8K0) Probable disease resistance protein At1g51480 75 5e-13
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 453 bits (1165), Expect = e-127
Identities = 253/630 (40%), Positives = 387/630 (61%), Gaps = 18/630 (2%)
Query: 11 SSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINA 70
SSS++S +Y VFLSFRGEDTRK FT HL L KGI TF+DDK LE G I +L A
Sbjct: 3 SSSSSSRWSYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCKA 62
Query: 71 IKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEE 130
I++S FAI + S +YA+S WCL+EL IMEC ++ V+P+FY VDPS VR+Q+ F +
Sbjct: 63 IEESQFAIVVFSENYATSRWCLNELVKIMECKTRFKQTVIPIFYDVDPSHVRNQKESFAK 122
Query: 131 AFRKHQEKFGQHSDRVDRWRDAFTQVASYSG-WDSKGQHEASLVENIAQHIHRKLVPKLP 189
AF +H+ K+ + + RWR A + A+ G D++ + +A + I I KL
Sbjct: 123 AFEEHETKYKDDVEGIQRWRIALNEAANLKGSCDNRDKTDADCIRQIVDQISSKLCKISL 182
Query: 190 SCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETI------RCE 243
S +N+VGI + +E++ L +G+N VR +GIWGMGG+GK+TIARA+++T+ +
Sbjct: 183 SYLQNIVGIDTHLEKIESLLEIGINGVRIMGIWGMGGVGKTTIARAIFDTLLGRMDSSYQ 242
Query: 244 FELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVL 303
F+ CFL++++E G+ LQ LLS L + ++++ DGK + + L KKVL+VL
Sbjct: 243 FDGACFLKDIKE--NKRGMHSLQNALLSELLREKANYNNEEDGKHQMASRLRSKKVLIVL 300
Query: 304 DDV-NELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFC 362
DD+ N+ + LE L G DWFG GSR+IITTRDKHL+ + + Y+ L H+++ LF
Sbjct: 301 DDIDNKDHYLEYLAGDLDWFGNGSRIIITTRDKHLIEKNDI--IYEVTALPDHESIQLFK 358
Query: 363 LKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPR 422
AF + P E + LS EVV+Y GLPLAL+V GS L+ + W SA++ +++ +
Sbjct: 359 QHAFGKEVPNENFEKLSLEVVNYAKGLPLALKVWGSLLHNLRLTEWKSAIEHMKNNSYSG 418
Query: 423 VQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLIT 482
+ D LKISYD L+ ++++FLDIACF +G + D ++ ILESC + G++ILI++SL+
Sbjct: 419 IIDKLKISYDGLEPKQQEMFLDIACFLRGEEKDYILQILESCHIGAEYGLRILIDKSLVF 478
Query: 483 LDSVNNKLGMHDLLQEMGRDIV-FQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSID 541
+ S N++ MHDL+Q+MG+ IV FQ+ DP RSRLW ++++ V++ N GT A+ +I
Sbjct: 479 I-SEYNQVQMHDLIQDMGKYIVNFQK---DPGERSRLWLAKEVEEVMSNNTGTMAMEAIW 534
Query: 542 MKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITT 601
+ ++ +A +L+ ++ + LP++L+ P ++ P T
Sbjct: 535 VSSYSS-TLRFSNQAVKNMKRLRVFNMGRSSTHYAIDYLPNNLRCFVCTNYPWESFPSTF 593
Query: 602 QLDELVDITLSHSKIEQLWQGVKVIDSLTK 631
+L LV + L H+ + LW K + SL +
Sbjct: 594 ELKMLVHLQLRHNSLRHLWTETKHLPSLRR 623
>WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY
DNA-binding protein 16)
Length = 1372
Score = 269 bits (688), Expect = 1e-71
Identities = 190/626 (30%), Positives = 307/626 (48%), Gaps = 70/626 (11%)
Query: 29 EDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAITILSPDYASS 88
E+ R F HL AL+RKG+ D D +S + + ++ + ++ IL +
Sbjct: 14 EEVRYSFVSHLSKALQRKGVNDVFIDSD----DSLSNESQSMVERARVSVMILP---GNR 66
Query: 89 TWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDR 148
T LD+L +++C + V+PV YGV S+ + F + SD
Sbjct: 67 TVSLDKLVKVLDCQKNKDQVVVPVLYGVRSSETEWLSALDSKGFSSVHHSRKECSD---- 122
Query: 149 WRDAFTQVASYSGWDSKGQHEASLVENIAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKF 208
+ LV+ + ++ KL +GI SK+ E+ K
Sbjct: 123 ---------------------SQLVKETVRDVYEKLFYM------ERIGIYSKLLEIEKM 155
Query: 209 LGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLEN-VREISETNGLVHLQR 267
+ D+R +GIWGM GIGK+T+A+AV++ + EF+ CF+E+ + I E L+
Sbjct: 156 INKQPLDIRCVGIWGMPGIGKTTLAKAVFDQMSGEFDAHCFIEDYTKAIQEKGVYCLLEE 215
Query: 268 QLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSR 327
Q L + + L +++ L K+VL+VLDDV +E+ +G DWFGP S
Sbjct: 216 QFLKENAGASGTVTKL----SLLRDRLNNKRVLVVLDDVRSPLVVESFLGGFDWFGPKSL 271
Query: 328 VIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCG 387
+IIT++DK + V++ Y+ L + +AL LF L A D ++ ++S +V+ Y
Sbjct: 272 IIITSKDKSVFRLCRVNQIYEVQGLNEKEALQLFSLCASIDDMAEQNLHEVSMKVIKYAN 331
Query: 388 GLPLALEVLGSYLYGRNIDV-WHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIA 446
G PLAL + G L G+ A KL+ P D +K SYD+L+ EK+IFLDIA
Sbjct: 332 GHPLALNLYGRELMGKKRPPEMEIAFLKLKECPPAIFVDAIKSSYDTLNDREKNIFLDIA 391
Query: 447 CFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQ 506
CFF+G D V+ +LE CG+FP +GI +L+E+SL+T+ N++ MH+L+Q++GR I+ +
Sbjct: 392 CFFQGENVDYVMQLLEGCGFFPHVGIDVLVEKSLVTIS--ENRVRMHNLIQDVGRQIINR 449
Query: 507 ESPNDPCRRSRLW------------SQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWNT 554
E+ RRSRLW Q + + T + + I+ L ++
Sbjct: 450 ET-RQTKRRSRLWEPCSIKYLLEDKEQNENEEQKTTFERAQVPEEIEGMFLDTSNLSFDI 508
Query: 555 E--AFSKTSQLKFLSLCEMQ---------LPLGLSCLPSSLKVLHWRGCPLKTLPITTQL 603
+ AF L+ + L LS LP+ L++LHW PL+ LP
Sbjct: 509 KHVAFDNMLNLRLFKIYSSNPEVHHVNNFLKGSLSSLPNVLRLLHWENYPLQFLPQNFDP 568
Query: 604 DELVDITLSHSKIEQLWQGVKVIDSL 629
LV+I + +S++++LW G K ++ L
Sbjct: 569 IHLVEINMPYSQLKKLWGGTKDLEML 594
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 248 bits (633), Expect = 3e-65
Identities = 190/630 (30%), Positives = 306/630 (48%), Gaps = 74/630 (11%)
Query: 29 EDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAITILSPDYASS 88
E+ R F HL AL RKGI D D++ ++ ++ I+ + ++ +L + S
Sbjct: 17 EEVRYSFVSHLSEALRRKGINNVVVDVDID--DLLFKESQAKIEKAGVSVMVLPGNCDPS 74
Query: 89 TWCLDELQMIMECSSKN-NLHVLPVFYGVDPSDVRHQ---RGCFEEAFRKHQEKFGQHSD 144
LD+ ++EC N + V+ V YG S +R Q F R HQ +
Sbjct: 75 EVWLDKFAKVLECQRNNKDQAVVSVLYG--DSLLRDQWLSELDFRGLSRIHQSR------ 126
Query: 145 RVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIHRKLVPKLPSCTENLVGIVSKVEE 204
K ++ LVE I + ++ +GI SK+ E
Sbjct: 127 --------------------KECSDSILVEEIVRDVYET------HFYVGRIGIYSKLLE 160
Query: 205 VNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISETNGLVH 264
+ + +R +GIWGM GIGK+T+A+AV++ + F+ +CF+E+ + GL
Sbjct: 161 IENMVNKQPIGIRCVGIWGMPGIGKTTLAKAVFDQMSSAFDASCFIEDYDKSIHEKGLYC 220
Query: 265 LQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGP 324
L + L + ND + ++++ L K+VL+VLDDV E+ + DW GP
Sbjct: 221 LLEEQL----LPGNDATIMK--LSSLRDRLNSKRVLVVLDDVCNALVAESFLEGFDWLGP 274
Query: 325 GSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKA-FKGDKPQEGYLDLSKEVV 383
GS +IIT+RDK + G+++ Y+ L + +A LF L A D ++ +LS V+
Sbjct: 275 GSLIIITSRDKQVFRLCGINQIYEVQGLNEKEARQLFLLSASIMEDMGEQNLHELSVRVI 334
Query: 384 DYCGGLPLALEVLGSYLYGRN-IDVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIF 442
Y G PLA+ V G L G+ + +A KL+ P ++ D K SYD+L EK+IF
Sbjct: 335 SYANGNPLAISVYGRELKGKKKLSEMETAFLKLKRRPPFKIVDAFKSSYDTLSDNEKNIF 394
Query: 443 LDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRD 502
LDIACFF+G + VI +LE CG+FP + I +L+++ L+T+ N++ +H L Q++GR+
Sbjct: 395 LDIACFFQGENVNYVIQLLEGCGFFPHVEIDVLVDKCLVTIS--ENRVWLHKLTQDIGRE 452
Query: 503 IVFQESPNDPCRRSRLWSQEDIDRVLTKN------------KGTEAINSIDMKLLQPYEA 550
I+ E+ RR RLW I +L N K + I+ L
Sbjct: 453 IINGETVQIE-RRRRLWEPWSIKYLLEYNEHKANGEPKTTFKRAQGSEEIEGLFLDTSNL 511
Query: 551 HWNTE--AFSKTSQLKFLSL-CE-------MQLPLG-LSCLPSSLKVLHWRGCPLKTLPI 599
++ + AF L+ L + C + P G L LP+ L++LHW PLK+LP
Sbjct: 512 RFDLQPSAFKNMLNLRLLKIYCSNPEVHPVINFPTGSLHSLPNELRLLHWENYPLKSLPQ 571
Query: 600 TTQLDELVDITLSHSKIEQLWQGVKVIDSL 629
LV+I + +S++++LW G K ++ L
Sbjct: 572 NFDPRHLVEINMPYSQLQKLWGGTKNLEML 601
Score = 50.1 bits (118), Expect = 2e-05
Identities = 28/82 (34%), Positives = 48/82 (58%), Gaps = 1/82 (1%)
Query: 427 LKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSV 486
L++SYD L M+K +FL IA F D V ++ G+++L + SLI++ S
Sbjct: 1089 LRVSYDDLQEMDKVLFLYIASLFNDEDVDFVAPLIAGIDLDVSSGLKVLADVSLISVSS- 1147
Query: 487 NNKLGMHDLLQEMGRDIVFQES 508
N ++ MH L ++MG++I+ +S
Sbjct: 1148 NGEIVMHSLQRQMGKEILHGQS 1169
>WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY
DNA-binding protein 19)
Length = 1895
Score = 246 bits (627), Expect = 2e-64
Identities = 181/628 (28%), Positives = 312/628 (48%), Gaps = 49/628 (7%)
Query: 20 YHVFLSF-RGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
Y V + + R + + + F HL A+L R+GI+ ++ + ++A+ I
Sbjct: 668 YDVVIRYGRADISNEDFISHLRASLCRRGISVYEKFNE-----------VDALPKCRVLI 716
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
+L+ Y S L I+E + V P+FY + P D +E + + + K
Sbjct: 717 IVLTSTYVPSN-----LLNILEHQHTEDRVVYPIFYRLSPYDFVCNSKNYERFYLQDEPK 771
Query: 139 FGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIHRKLVPKLPSCTENLVGI 198
+W+ A ++ G+ + E+ L++ I + + L + N++G+
Sbjct: 772 ---------KWQAALKEITQMPGYTLTDKSESELIDEIVRDALKVLCS---ADKVNMIGM 819
Query: 199 VSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISE 258
+VEE+ L + DVR IGIWG GIGK+TIA ++ I ++E L+++ + E
Sbjct: 820 DMQVEEILSLLCIESLDVRSIGIWGTVGIGKTTIAEEIFRKISVQYETCVVLKDLHKEVE 879
Query: 259 TNGLVHLQRQLLSHLSISRNDFHDLYDGKKT-IQNSLCRKKVLLVLDDVNELNQLENLVG 317
G ++ LS + + D K + +++ L RK++L++LDDVN+ ++ +G
Sbjct: 880 VKGHDAVRENFLSEVLEVEPHVIRISDIKTSFLRSRLQRKRILVILDDVNDYRDVDTFLG 939
Query: 318 KQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLD 377
++FGPGSR+I+T+R++ + + + Y+ L +L+L + E Y
Sbjct: 940 TLNYFGPGSRIIMTSRNRRVFVLCKIDHVYEVKPLDIPKSLLLLDRGTCQIVLSPEVYKT 999
Query: 378 LSKEVVDYCGGLPLALEVLGSYLYGRNID-VWHSAVKKLRSFPHPRVQDNLKISYDSLDT 436
LS E+V + G P L+ L S ID W+ +++++ + + S LD
Sbjct: 1000 LSLELVKFSNGNPQVLQFLSS------IDREWNKLSQEVKTTSPIYIPGIFEKSCCGLDD 1053
Query: 437 MEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHDLL 496
E+ IFLDIACFF + D V +L+ CG+ +G + L+++SL+T+ S +N + M +
Sbjct: 1054 NERGIFLDIACFFNRIDKDNVAMLLDGCGFSAHVGFRGLVDKSLLTI-SQHNLVDMLSFI 1112
Query: 497 QEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQ-PYEAHWNTE 555
Q GR+IV QES + P RSRLW+ + I V + GT AI I + +L ++A N
Sbjct: 1113 QATGREIVRQESADRPGDRSRLWNADYIRHVFINDTGTSAIEGIFLDMLNLKFDA--NPN 1170
Query: 556 AFSKTSQLKFLSL-CE-------MQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELV 607
F K L+ L L C + P GL LPS L++LHW PL +LP + + LV
Sbjct: 1171 VFEKMCNLRLLKLYCSKAEEKHGVSFPQGLEYLPSKLRLLHWEYYPLSSLPKSFNPENLV 1230
Query: 608 DITLSHSKIEQLWQGVKVIDSLTKNSIK 635
++ L S ++LW+G K T +S++
Sbjct: 1231 ELNLPSSCAKKLWKGKKARFCTTNSSLE 1258
>DR27_ARATH (Q9T048) Putative disease resistance protein At4g27190
Length = 985
Score = 89.7 bits (221), Expect = 2e-17
Identities = 92/349 (26%), Positives = 165/349 (46%), Gaps = 33/349 (9%)
Query: 210 GMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCE-----FELTCFLENVREISETNGLVH 264
G+ + IG+WGMGG+GK+T+ R + +R E F L F+ +E
Sbjct: 158 GLTSEKAQKIGVWGMGGVGKTTLVRTLNNKLREEGATQPFGLVIFVIVSKEFDPR----E 213
Query: 265 LQRQLLSHLSI-SRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFG 323
+Q+Q+ L I ++ + + ++ + +K LL+LDDV + L+ L +
Sbjct: 214 VQKQIAERLDIDTQMEESEEKLARRIYVGLMKERKFLLILDDVWKPIDLDLLGIPRTEEN 273
Query: 324 PGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYL-DLSKEV 382
GS+VI+T+R + + + L + DA LFC A GD + ++ ++K V
Sbjct: 274 KGSKVILTSRFLEVCRSMKTDLDVRVDCLLEEDAWELFCKNA--GDVVRSDHVRKIAKAV 331
Query: 383 VDYCGGLPLALEVLGSYLYG-RNIDVWHSAVKKL-RSFP-----HPRVQDNLKISYDSLD 435
CGGLPLA+ +G+ + G +N+ +W+ + KL +S P ++ LK+SYD L+
Sbjct: 332 SQECGGLPLAIITVGTAMRGKKNVKLWNHVLSKLSKSVPWIKSIEEKIFQPLKLSYDFLE 391
Query: 436 TMEKDIFLDIACFFK--GMKGDKVIDILESCGYFPQIGIQ-ILIERSLITLDSVNNKLGM 492
K FL A F + ++ +V+ + G+ ++G Q + + T++S+ + +
Sbjct: 392 DKAKFCFLLCALFPEDYSIEVTEVVRYWMAEGFMEELGSQEDSMNEGITTVESLKDYCLL 451
Query: 493 HDLLQEMGRDIVFQESPNDPCRRSRLW----SQEDIDRVLTKNKGTEAI 537
D RD V +D R +W SQ+D ++ G + I
Sbjct: 452 ED---GDRRDTV---KMHDVVRDFAIWIMSSSQDDSHSLVMSGTGLQDI 494
>DR32_ARATH (Q9FG91) Probable disease resistance protein At5g43730
Length = 848
Score = 87.0 bits (214), Expect = 1e-16
Identities = 92/316 (29%), Positives = 151/316 (47%), Gaps = 37/316 (11%)
Query: 214 NDVRFIGIWGMGGIGKSTIARAV---YETIRCEFELTCFLENVREISETNGLVHLQRQLL 270
+++R +G++GMGGIGK+T+ ++ + + EF++ ++ +S+ L +Q Q+L
Sbjct: 170 DEIRTLGLYGMGGIGKTTLLESLNNKFVELESEFDVVIWV----VVSKDFQLEGIQDQIL 225
Query: 271 SHLSISRNDFHDLYDGKKT--IQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRV 328
L + ++ + KK I N+L RKK +L+LDD+ L + GS++
Sbjct: 226 GRLRPDK-EWERETESKKASLINNNLKRKKFVLLLDDLWSEVDLIKIGVPPPSRENGSKI 284
Query: 329 IITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLD---LSKEVVDY 385
+ TTR K + K K L +A LF L GD + D L++ V
Sbjct: 285 VFTTRSKEVCKHMKADKQIKVDCLSPDEAWELFRLTV--GDIILRSHQDIPALARIVAAK 342
Query: 386 CGGLPLALEVLGSYLYGR-NIDVWHSAVKKLRS----FP--HPRVQDNLKISYDSLDTME 438
C GLPLAL V+G + + + W A+ L S FP R+ LK SYDSL E
Sbjct: 343 CHGLPLALNVIGKAMVCKETVQEWRHAINVLNSPGHKFPGMEERILPILKFSYDSLKNGE 402
Query: 439 -KDIFLDIACFFKG--MKGDKVIDILESCGY-----FPQIG-------IQILIERSLITL 483
K FL + F + ++ DK+I+ GY + G I +L+ L+
Sbjct: 403 IKLCFLYCSLFPEDFEIEKDKLIEYWICEGYINPNRYEDGGTNQGYDIIGLLVRAHLLIE 462
Query: 484 DSVNNKLGMHDLLQEM 499
+ +K+ MHD+++EM
Sbjct: 463 CELTDKVKMHDVIREM 478
>DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400
Length = 874
Score = 81.6 bits (200), Expect = 6e-15
Identities = 85/329 (25%), Positives = 146/329 (43%), Gaps = 34/329 (10%)
Query: 193 ENLVGIVSKVEEV-NKFLGMGLNDVRFIGIWGMGGIGKSTIARAV---YETIRCEFELTC 248
+ +VG + VE N + +G V +GI+GMGG+GK+T+ + + T+ +F++
Sbjct: 154 QEIVGQEAIVESTWNSMMEVG---VGLLGIYGMGGVGKTTLLSQINNKFRTVSNDFDIAI 210
Query: 249 FLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGK--KTIQNSLCRKKVLLVLDDV 306
++ +S+ + +Q + L + + + + TI+ SL KK +L+LDD+
Sbjct: 211 WV----VVSKNPTVKRIQEDIGKRLDLYNEGWEQKTENEIASTIKRSLENKKYMLLLDDM 266
Query: 307 NELNQLENL---VGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCL 363
L N+ V K++ GS++ T+R + GV K + L DA LF
Sbjct: 267 WTKVDLANIGIPVPKRN----GSKIAFTSRSNEVCGKMGVDKEIEVTCLMWDDAWDLFTR 322
Query: 364 KAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYL-YGRNIDVWHSAVKKLRSFPHPR 422
+ + +++K + C GLPLAL V+G + ++I+ WH AV
Sbjct: 323 NMKETLESHPKIPEVAKSIARKCNGLPLALNVIGETMARKKSIEEWHDAVGVFSGI-EAD 381
Query: 423 VQDNLKISYDSLDTME-KDIFLDIACFFKGMK-----------GDKVIDILESCGYFPQI 470
+ LK SYD L + K FL A F + + G +I + Y
Sbjct: 382 ILSILKFSYDDLKCEKTKSCFLFSALFPEDYEIGKDDLIEYWVGQGIILGSKGINYKGYT 441
Query: 471 GIQILIERSLITLDSVNNKLGMHDLLQEM 499
I L L+ K+ MHD+++EM
Sbjct: 442 IIGTLTRAYLLKESETKEKVKMHDVVREM 470
>DR30_ARATH (Q9LZ25) Probable disease resistance protein At5g04720
Length = 811
Score = 81.3 bits (199), Expect = 7e-15
Identities = 75/263 (28%), Positives = 122/263 (45%), Gaps = 31/263 (11%)
Query: 196 VGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVY--ETIRCEF-ELTCFLEN 252
VG+ +V + L ++ R IGI GM G GK+T+A+ + E +R F FL
Sbjct: 180 VGLDLGKRKVKEMLFKSIDGERLIGISGMSGSGKTTLAKELARDEEVRGHFGNKVLFLT- 238
Query: 253 VREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQL 312
+S++ L L+ + L+ + + +L + L++LDDV L
Sbjct: 239 ---VSQSPNLEELRAHIWGFLT----------SYEAGVGATLPESRKLVILDDVWTRESL 285
Query: 313 ENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQ 372
+ L+ + PG+ ++ +R K TY +L +H+A LFCL F
Sbjct: 286 DQLMFENI---PGTTTLVVSRSK----LADSRVTYDVELLNEHEATALFCLSVFNQKLVP 338
Query: 373 EGYLD-LSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKL-RSFP-----HPRVQD 425
G+ L K+VV C GLPL+L+V+G+ L R W AV++L R P RV
Sbjct: 339 SGFSQSLVKQVVGECKGLPLSLKVIGASLKERPEKYWEGAVERLSRGEPADETHESRVFA 398
Query: 426 NLKISYDSLDTMEKDIFLDIACF 448
++ + ++LD +D FL + F
Sbjct: 399 QIEATLENLDPKTRDCFLVLGAF 421
>DR39_ARATH (Q9LVT1) Putative disease resistance protein At5g47280
Length = 623
Score = 80.9 bits (198), Expect = 1e-14
Identities = 77/251 (30%), Positives = 116/251 (45%), Gaps = 27/251 (10%)
Query: 213 LND-VRFIGIWGMGGIGKSTIARAVY--ETIRCEFELTCFLENVREISETNGLVHLQRQL 269
LND R IGI GM G GK+ +A+ + E +R F V + L L R
Sbjct: 5 LNDEARIIGISGMIGSGKTILAKELARDEEVRGHFANRVLFLTVSQSPNLEELRSLIRDF 64
Query: 270 LSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVI 329
L+ H+ G + S+ + L++LDDV L+ L+ PG+ +
Sbjct: 65 LTG--------HEAGFGT-ALPESVGHTRKLVILDDVRTRESLDQLMFNI----PGTTTL 111
Query: 330 ITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYL-DLSKEVVDYCGG 388
+ ++ K + TY +L +HDA LFCL AF G+ L K+VV G
Sbjct: 112 VVSQSKLV----DPRTTYDVELLNEHDATSLFCLSAFNQKSVPSGFSKSLVKQVVGESKG 167
Query: 389 LPLALEVLGSYLYGRNIDVWHSAVKKL-RSFP-----HPRVQDNLKISYDSLDTMEKDIF 442
LPL+L+VLG+ L R W AV++L R P +V ++ + ++LD K+ F
Sbjct: 168 LPLSLKVLGASLNDRPETYWAIAVERLSRGEPVDETHESKVFAQIEATLENLDPKTKECF 227
Query: 443 LDIACFFKGMK 453
LD+ F +G K
Sbjct: 228 LDMGAFPEGKK 238
>RPS2_ARATH (Q42484) Disease resistance protein RPS2 (Resistance to
Pseudomonas syringae protein 2)
Length = 909
Score = 80.1 bits (196), Expect = 2e-14
Identities = 75/267 (28%), Positives = 130/267 (48%), Gaps = 16/267 (5%)
Query: 193 ENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRC---EFELTCF 249
+++VG + +E+V +FL + IG++G GG+GK+T+ +++ + ++++ +
Sbjct: 153 KSVVGNTTMMEQVLEFLSEE-EERGIIGVYGPGGVGKTTLMQSINNELITKGHQYDVLIW 211
Query: 250 LENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNEL 309
++ RE E +Q+ + + L +S ++ + I +L +K+ LL+LDDV E
Sbjct: 212 VQMSREFGECT----IQQAVGARLGLSWDEKETGENRALKIYRALRQKRFLLLLDDVWEE 267
Query: 310 NQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGD 369
LE + +V+ TTR L G + L K A LFC K ++ D
Sbjct: 268 IDLEKTGVPRPDRENKCKVMFTTRSIALCNNMGAEYKLRVEFLEKKHAWELFCSKVWRKD 327
Query: 370 KPQEGYLD-LSKEVVDYCGGLPLALEVLGSYLYGRNI-DVWHSAVKKLRSFPHPRVQDN- 426
+ + L++ +V CGGLPLAL LG + R + W A + L FP N
Sbjct: 328 LLESSSIRRLAEIIVSKCGGLPLALITLGGAMAHRETEEEWIHASEVLTRFPAEMKGMNY 387
Query: 427 ----LKISYDSLDT-MEKDIFLDIACF 448
LK SYD+L++ + + FL A F
Sbjct: 388 VFALLKFSYDNLESDLLRSCFLYCALF 414
>ADR1_ARATH (Q9FW44) Disease resistance protein ADR1 (Activated
disease resistance protein 1)
Length = 787
Score = 79.3 bits (194), Expect = 3e-14
Identities = 70/242 (28%), Positives = 111/242 (44%), Gaps = 36/242 (14%)
Query: 215 DVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLS 274
D GI GM G GK+T+A E+S+ + + L + + L+
Sbjct: 185 DTHLFGISGMSGSGKTTLAI--------------------ELSKDDDVRGLFKNKVLFLT 224
Query: 275 ISRN-DFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTR 333
+SR+ +F +L + ++ L++LDDV L+ L+ K GS ++ +R
Sbjct: 225 VSRSPNFENLESCIREFLYDGVHQRKLVILDDVWTRESLDRLMSKIR----GSTTLVVSR 280
Query: 334 DKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLD-LSKEVVDYCGGLPLA 392
K TY +L K +A+ L CL AF+ P + L K+VVD C GLPL+
Sbjct: 281 SK----LADPRTTYNVELLKKDEAMSLLCLCAFEQKSPPSPFNKYLVKQVVDECKGLPLS 336
Query: 393 LEVLGSYLYGRNIDVWHSAVKKL------RSFPHPRVQDNLKISYDSLDTMEKDIFLDIA 446
L+VLG+ L + W VK+L RV +++ S ++LD +D FLD+
Sbjct: 337 LKVLGASLKNKPERYWEGVVKRLLRGEAADETHESRVFAHMEESLENLDPKIRDCFLDMG 396
Query: 447 CF 448
F
Sbjct: 397 AF 398
Score = 33.5 bits (75), Expect = 1.8
Identities = 25/99 (25%), Positives = 40/99 (40%), Gaps = 26/99 (26%)
Query: 518 LWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQ----- 572
+W++E +DR+++K +G+ + KL P +N E K + L LC +
Sbjct: 257 VWTRESLDRLMSKIRGSTTLVVSRSKLADP-RTTYNVELLKKDEAMSLLCLCAFEQKSPP 315
Query: 573 -----------------LPLGLSCLPSSLK---VLHWRG 591
LPL L L +SLK +W G
Sbjct: 316 SPFNKYLVKQVVDECKGLPLSLKVLGASLKNKPERYWEG 354
>DR37_ARATH (Q9LVT4) Probable disease resistance protein At5g47250
Length = 843
Score = 79.0 bits (193), Expect = 4e-14
Identities = 84/338 (24%), Positives = 159/338 (46%), Gaps = 36/338 (10%)
Query: 193 ENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAV---YETIRCEFELTCF 249
+ VG+ + +E+ + L N R +GI+GMGG+GK+T+ + + + ++++ +
Sbjct: 155 QQTVGLDTTLEKTWESLRKDEN--RMLGIFGMGGVGKTTLLTLINNKFVEVSDDYDVVIW 212
Query: 250 LENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLC----RKKVLLVLDD 305
+E+ ++ + +Q + L I N++ GKK + S + + +L+LDD
Sbjct: 213 VESSKDAD----VGKIQDAIGERLHICDNNWSTYSRGKKASEISRVLRDMKPRFVLLLDD 268
Query: 306 VNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKA 365
+ E L + G +V+ TTR K + ++ + L ++DA LF +K
Sbjct: 269 LWEDVSLTAI--GIPVLGKKYKVVFTTRSKDVCSVMRANEDIEVQCLSENDAWDLFDMKV 326
Query: 366 FKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDV-WHSAVKKLRSF------ 418
+ D++K++V C GLPLALEV+ + ++ + W A+ L S+
Sbjct: 327 HCDGLNEIS--DIAKKIVAKCCGLPLALEVIRKTMASKSTVIQWRRALDTLESYRSEMKG 384
Query: 419 PHPRVQDNLKISYDSLDTMEKDIFLDIACFFKG--MKGDKVIDILESCGYFPQ-IGIQIL 475
+ LK+SYD L T FL A F K +K D++++ G+ + G +
Sbjct: 385 TEKGIFQVLKLSYDYLKTKNAKCFLYCALFPKAYYIKQDELVEYWIGEGFIDEKDGRERA 444
Query: 476 IERSLITLDSV---------NNKLGMHDLLQEMGRDIV 504
+R +D++ N K+ MHD++++M IV
Sbjct: 445 KDRGYEIIDNLVGAGLLLESNKKVYMHDMIRDMALWIV 482
>RGA4_SOLBU (Q7XA39) Putative disease resistance protein RGA4
(RGA4-blb)
Length = 988
Score = 78.6 bits (192), Expect = 5e-14
Identities = 81/302 (26%), Positives = 138/302 (44%), Gaps = 26/302 (8%)
Query: 203 EEVNKFLGMGLN---DVRFIGIWGMGGIGKSTIARAVY--ETIRCEFELTCFLENVREIS 257
+E+ K L +N ++ I GMGG+GK+T+A+ ++ E + F ++ +
Sbjct: 161 DEIVKILINNVNVAEELPVFPIIGMGGLGKTTLAQMIFNDERVTKHFNPKIWVCVSDDFD 220
Query: 258 ETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDV--NELNQLENL 315
E L + ++ ++ S DL +K +Q L K+ LLVLDDV ++L + L
Sbjct: 221 EKR----LIKTIIGNIERSSPHVEDLASFQKKLQELLNGKRYLLVLDDVWNDDLEKWAKL 276
Query: 316 VGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGY 375
G+ ++ TTR + + G + Y L HD+L+LF +AF K
Sbjct: 277 RAVLTVGARGASILATTRLEKVGSIMGTLQPYHLSNLSPHDSLLLFMQRAFGQQKEANPN 336
Query: 376 L-DLSKEVVDYCGGLPLALEVLGSYL-YGRNIDVW-HSAVKKLRSFPHPR--VQDNLKIS 430
L + KE+V CGG+PLA + LG L + R W H ++ S P + L++S
Sbjct: 337 LVAIGKEIVKKCGGVPLAAKTLGGLLRFKREESEWEHVRDNEIWSLPQDESSILPALRLS 396
Query: 431 YDSLDTMEKDIFLDIACFFKGMK--GDKVIDILESCGYFPQIGIQILIERSLITLDSVNN 488
Y L + F A F K K + +I + + G+ L+ + + L+ V N
Sbjct: 397 YHHLPLDLRQCFAYCAVFPKDTKMIKENLITLWMAHGF--------LLSKGNLELEDVGN 448
Query: 489 KL 490
++
Sbjct: 449 EV 450
>DRL1_ARATH (P60838) Probable disease resistance protein At1g12280
Length = 894
Score = 77.8 bits (190), Expect = 8e-14
Identities = 79/311 (25%), Positives = 152/311 (48%), Gaps = 36/311 (11%)
Query: 219 IGIWGMGGIGKSTIARAVYETI--RCE-FELTCFLENVREISETNGLVHLQRQLLSHLSI 275
+G++GMGG+GK+T+ + +C F + ++ +S++ + +Q + L +
Sbjct: 179 VGLYGMGGVGKTTLLTRINNKFSEKCSGFGVVIWV----VVSKSPDIHRIQGDIGKRLDL 234
Query: 276 SRNDFHDLYDGKKT--IQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTR 333
++ ++ + ++ I N L ++K +L+LDD+ E LE L G +V+ TTR
Sbjct: 235 GGEEWDNVNENQRALDIYNVLGKQKFVLLLDDIWEKVNLEVLGVPYPSRQNGCKVVFTTR 294
Query: 334 DKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLD---LSKEVVDYCGGLP 390
+ + V + L ++A LF +K G+ +G+ D L+++V C GLP
Sbjct: 295 SRDVCGRMRVDDPMEVSCLEPNEAWELFQMKV--GENTLKGHPDIPELARKVAGKCCGLP 352
Query: 391 LALEVLGSYL-YGRNIDVWHSAVKKLRSFP-----HPRVQDNLKISYDSLDTME-KDIFL 443
LAL V+G + R + W +A+ L S+ ++ LK SYD+L+ + K FL
Sbjct: 353 LALNVIGETMACKRMVQEWRNAIDVLSSYAAEFPGMEQILPILKYSYDNLNKEQVKPCFL 412
Query: 444 DIACFFKG--MKGDKVIDILESCGYFPQIG------------IQILIERSLITLDSVN-N 488
+ F + M+ +++ID G+ + I IL+ L+ +++N
Sbjct: 413 YCSLFPEDYRMEKERLIDYWICEGFIDENESRERALSQGYEIIGILVRACLLLEEAINKE 472
Query: 489 KLGMHDLLQEM 499
++ MHD+++EM
Sbjct: 473 QVKMHDVVREM 483
>DR33_ARATH (Q9FG90) Probable disease resistance protein At5g43740
Length = 862
Score = 77.8 bits (190), Expect = 8e-14
Identities = 99/355 (27%), Positives = 160/355 (44%), Gaps = 44/355 (12%)
Query: 176 IAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMGGIGKSTIAR 234
+AQ I K+ KL T VG+ VE L +ND + +G++GMGG+GK+T+
Sbjct: 136 VAQEIIHKVEKKLIQTT---VGLDKLVEMAWSSL---MNDEIGTLGLYGMGGVGKTTLLE 189
Query: 235 AV---YETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKT-- 289
++ + + EF++ ++ +S+ +Q Q+L L S ++ + KK
Sbjct: 190 SLNNKFVELESEFDVVIWV----VVSKDFQFEGIQDQILGRLR-SDKEWERETESKKASL 244
Query: 290 IQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKT 349
I N+L RKK +L+LDD+ + + GS+++ TTR + K K
Sbjct: 245 IYNNLERKKFVLLLDDLWSEVDMTKIGVPPPTRENGSKIVFTTRSTEVCKHMKADKQIKV 304
Query: 350 GMLCKHDALVLFCLKAFKGDKPQEGYLD---LSKEVVDYCGGLPLALEVLGSYLYGR-NI 405
L +A LF L GD + D L++ V C GLPLAL V+G + + I
Sbjct: 305 ACLSPDEAWELFRLTV--GDIILRSHQDIPALARIVAAKCHGLPLALNVIGKAMSCKETI 362
Query: 406 DVWHSAVKKLRSFPH------PRVQDNLKISYDSLDTME-KDIFLDIACFFKG--MKGDK 456
W A+ L S H R+ LK SYDSL E K FL + F + + +K
Sbjct: 363 QEWSHAINVLNSAGHEFPGMEERILPILKFSYDSLKNGEIKLCFLYCSLFPEDSEIPKEK 422
Query: 457 VIDILESCGY-----FPQIG-------IQILIERSLITLDSVNNKLGMHDLLQEM 499
I+ G+ + G I +L+ L+ + + + MHD+++EM
Sbjct: 423 WIEYWICEGFINPNRYEDGGTNHGYDIIGLLVRAHLLIECELTDNVKMHDVIREM 477
>R133_ARATH (Q9STE7) Putative disease resistance RPP13-like protein
3
Length = 847
Score = 77.4 bits (189), Expect = 1e-13
Identities = 100/389 (25%), Positives = 172/389 (43%), Gaps = 54/389 (13%)
Query: 174 ENIAQHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMGLNDVR-----FIGIWGMGGIG 228
ENI R+L P E LV V ++V L L+D I I+GMGG+G
Sbjct: 140 ENITNVRVRQLRRAPPVDQEELV--VGLEDDVKILLVKLLSDNEKDKSYIISIFGMGGLG 197
Query: 229 KSTIARAVYET--IRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDG 286
K+ +AR +Y + ++ F+ + +E + L+ + R L +S + +++
Sbjct: 198 KTALARKLYNSGDVKRRFDCRAWTYVSQEYKTRDILIRIIRSL-GIVSAEEMEKIKMFEE 256
Query: 287 KKTIQ----NSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLM-TH 341
+ ++ L K ++V+DDV + + E+L GS+VIITTR + +
Sbjct: 257 DEELEVYLYGLLEGKNYMVVVDDVWDPDAWESLKRALPCDHRGSKVIITTRIRAIAEGVE 316
Query: 342 GVHKTYKTGMLCKHDALVLFCLKAFKG-DKPQEGYLDLSKEVVDYCGGLPLALEVLGSYL 400
G +K L ++ LF KAF +K E KE+V CGGLPLA+ VL L
Sbjct: 317 GTVYAHKLRFLTFEESWTLFERKAFSNIEKVDEDLQRTGKEMVKKCGGLPLAIVVLSGLL 376
Query: 401 YGRNIDVWHSAVKKLRSFPHPRVQDN-------LKISYDSLDTMEKDIFLDIACFFKG-- 451
+ + WH L R++DN +S+ + K FL + F +
Sbjct: 377 SRKRTNEWHEVCASL----WRRLKDNSIHISTVFDLSFKEMRHELKLCFLYFSVFPEDYE 432
Query: 452 MKGDKVIDILESCGYFPQ-----------IGIQILIERSLITLDSVNN----KLGMHDLL 496
+K +K+I +L + G+ + I L++RSL+ + + +HDLL
Sbjct: 433 IKVEKLIHLLVAEGFIQEDEEMMMEDVARCYIDELVDRSLVKAERIERGKVMSCRIHDLL 492
Query: 497 QEM----GRDIVF------QESPNDPCRR 515
+++ +++ F ++ +D CRR
Sbjct: 493 RDLAIKKAKELNFVNVYNEKQHSSDICRR 521
>DR23_ARATH (O82484) Putative disease resistance protein At4g10780
Length = 892
Score = 77.4 bits (189), Expect = 1e-13
Identities = 84/309 (27%), Positives = 142/309 (45%), Gaps = 31/309 (10%)
Query: 219 IGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRN 278
+G++GMGG+GK+T+ ++ T+ + V +S + +Q + L
Sbjct: 176 MGLYGMGGVGKTTLLTQIHNTLHDTKNGVDIVIWV-VVSSDLQIHKIQEDIGEKLGFIGK 234
Query: 279 DFHDLYDGKKTIQ--NSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKH 336
+++ + +K + N L +K+ +L+LDD+ + L + +V+ TTR
Sbjct: 235 EWNKKQESQKAVDILNCLSKKRFVLLLDDIWKKVDLTKIGIPSQTRENKCKVVFTTRSLD 294
Query: 337 LLMTHGVHKTYKTGMLCKHDALVLFCLKAFK---GDKPQEGYLDLSKEVVDYCGGLPLAL 393
+ GVH + L +DA LF K + G P L+L+K+V C GLPLAL
Sbjct: 295 VCARMGVHDPMEVQCLSTNDAWELFQEKVGQISLGSHPD--ILELAKKVAGKCRGLPLAL 352
Query: 394 EVLGSYLYG-RNIDVWHSAVKKLRSF--PHPRVQDN----LKISYDSL-DTMEKDIFLDI 445
V+G + G R + WH AV L S+ + D+ LK SYD+L D + F
Sbjct: 353 NVIGETMAGKRAVQEWHHAVDVLTSYAAEFSGMDDHILLILKYSYDNLNDKHVRSCFQYC 412
Query: 446 ACFFK--GMKGDKVIDILESCGYFP---------QIGIQIL--IERSLITLDSVNNKL-- 490
A + + +K ++ID G+ G +IL + R+ + + NKL
Sbjct: 413 ALYPEDYSIKKYRLIDYWICEGFIDGNIGKERAVNQGYEILGTLVRACLLSEEGKNKLEV 472
Query: 491 GMHDLLQEM 499
MHD+++EM
Sbjct: 473 KMHDVVREM 481
>RGA3_SOLBU (Q7XA40) Putative disease resistance protein RGA3
(RGA1-blb) (Blight resistance protein B149)
Length = 947
Score = 76.3 bits (186), Expect = 2e-13
Identities = 102/412 (24%), Positives = 176/412 (41%), Gaps = 39/412 (9%)
Query: 215 DVRFIGIWGMGGIGKSTIARAVYETIRC--EFELTCFLENVREISETNGLVHLQRQLLSH 272
+V + I GMGG+GK+T+A+ V+ R F L ++ V + + L+ + +
Sbjct: 174 EVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWV-CVSDDFDEKRLIKAIVESIEG 232
Query: 273 LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDV--NELNQLENLVGKQDWFGPGSRVII 330
S+ D L +K +Q L K+ LVLDDV + + +NL G+ ++I
Sbjct: 233 KSLGDMDLAPL---QKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILI 289
Query: 331 TTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAF-KGDKPQEGYLDLSKEVVDYCGGL 389
TTR + + G + Y+ L + D +LF +AF + +++ KE+V CGG+
Sbjct: 290 TTRLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGV 349
Query: 390 PLALEVLGSYL-YGRNIDVW-HSAVKKLRSFPHPR--VQDNLKISYDSLDTMEKDIFLDI 445
PLA + LG L + R W H ++ + P V L++SY L + F
Sbjct: 350 PLAAKTLGGLLRFKREESEWEHVRDSEIWNLPQDENSVLPALRLSYHHLPLDLRQCFAYC 409
Query: 446 ACFFKGMKGDK--------VIDILESCG--YFPQIGIQILIERSL------ITLDSVNNK 489
A F K K +K L S G +G ++ E L I + S
Sbjct: 410 AVFPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQEIEVKSGKTY 469
Query: 490 LGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYE 549
MHDL+ ++ + + + R+ + ED+ ++T K +I ++
Sbjct: 470 FKMHDLIHDLATSMFSASASSRSIRQINVKDDEDMMFIVTNYKDMMSIGFSEV------V 523
Query: 550 AHWNTEAFSKTSQLKFLSLCEM---QLPLGLSCLPSSLKVLHWRGCPLKTLP 598
+ ++ F + L+ L+L QLP + L L+ L G + +LP
Sbjct: 524 SSYSPSLFKRFVSLRVLNLSNSEFEQLPSSVGDL-VHLRYLDLSGNKICSLP 574
>DRL3_ARATH (Q9LMP6) Probable disease resistance protein At1g15890
Length = 921
Score = 75.5 bits (184), Expect = 4e-13
Identities = 84/313 (26%), Positives = 144/313 (45%), Gaps = 37/313 (11%)
Query: 217 RFIGIWGMGGIGKSTIARAVYETI---RCEFELTCFLENVREISETNGLVHLQRQLLSHL 273
R +G++GMGG+GK+T+ ++ F+L ++ +++ +Q Q+L L
Sbjct: 245 RTLGLYGMGGVGKTTLLASINNKFLEGMNGFDLVIWVVVSKDLQNEG----IQEQILGRL 300
Query: 274 SISRNDFHDLYDGKKT--IQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIIT 331
+ R + + + +K I N L KK +L+LDD+ LE + GS+++ T
Sbjct: 301 GLHRG-WKQVTEKEKASYICNILNVKKFVLLLDDLWSEVDLEKIGVPPLTRENGSKIVFT 359
Query: 332 TRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLD---LSKEVVDYCGG 388
TR K + V K L +A LF K G P + + D L+++V + C G
Sbjct: 360 TRSKDVCRDMEVDGEMKVDCLPPDEAWELFQKKV--GPIPLQSHEDIPTLARKVAEKCCG 417
Query: 389 LPLALEVLGSYLYGR-NIDVWHSAVKKLRSFPH--PRVQDN----LKISYDSL-DTMEKD 440
LPLAL V+G + R + W + L S H P +++ LK SYD L D K
Sbjct: 418 LPLALSVIGKAMASRETVQEWQHVIHVLNSSSHEFPSMEEKILPVLKFSYDDLKDEKVKL 477
Query: 441 IFLDIACFFKG--MKGDKVIDILESCGYF----PQIG--------IQILIERSLITLDSV 486
FL + F + ++ +++I+ G+ + G I L+ L+ +
Sbjct: 478 CFLYCSLFPEDYEVRKEELIEYWMCEGFIDGNEDEDGANNKGHDIIGSLVRAHLLMDGEL 537
Query: 487 NNKLGMHDLLQEM 499
K+ MHD+++EM
Sbjct: 538 TTKVKMHDVIREM 550
>DRL5_ARATH (Q9C8K0) Probable disease resistance protein At1g51480
Length = 854
Score = 75.1 bits (183), Expect = 5e-13
Identities = 96/351 (27%), Positives = 159/351 (44%), Gaps = 41/351 (11%)
Query: 181 HRKLVPKLPSCT-ENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMGGIGKSTIARAV-- 236
H+ VPK+ VG+ + VE K L +ND +R + + GMGG+GK+T+ +
Sbjct: 139 HKIPVPKVEEKNIHTTVGLYAMVEMAWKSL---MNDEIRTLCLHGMGGVGKTTLLACINN 195
Query: 237 -YETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKT-IQNSL 294
+ + EF++ ++ +S+ L +Q Q+L L + + + + K + I N+L
Sbjct: 196 KFVELESEFDVVIWV----VVSKDFQLEGIQDQILGRLRLDKEWERETENKKASLINNNL 251
Query: 295 CRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCK 354
RKK +L+LDD+ L + G++++ T R K + K L
Sbjct: 252 KRKKFVLLLDDLWSEVDLNKIGVPPPTRENGAKIVFTKRSKEVSKYMKADMQIKVSCLSP 311
Query: 355 HDALVLFCLKAFKGDKPQEGYLD---LSKEVVDYCGGLPLALEVLGSYLYGR-NIDVWHS 410
+A LF + D + D L++ V C GLPLAL V+G + + I WH
Sbjct: 312 DEAWELFRITV--DDVILSSHEDIPALARIVAAKCHGLPLALIVIGEAMACKETIQEWHH 369
Query: 411 AVKKLRS-----FP--HPRVQDNLKISYDSLDTME-KDIFLDIACFFKG--MKGDKVIDI 460
A+ L S FP R+ LK SYDSL E K FL + F + ++ +K+I+
Sbjct: 370 AINVLNSPAGHKFPGMEERILLVLKFSYDSLKNGEIKLCFLYCSLFPEDFEIEKEKLIEY 429
Query: 461 LESCGY-----FPQIG-------IQILIERSLITLDSVNNKLGMHDLLQEM 499
GY + G I +L+ L+ + K+ MH +++EM
Sbjct: 430 WICEGYINPNRYEDGGTNQGYDIIGLLVRAHLLIECELTTKVKMHYVIREM 480
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.137 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,583,219
Number of Sequences: 164201
Number of extensions: 3351671
Number of successful extensions: 9487
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 9286
Number of HSP's gapped (non-prelim): 132
length of query: 640
length of database: 59,974,054
effective HSP length: 117
effective length of query: 523
effective length of database: 40,762,537
effective search space: 21318806851
effective search space used: 21318806851
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149204.4


