

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.13 - phase: 0
(163 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
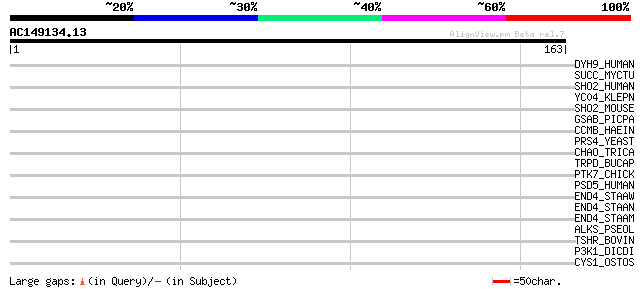
Sequences producing significant alignments: (bits) Value
DYH9_HUMAN (Q9NYC9) Ciliary dynein heavy chain 9 (Axonemal beta ... 33 0.28
SUCC_MYCTU (P71559) Succinyl-CoA synthetase beta chain (EC 6.2.1... 32 0.83
SHO2_HUMAN (Q9UQ13) Leucine-rich repeat protein SHOC-2 (Ras-bind... 32 0.83
YC04_KLEPN (Q48450) Putative capsule polysaccharide export prote... 30 2.4
SHO2_MOUSE (O88520) Leucine-rich repeat protein SHOC-2 (Ras-bind... 30 3.1
GSAB_PICPA (Q9HFR4) Pexophagy regulatory protein Gsa11 30 3.1
CCMB_HAEIN (P45033) Heme exporter protein B (Cytochrome c-type b... 30 3.1
PRS4_YEAST (P40327) 26S protease regulatory subunit 4 homolog (T... 29 4.1
CHAO_TRICA (P82963) Chaoptin (Photoreceptor cell-specific membra... 29 4.1
TRPD_BUCAP (P42392) Anthranilate phosphoribosyltransferase (EC 2... 29 5.4
PTK7_CHICK (Q91048) Tyrosine-protein kinase-like 7 precursor (Ki... 28 7.0
PSD5_HUMAN (Q16401) 26S proteasome non-ATPase regulatory subunit... 28 7.0
END4_STAAW (P63539) Probable endonuclease IV (EC 3.1.21.2) (Endo... 28 7.0
END4_STAAN (P63538) Probable endonuclease IV (EC 3.1.21.2) (Endo... 28 7.0
END4_STAAM (P63537) Probable endonuclease IV (EC 3.1.21.2) (Endo... 28 7.0
ALKS_PSEOL (P17051) Regulatory protein alkS 28 7.0
TSHR_BOVIN (Q27987) Thyrotropin receptor precursor (TSH-R) (Thyr... 28 9.1
P3K1_DICDI (P54673) Phosphatidylinositol 3-kinase 1 (EC 2.7.1.13... 28 9.1
CYS1_OSTOS (P25802) Cathepsin B-like cysteine proteinase 1 precu... 28 9.1
>DYH9_HUMAN (Q9NYC9) Ciliary dynein heavy chain 9 (Axonemal beta
dynein heavy chain 9)
Length = 4486
Score = 33.1 bits (74), Expect = 0.28
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 1/87 (1%)
Query: 56 PSFVFSSKTLSILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
P + FS K SI+ K + P +L A +D +TF+ Y+ ++ L C L YL
Sbjct: 3697 PMYQFSLKAFSIVFQKAVE-RAAPDESLRERVANLIDSITFSVYQYTIRGLFECDKLTYL 3755
Query: 116 GTNNLVVELPYSERPVISLSNLIRANI 142
+ L E + L L+R+ +
Sbjct: 3756 AQLTFQILLMNREVNAVELDFLLRSPV 3782
>SUCC_MYCTU (P71559) Succinyl-CoA synthetase beta chain (EC 6.2.1.5)
(SCS-beta)
Length = 387
Score = 31.6 bits (70), Expect = 0.83
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 7/62 (11%)
Query: 38 VENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQI------TLNEV-PFVNLPSLKALY 90
V+ +++DF+ S+ Q LP+ V + ++I KL ++ TL EV P V P K L
Sbjct: 144 VKGVDLDFARSIAEQGHLPAEVLDTAAVTIAKLWELFVAEDATLVEVNPLVRTPDHKILA 203
Query: 91 LD 92
LD
Sbjct: 204 LD 205
>SHO2_HUMAN (Q9UQ13) Leucine-rich repeat protein SHOC-2 (Ras-binding
protein Sur-8)
Length = 582
Score = 31.6 bits (70), Expect = 0.83
Identities = 31/100 (31%), Positives = 45/100 (45%), Gaps = 7/100 (7%)
Query: 51 SQMTLPSFVFSSKTLSILKLKQITLNEVPFV--NLPSLKALYLDVVTFTYYELILKLLSG 108
S +LP + + K L +L L+ L E+P V L SL LYL T E +K LS
Sbjct: 157 SLTSLPDSLDNLKKLRMLDLRHNKLREIPSVVYRLDSLTTLYLRFNRITTVEKDIKNLSK 216
Query: 109 CPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIE 148
+L N + +LP + L NLI ++ +E
Sbjct: 217 LSMLSI--RENKIKQLP---AEIGELCNLITLDVAHNQLE 251
>YC04_KLEPN (Q48450) Putative capsule polysaccharide export protein
precursor (ORF4)
Length = 378
Score = 30.0 bits (66), Expect = 2.4
Identities = 19/48 (39%), Positives = 27/48 (55%), Gaps = 3/48 (6%)
Query: 93 VVTFTYYELILKLLSGCPILQYLGTNNL---VVELPYSERPVISLSNL 137
+V F+ L + LSGC I+ G N+L VVELP S+ + L N+
Sbjct: 5 IVRFSALALAIGFLSGCTIVPGQGLNSLRKNVVELPDSDYDLDKLVNV 52
>SHO2_MOUSE (O88520) Leucine-rich repeat protein SHOC-2 (Ras-binding
protein Sur-8)
Length = 582
Score = 29.6 bits (65), Expect = 3.1
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 7/100 (7%)
Query: 51 SQMTLPSFVFSSKTLSILKLKQITLNEVPFV--NLPSLKALYLDVVTFTYYELILKLLSG 108
S +LP + + K L +L L+ L E+P V L SL LYL T E +K L
Sbjct: 157 SLTSLPDSLDNLKKLRMLDLRHNKLREIPSVVYRLDSLTTLYLRFNRITTVEKDIKNLPK 216
Query: 109 CPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIE 148
+L N + +LP + L NLI ++ +E
Sbjct: 217 LSMLSI--RENKIKQLP---AEIGELCNLITLDVAHNQLE 251
>GSAB_PICPA (Q9HFR4) Pexophagy regulatory protein Gsa11
Length = 1862
Score = 29.6 bits (65), Expect = 3.1
Identities = 31/106 (29%), Positives = 46/106 (43%), Gaps = 14/106 (13%)
Query: 13 FHLQCWNDYDCR---------DIYNFLYIAIQRGVENLNID-FSHSLFSQMTLPSFVFSS 62
F Q ++YD R D+YN + + GV NLN + +H L +PS + S
Sbjct: 472 FSFQPGSNYDVRFEMILDDDNDLYNITILCPKPGVFNLNEEVLNHLLSFCSKIPSTLHSL 531
Query: 63 KTLSILKLKQITL-NEVPFVNLPSLKALYLDVVTFTYYELILKLLS 107
++ K K +L N N +YL +F E IL+L S
Sbjct: 532 QSYQYQKAKLASLSNPQSNSNNDQENEIYLQTASF---EFILQLTS 574
>CCMB_HAEIN (P45033) Heme exporter protein B (Cytochrome c-type
biogenesis protein ccmB)
Length = 221
Score = 29.6 bits (65), Expect = 3.1
Identities = 18/71 (25%), Positives = 35/71 (48%), Gaps = 4/71 (5%)
Query: 50 FSQMTLPSFVFSSKTLSILKLKQIT----LNEVPFVNLPSLKALYLDVVTFTYYELILKL 105
F +L + +++ L + L ++ L +P + L + AL L + ++ L+L L
Sbjct: 74 FIDGSLEQLMLTAQPLPMTALAKVVAHWLLTGLPLILLSPIAALLLSLEVNIWWALVLTL 133
Query: 106 LSGCPILQYLG 116
L G P+L +G
Sbjct: 134 LLGTPVLSCIG 144
>PRS4_YEAST (P40327) 26S protease regulatory subunit 4 homolog
(TAT-binding homolog 5)
Length = 437
Score = 29.3 bits (64), Expect = 4.1
Identities = 20/73 (27%), Positives = 38/73 (51%), Gaps = 3/73 (4%)
Query: 65 LSILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVEL 124
LSI L++I ++ V P++ Y+ +++F EL L GC +L + T ++V L
Sbjct: 104 LSIGTLEEIIDDDHAIVTSPTMPDYYVSILSFVDKEL---LEPGCSVLLHHKTMSIVGVL 160
Query: 125 PYSERPVISLSNL 137
P++S+ +
Sbjct: 161 QDDADPMVSVMKM 173
>CHAO_TRICA (P82963) Chaoptin (Photoreceptor cell-specific membrane
protein) (Fragment)
Length = 782
Score = 29.3 bits (64), Expect = 4.1
Identities = 28/111 (25%), Positives = 49/111 (43%), Gaps = 16/111 (14%)
Query: 40 NLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFVNLPSLKALYL-------- 91
+LN+ + L ++ SF TL L L ++L++VP ++ P+L +L L
Sbjct: 422 SLNLGHNSQLTLEINGLSFQGLEYTLLHLNLDNVSLSQVPALSTPNLLSLSLAFNSLPTV 481
Query: 92 ------DVVTFTYYELILKLLSGCPILQYLGTNNLVVELPYSERPVISLSN 136
++ + Y L LS PI+ + T + L P+ +LSN
Sbjct: 482 ALEVAGNISSLRYLNLDYNDLSAVPIVTHSLTE--LRHLSLEGNPITTLSN 530
>TRPD_BUCAP (P42392) Anthranilate phosphoribosyltransferase (EC
2.4.2.18)
Length = 335
Score = 28.9 bits (63), Expect = 5.4
Identities = 28/74 (37%), Positives = 36/74 (47%), Gaps = 11/74 (14%)
Query: 25 DIYNFLYIAIQRGVEN-------LNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNE 77
D+ N I + +EN LNI F LF+ F ++SKT SILK+K I
Sbjct: 121 DLLNKFKINLNTSLENSKKILDKLNICF---LFAPKYHSVFKYASKTRSILKIKTIFNLL 177
Query: 78 VPFVNLPSLKALYL 91
PF+N PS L L
Sbjct: 178 GPFLN-PSRPPLTL 190
>PTK7_CHICK (Q91048) Tyrosine-protein kinase-like 7 precursor
(Kinase like protein)
Length = 1051
Score = 28.5 bits (62), Expect = 7.0
Identities = 12/31 (38%), Positives = 20/31 (63%), Gaps = 1/31 (3%)
Query: 125 PYSERPVISLSNLIRANICD-IHIEFDWLQN 154
PYS+ + S ++R + + H+EF+WLQN
Sbjct: 25 PYSQDALHGRSAILRCEVEEPAHVEFEWLQN 55
>PSD5_HUMAN (Q16401) 26S proteasome non-ATPase regulatory subunit 5
(26S proteasome subunit S5B) (26S protease subunit S5
basic)
Length = 503
Score = 28.5 bits (62), Expect = 7.0
Identities = 24/66 (36%), Positives = 36/66 (54%), Gaps = 4/66 (6%)
Query: 63 KTLSILKLKQITLNEVPFVNL-PSLKALYL--DVVTFTYYELILKLLSGCP-ILQYLGTN 118
K+LS + L Q L + NL LK++ D+V + YELI+++ S P L Y T+
Sbjct: 146 KSLSRISLTQAGLEALFESNLLDDLKSVMKTNDIVRYRVYELIIEISSVSPESLNYCTTS 205
Query: 119 NLVVEL 124
LV +L
Sbjct: 206 GLVTQL 211
>END4_STAAW (P63539) Probable endonuclease IV (EC 3.1.21.2)
(Endodeoxyribonuclease IV)
Length = 296
Score = 28.5 bits (62), Expect = 7.0
Identities = 15/58 (25%), Positives = 32/58 (54%), Gaps = 5/58 (8%)
Query: 103 LKLLSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFDWLQNVERLRA 160
L + G +++ G +N+VV PY +I+++N + ++ ++F Q +ER +A
Sbjct: 48 LNITKGHEVMEKYGLSNIVVHAPY----IINIANTTKPETFNLGVDF-LQQEIERTQA 100
>END4_STAAN (P63538) Probable endonuclease IV (EC 3.1.21.2)
(Endodeoxyribonuclease IV)
Length = 296
Score = 28.5 bits (62), Expect = 7.0
Identities = 15/58 (25%), Positives = 32/58 (54%), Gaps = 5/58 (8%)
Query: 103 LKLLSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFDWLQNVERLRA 160
L + G +++ G +N+VV PY +I+++N + ++ ++F Q +ER +A
Sbjct: 48 LNITKGHEVMEKYGLSNIVVHAPY----IINIANTTKPETFNLGVDF-LQQEIERTQA 100
>END4_STAAM (P63537) Probable endonuclease IV (EC 3.1.21.2)
(Endodeoxyribonuclease IV)
Length = 296
Score = 28.5 bits (62), Expect = 7.0
Identities = 15/58 (25%), Positives = 32/58 (54%), Gaps = 5/58 (8%)
Query: 103 LKLLSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFDWLQNVERLRA 160
L + G +++ G +N+VV PY +I+++N + ++ ++F Q +ER +A
Sbjct: 48 LNITKGHEVMEKYGLSNIVVHAPY----IINIANTTKPETFNLGVDF-LQQEIERTQA 100
>ALKS_PSEOL (P17051) Regulatory protein alkS
Length = 882
Score = 28.5 bits (62), Expect = 7.0
Identities = 17/64 (26%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Query: 91 LDVVTFTYYELILKLLSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFD 150
LD VT Y + K ++G ++YL TN +++ E +L ++R + E
Sbjct: 279 LDFVTPDQYNYVFKCVNGVSCIKYLSTNYMLLRHVSGEPAQFTLHPVLR----NFLREIT 334
Query: 151 WLQN 154
W +N
Sbjct: 335 WTEN 338
>TSHR_BOVIN (Q27987) Thyrotropin receptor precursor (TSH-R)
(Thyroid stimulating hormone receptor)
Length = 763
Score = 28.1 bits (61), Expect = 9.1
Identities = 14/50 (28%), Positives = 26/50 (52%)
Query: 44 DFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFVNLPSLKALYLDV 93
DF + ++PS S++TL ++ T+ F NLP++ +YL +
Sbjct: 36 DFRVTCKDIQSIPSLPPSTQTLKFIETHLKTIPSRAFSNLPNISRIYLSI 85
>P3K1_DICDI (P54673) Phosphatidylinositol 3-kinase 1 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K)
Length = 1570
Score = 28.1 bits (61), Expect = 9.1
Identities = 18/70 (25%), Positives = 32/70 (45%), Gaps = 6/70 (8%)
Query: 15 LQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQIT 74
L C D D + + I++ G+E FS + + P F F ++TLS+ + +
Sbjct: 867 LSCIKDIDSSSV--IVSISLYHGIEC----FSKAFTQPIIPPPFAFLAETLSVDWCEWLV 920
Query: 75 LNEVPFVNLP 84
+ + NLP
Sbjct: 921 FTNIDYSNLP 930
>CYS1_OSTOS (P25802) Cathepsin B-like cysteine proteinase 1
precursor (EC 3.4.22.-)
Length = 341
Score = 28.1 bits (61), Expect = 9.1
Identities = 16/53 (30%), Positives = 30/53 (56%), Gaps = 5/53 (9%)
Query: 106 LSGCPILQYLGTNNLVVE-----LPYSERPVISLSNLIRANICDIHIEFDWLQ 153
LSG P+++YL N + E +PY ++ ++ L + + NI D +E + L+
Sbjct: 33 LSGLPLVEYLQKNQRLFEVTATPVPYFKQRLMDLKYIDQNNIPDEEVEDEELE 85
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.142 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,710,005
Number of Sequences: 164201
Number of extensions: 684995
Number of successful extensions: 1901
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 1898
Number of HSP's gapped (non-prelim): 20
length of query: 163
length of database: 59,974,054
effective HSP length: 102
effective length of query: 61
effective length of database: 43,225,552
effective search space: 2636758672
effective search space used: 2636758672
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149134.13


