

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.1 - phase: 0
(555 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
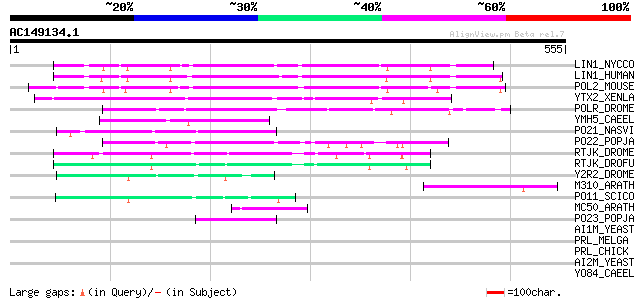
Score E
Sequences producing significant alignments: (bits) Value
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 164 7e-40
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 154 6e-37
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 150 8e-36
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 117 6e-26
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 92 5e-18
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 75 4e-13
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 75 4e-13
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 69 2e-11
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 67 9e-11
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 57 1e-07
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 55 5e-07
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 53 2e-06
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 52 3e-06
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 52 5e-06
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 50 1e-05
AI1M_YEAST (P03875) COX1/OXI3 intron 1 protein 42 0.003
PRL_MELGA (P17572) Prolactin precursor (PRL) 39 0.035
PRL_CHICK (P14676) Prolactin precursor (PRL) 38 0.061
AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein 37 0.10
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 37 0.13
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 164 bits (414), Expect = 7e-40
Identities = 133/455 (29%), Positives = 215/455 (47%), Gaps = 35/455 (7%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKN----GWLPNNF 99
+ E+ + + +L +PGPD F + F+Q + K+++ +L+ F+N G LPN F
Sbjct: 453 SSSEIASTIQNLPKKKSPGPDGFTSEFYQTF----KEELVPILLNLFQNIEKEGILPNTF 508
Query: 100 NANSIILIPKTPNADSV--DQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQ 157
+I LIPK P D + YR I+L+N KI+NK+L +R+ + + II +Q GF+
Sbjct: 509 YEANITLIPK-PGKDPTRKENYRPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIP 567
Query: 158 GR----NIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNE 213
G NIR I + IN L NK ++ L ID KAFD + F++ LK G
Sbjct: 568 GSQGWFNIRKSINVIQH-INKLKNKD---HMILSIDAEKAFDNIQHPFMIRTLKKIGIEG 623
Query: 214 LFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILAD 273
F I+ I I +NG + F G RQG PLSPLLF IV EVL +I+I +
Sbjct: 624 TFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEVL--AIAIREE 681
Query: 274 KGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSF 333
K + + S L + DD++V+ + S L + Y++ SG +N KS
Sbjct: 682 KAIKGIHIGSEEIKL---SLFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSV 738
Query: 334 IFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIH---FQPIADKVKAKLAK 390
F + + + + F V YLG + K K ++ ++ + ++ + K
Sbjct: 739 AFIYTNNNQAEKTVKDSIPFTVVPKKMKYLGVYLTK-DVKDLYKENYETLRKEIAEDVNK 797
Query: 391 WKASLLSIAGRIQLVKSVVQSMLVHTMSI--YSWPIKILKEMEKWIKNFIWSGDVTKRKM 448
WK S GRI +VK + ++ + P+ K++EK I +FIW+ +K
Sbjct: 798 WKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIWN-----QKK 852
Query: 449 VTVAWRKICADYEEGGLGVKSLICLNEATNLKICW 483
+A + + GG+ + L ++ +K W
Sbjct: 853 PQIAKTLLSNKNKAGGITLPDLRLYYKSIVIKTAW 887
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 154 bits (389), Expect = 6e-37
Identities = 130/465 (27%), Positives = 216/465 (45%), Gaps = 35/465 (7%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFF----KNGWLPNNF 99
T E++ + L + +PGP+ F A F+Q Y K+++ +L F K G LPN+F
Sbjct: 453 TSSEIEAIINSLPNKKSPGPEGFTAEFYQRY----KEELVPFLLKLFQSIEKEGILPNSF 508
Query: 100 NANSIILIPKTPNADSV--DQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQ 157
SIILIPK P D+ + +R I+L+N KI+NK+LA+++ + + +I +Q GF+
Sbjct: 509 YEASIILIPK-PGRDTTKKENFRPISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIP 567
Query: 158 GR----NIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNE 213
NIR I + D ++ + ID KAFD + F+L L G +
Sbjct: 568 AMQGWFNIRKSINIIQHINRTKDT----NHMIISIDAEKAFDKIQQPFMLKPLNKLGIDG 623
Query: 214 LFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILAD 273
+ I+ I I +NG + G RQG PLSPLL IV EVL+R+I +
Sbjct: 624 TYLKIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLSPLLPNIVLEVLARAIR--QE 681
Query: 274 KGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSF 333
K + + L + DD++V+ + + S L L + ++ SG +N++KS
Sbjct: 682 KEIKGIQLGKEEVKL---SLFADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQ 738
Query: 334 IFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGI--HFQPIADKVKAKLAKW 391
F + I++ L F + S YLG + + +++P+ +++K KW
Sbjct: 739 AFLYTNNRQTESQIMSELPFTIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKW 798
Query: 392 KASLLSIAGRIQLVKSVVQSMLVHTMSI--YSWPIKILKEMEKWIKNFIWSGDVTKRKMV 449
K S GRI +VK + +++ + P+ E+EK FIW+ +K
Sbjct: 799 KNIPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIWN-----QKRA 853
Query: 450 TVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSD--EQW 492
+A + + GG+ + +AT K W Q+ +QW
Sbjct: 854 HIAKSTLSQKNKAGGITLPDFKLYYKATVTKTAWYWYQNRDIDQW 898
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 150 bits (379), Expect = 8e-36
Identities = 141/496 (28%), Positives = 225/496 (44%), Gaps = 42/496 (8%)
Query: 19 DHLLVEEAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIV 78
D L +PKL + L + + KE ++ + L + +PGPD F A F+Q +
Sbjct: 456 DKFLDRYQVPKLNQDQVDHLNSPISPKE-IEAVINSLPTKKSPGPDGFSAEFYQTF---- 510
Query: 79 KKDVYEAVLDFFKN----GWLPNNFNANSIILIPKTPNAD--SVDQYRTIALVNFKFKII 132
K+D+ + F G LPN+F +I LIPK P D ++ +R I+L+N KI+
Sbjct: 511 KEDLIPILHKLFHKIEVEGTLPNSFYEATITLIPK-PQKDPTKIENFRPISLMNIDAKIL 569
Query: 133 NKVLADRLAKILPSIISKEQRGFVQGR----NIRDCIALTSEAINVLDNKSFGGNLALKI 188
NK+LA+R+ + + +II +Q GF+ G NIR I + IN L +K+ ++ + +
Sbjct: 570 NKILANRIQEHIKAIIHPDQVGFIPGMQGWFNIRKSINVI-HYINKLKDKN---HMIISL 625
Query: 189 DVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQ 248
D KAFD + F++ VL+ G + N IK I I +NG + G RQ
Sbjct: 626 DAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQ 685
Query: 249 GDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSS 308
G PLSP LF IV EVL+R+I + I + L DD++V+ +S
Sbjct: 686 GCPLSPYLFNIVLEVLARAIRQQKEIKGIQIGKEEVKISL-----LADDMIVYISDPKNS 740
Query: 309 LIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIF 368
L +L + + G +N KS F I F++ + YLG +
Sbjct: 741 TRELLNLINSFGEVVGYKINSNKSMAFLYTKNKQAEKEIRETTPFSIVTNNIKYLGVTLT 800
Query: 369 KGKPKGIH---FQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIK 425
K + K ++ F+ + ++K L +WK S GRI +VK + ++ + + PIK
Sbjct: 801 K-EVKDLYDKNFKSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFN--AIPIK 857
Query: 426 I----LKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKI 481
I E+E I F+W+ K +A + GG+ + L A +K
Sbjct: 858 IPTQFFNELEGAICKFVWN-----NKKPRIAKSLLKDKRTSGGITMPDLKLYYRAIVIKT 912
Query: 482 CWNLMQSD--EQWANI 495
W + +QW I
Sbjct: 913 AWYWYRDRQVDQWNRI 928
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 117 bits (294), Expect = 6e-26
Identities = 103/423 (24%), Positives = 199/423 (46%), Gaps = 18/423 (4%)
Query: 25 EAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYE 84
+ +P + + RL T + T +E+ A+ + + +PG D FFQ +W+ + D +
Sbjct: 432 DGLPVVSERRKERLETPI-TLDELSQALRLMPHNKSPGLDGLTIEFFQFFWDTLGPDFHR 490
Query: 85 AVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKIL 144
+ + FK G LP + + L+PK + + +R ++L++ +KI+ K ++ RL +L
Sbjct: 491 VLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVL 550
Query: 145 PSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGGNLA-LKIDVTKAFDTLNWDFLL 203
+I +Q V GR I D + L + ++ + G +LA L +D KAFD ++ +L+
Sbjct: 551 AEVIHPDQSYTVPGRTIFDNVFLIRDLLHFA--RRTGLSLAFLSLDQEKAFDRVDHQYLI 608
Query: 204 LVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEV 263
L+ + F F ++KT+ S++ + +N + RGVRQG PLS L+ + E
Sbjct: 609 GTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIE- 667
Query: 264 LSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCS 323
L K L L+ + + Y DD+++ + + L + YA S
Sbjct: 668 ---PFLCLLRKRLTGLVLKEPDMRVVLSA-YADDVILVAQ-DLVDLERAQECQEVYAAAS 722
Query: 324 GQIMNIRKSF-IFAGGITDTRMNNIVNILGFNVGSLPF--TYLGAPIFKGKPKGIHFQPI 380
+N KS + G + + + + + + YL A + P +F +
Sbjct: 723 SARINWSKSSGLLEGSLKVDFLPPAFRDISWESKIIKYLGVYLSAEEY---PVSQNFIEL 779
Query: 381 ADKVKAKLAKWK--ASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFI 438
+ V +L KWK A +LS+ GR ++ +V S + + + S + + ++++ + +F+
Sbjct: 780 EECVLTRLGKWKGFAKVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFL 839
Query: 439 WSG 441
W G
Sbjct: 840 WIG 842
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 91.7 bits (226), Expect = 5e-18
Identities = 99/417 (23%), Positives = 182/417 (42%), Gaps = 32/417 (7%)
Query: 93 GWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQ 152
G LP++ + IPKT A +R I++ + + +N +LA RL + Q
Sbjct: 389 GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSINW--DPRQ 446
Query: 153 RGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFN 212
RGF+ D + + +K F +DV+KAFD+L+ + L+ +G
Sbjct: 447 RGFLPTDGCADNATIVDLVLRH-SHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYGAP 505
Query: 213 ELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILA 272
+ F ++++ ++ +G F RGV+QGDPLSP+LF +V + L R+
Sbjct: 506 KGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRT----- 560
Query: 273 DKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKS 332
L I A N + + DDL++F + +M ++L + G +N K
Sbjct: 561 ---LPSEIGAKVGNAITNAAAFADDLVLFAETRMGLQVLLDKTLD-FLSIVGLKLNADKC 616
Query: 333 FIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPI-------ADKVK 385
F + ++ F VGS L + K GI+F A+ +
Sbjct: 617 FTVGIKGQPKQKCTVLEAQSFYVGSSEIPSL-KRTDEWKYLGINFTATGRVRCNPAEDIG 675
Query: 386 AKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFI--WSGDV 443
KL + + L R+ +++V+ L H +++ S I +L++ +K I+ ++ W +
Sbjct: 676 PKLQRLTKAPLKPQQRLFALRTVLIPQLYHKLALGSVAIGVLRKTDKLIRYYVRRW---L 732
Query: 444 TKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDEQWANIIRSRV 500
V +A+ + A + GGLG+ SL + L+ N+ +W ++ ++ V
Sbjct: 733 NLPLDVPIAF--VHAPPKSGGLGIPSLRWVAPMLRLRRLSNI-----KWPHLTQNEV 782
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 75.1 bits (183), Expect = 4e-13
Identities = 51/173 (29%), Positives = 81/173 (46%), Gaps = 6/173 (3%)
Query: 90 FKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIIS 149
F + +PN + II IPK N S YR I+L + +I+ +++ R+ ++S
Sbjct: 656 FSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLS 715
Query: 150 KEQRGFVQGRNIRDCIALTSEAINVLDN--KSFGGNLALKIDVTKAFDTLNWDFLLLVLK 207
Q GF+ N R C + +I++ + K+ L D KAFD ++ LL L
Sbjct: 716 PHQHGFL---NFRSCPSSLVRSISLYHSILKNEKSLDILFFDFAKAFDKVSHPILLKKLA 772
Query: 208 TFGFNELFCNWIKTILHSSKMFISMNG-AQHGFFNCNRGVRQGDPLSPLLFCI 259
FG ++L C+W K LH + +N + + GV QG PLLF +
Sbjct: 773 LFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFIL 825
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 75.1 bits (183), Expect = 4e-13
Identities = 60/231 (25%), Positives = 107/231 (45%), Gaps = 18/231 (7%)
Query: 47 EVKNAVFDLNSDD----------APGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLP 96
E+KN +++D+ A GPD WN + + + +G P
Sbjct: 311 EIKNLWRPISNDEIKEVEACKRTAAGPDGMTTTA----WNSIDECIKSLFNMIMYHGQCP 366
Query: 97 NNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFV 156
+ + +LIPK P +R +++ + + +++LA+R+ + ++ QR F+
Sbjct: 367 RRYLDSRTVLIPKEPGTMDPACFRPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFI 424
Query: 157 QGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFC 216
+ + +L S I K G +A+ +DV KAFD++ +L L+
Sbjct: 425 VADGVAENTSLLSAMIKEARMKIKGLYIAI-LDVKKAFDSVEHRSILDALRRKKLPLEMR 483
Query: 217 NWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLF-CIVEEVLSR 266
N+I + +SK + + + + RGVRQGDPLSPLLF C+++ VL R
Sbjct: 484 NYIMWVYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSPLLFNCVMDAVLRR 534
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 69.3 bits (168), Expect = 2e-11
Identities = 85/367 (23%), Positives = 162/367 (43%), Gaps = 47/367 (12%)
Query: 93 GWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQ 152
G +P + A LIPK + ++ +R I + + ++++++LA RL + + Q
Sbjct: 50 GHVPTPWTAMRTTLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVE--LHPAQ 107
Query: 153 RGF--VQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFG 210
+G+ + G + + T + K++ + +DV KAFDT++ + L+ G
Sbjct: 108 KGYARIDGTLVNSLLLDTYISSRREQRKTYN---VVSLDVRKAFDTVSHSSICRALQRLG 164
Query: 211 FNELFCNWIKTILHSSKMFISMN-GAQHGFFNCNRGVRQGDPLSPLLF-CIVEEVLSRSI 268
+E N+I L S I + G+Q RGV+QGDPLSP LF +++E+L
Sbjct: 165 IDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELL---C 221
Query: 269 SILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFT--RYADCSGQI 326
S+ + G+ I + L F DDL++ + + +++ +L T + G
Sbjct: 222 SLQSTPGIGGTIGEEKIPVLAF----ADDLLLL---EDNDVLLPTTLATVANFFRLRGMS 274
Query: 327 MNIRKSFIF----AGGITDTRMNNIVNI----LGFNVGSLPFTYLGAPIFKGKPKGIHFQ 378
+N +KS +GG+ R + + L F YLG HF
Sbjct: 275 LNAKKSVSISVAASGGVCIPRTKPFLRVDNVLLPVTDRMQTFRYLG-----------HFF 323
Query: 379 PIADKVKA---KLAKW----KASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEME 431
++ K L++W +A+ L ++ L++ V L++ + S + L++ +
Sbjct: 324 GLSGAAKPTVYNLSRWLKCVEAAPLKPEQKLSLIREHVVPKLLYGLQNPSVTARTLRDAD 383
Query: 432 KWIKNFI 438
K I+ +
Sbjct: 384 KLIRTTV 390
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 67.4 bits (163), Expect = 9e-11
Identities = 94/397 (23%), Positives = 163/397 (40%), Gaps = 40/397 (10%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKD---VYEAVLDFFKNGWLPNNFN 100
T EEVKN + L APG D+ ++ + + ++ +VLD G+ P +
Sbjct: 439 TLEEVKNLIAKLPLKKAPGEDLLDNRTIRLLPDQALQFLALIFNSVLDV---GYFPKAWK 495
Query: 101 ANSIILIPKTPNADS-VDQYRTIALVNFKFKIINKVLADRL--AKILPSIISKEQRGF-V 156
+ SII+I KT + VD YR +L+ KI+ +++ +RL K + I K Q GF +
Sbjct: 496 SASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTCKDVTKAIPKFQFGFRL 555
Query: 157 QGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFC 216
Q + + A+ ++NK + + +D+ +AFD + LL K +L+
Sbjct: 556 QHGTPEQLHRVVNFALEAMENKEYA--VGAFLDIQQAFDRVWHPGLLYKAKRLFPPQLYL 613
Query: 217 NWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGL 276
+K+ L +S++G + GV QG L P L+ + + +
Sbjct: 614 V-VKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSVFASDMPTHTPVTEVDEE 672
Query: 277 IDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQ---IMNIRKSF 333
LIA Y DD V K+K S++ S Y D Q N+R +
Sbjct: 673 DVLIAT-----------YADDTAVLTKSK--SILAATSGLQEYLDAFQQWAENWNVRINA 719
Query: 334 IFAGGITDTRMNNIVNILGFNVGSL-----PFTYLGAPIFKGKPKGIHFQPIADKVKAKL 388
+T + N G L + YLG + + H I + K+
Sbjct: 720 EKCANVTFANRTGSCPGVSLN-GRLIRHHQAYKYLGITLDRKLTFSRHITNIQQAFRTKV 778
Query: 389 AK--W---KASLLSIAGRIQLVKSVVQSMLVHTMSIY 420
A+ W + LS+ ++ + KS++ L + + +Y
Sbjct: 779 ARMSWLIAPRNKLSLGCKVNIYKSILAPCLFYGLQVY 815
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 57.4 bits (137), Expect = 1e-07
Identities = 87/393 (22%), Positives = 154/393 (39%), Gaps = 32/393 (8%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
T EEVK V L APG D+ ++ + + + G+ P S
Sbjct: 439 TLEEVKELVSKLKPKKAPGEDLLDNRTIRLLPDQALLYLVLIFNSILRVGYFPKARPTAS 498
Query: 104 IILIPKTPNAD-SVDQYRTIALVNFKFKIINKVLADRL--AKILPSIISKEQRGF-VQGR 159
II+I K VD YR +L+ K++ +++ +R+ ++ + I K Q GF +Q
Sbjct: 499 IIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRILTSEEVTRAIPKFQFGFRLQHG 558
Query: 160 NIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWI 219
+ + A+ L+ K + G+ L D+ +AFD + LL K+ +LF I
Sbjct: 559 TPEQLHRVVNFALEALEKKEYAGSCFL--DIQQAFDRVWHPGLLYKAKSLLSPQLF-QLI 615
Query: 220 KTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDL 279
K+ K ++ +G + GV QG L P L+ I + ++ L
Sbjct: 616 KSFWEGRKFSVTADGCRSSVKFIEAGVPQGSVLGPTLYSIFTADMPNQNAVTGLAEGEVL 675
Query: 280 IAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGI 339
IA Y DD+ V K+ + ++ Y D + I AG
Sbjct: 676 IAT-----------YADDIAVLTKS--TCIVEATDALQEYLDAFQEWAVKWNVSINAGKC 722
Query: 340 TDTRMNNIV-NILGFNV-GSL-----PFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWK 392
+ N + + G + GSL + YLG + + H + + ++ K
Sbjct: 723 ANVTFTNAIRDCPGVTINGSLLSHTHEYKYLGVILDRSLTFRRHITSLQHSFRTRITKMN 782
Query: 393 ASL-----LSIAGRIQLVKSVVQSMLVHTMSIY 420
L LS+ ++++ K +V L + + +Y
Sbjct: 783 WLLAARNKLSLDNKVKIYKCIVAPGLFYAIQVY 815
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 55.1 bits (131), Expect = 5e-07
Identities = 50/225 (22%), Positives = 90/225 (39%), Gaps = 19/225 (8%)
Query: 47 EVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIIL 106
EV V L S +PG D + W + + + + G+ P + ++
Sbjct: 438 EVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPRVVS 497
Query: 107 IPKTPNADSVD--QYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDC 164
+ K P+ D + YR I L+ K++ ++ +R+ ++LP + Q GF QGR + D
Sbjct: 498 LLKGPDKDKCEPSSYRGICLLPVFGKVLEAIMVNRVREVLPE-GCRWQFGFRQGRCVEDA 556
Query: 165 IALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNEL-----FCNWI 219
++ + G +D AFD + W L L G E+ F +
Sbjct: 557 WRHVKSSVGASAAQYVLGTF---VDFKGAFDNVEWSAALSRLADLGCREMGLWQSFFSGR 613
Query: 220 KTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVL 264
+ ++ SS + + RG QG P ++ I+ +VL
Sbjct: 614 RAVIRSSSGTVEV--------PVTRGCPQGSISGPFIWDILMDVL 650
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 53.1 bits (126), Expect = 2e-06
Identities = 37/139 (26%), Positives = 60/139 (42%), Gaps = 5/139 (3%)
Query: 414 VHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYE-EGGLGVKSLIC 472
V+ MS + + K++ + F WS KRK+ VAW+K+C E +GGLG + L
Sbjct: 5 VYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRDLGW 64
Query: 473 LNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCMNHHIS---SSIWSGVKAEFHVIKD 528
N+A K + ++ Q + ++RSR M + S W + ++
Sbjct: 65 FNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGRELLSR 124
Query: 529 NCSWIVGDGKQINFWSDSW 547
+GDG W D W
Sbjct: 125 GLLRTIGDGIHTKVWLDRW 143
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 52.4 bits (124), Expect = 3e-06
Identities = 58/250 (23%), Positives = 98/250 (39%), Gaps = 16/250 (6%)
Query: 46 EEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSII 105
+EV ++V +PGPD + W + + ++ + P + S++
Sbjct: 408 DEVSDSVRRCKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLYKQCLLESYFPQKWKIASLV 467
Query: 106 LIPKTPNADSVD--QYRTIALVNFKFKIINKVLADRL-AKILPSIISKEQRGFVQGRNIR 162
++ K + D YR I L++ K++ ++ RL K++ +S Q F G++
Sbjct: 468 ILLKLLDRIRSDPGSYRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKSTE 527
Query: 163 DCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTI 222
D + + K G L ID AFD L W +LL L NE F W+
Sbjct: 528 DAWRCVQRHVECSEMKYVIG---LNIDFQGAFDNLGWLSMLLKLDEAQSNE-FGLWMSYF 583
Query: 223 LHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRS-------ISILADKG 275
++ G + RG QG P ++ +V L + I AD G
Sbjct: 584 GGRKVYYVGKTGIVRK--DVTRGCPQGSKSGPAMWKLVMNELLLALVAAGFFIVAFADDG 641
Query: 276 LIDLIAASRN 285
I + A SR+
Sbjct: 642 TIVIGANSRS 651
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 51.6 bits (122), Expect = 5e-06
Identities = 30/76 (39%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Query: 222 ILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIA 281
+ H +FI +NGA G +RG+RQGDPLSP LF + EVLS ++G + I
Sbjct: 5 VCHFYLLFI-INGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIR 63
Query: 282 ASRNNCLPFHCFYVDD 297
S N+ H + DD
Sbjct: 64 VSNNSPRINHLLFADD 79
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 50.4 bits (119), Expect = 1e-05
Identities = 27/82 (32%), Positives = 49/82 (58%), Gaps = 1/82 (1%)
Query: 186 LKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRG 245
+ +D+ KAFDT++ +L ++ FG ++ ++I + + + I + G G
Sbjct: 26 VSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNIVVGGRTTNKIYIRNG 85
Query: 246 VRQGDPLSPLLFCIV-EEVLSR 266
V+QGDPLSP+LF IV +E+++R
Sbjct: 86 VKQGDPLSPVLFNIVLDELVTR 107
>AI1M_YEAST (P03875) COX1/OXI3 intron 1 protein
Length = 834
Score = 42.4 bits (98), Expect = 0.003
Identities = 37/141 (26%), Positives = 65/141 (45%), Gaps = 12/141 (8%)
Query: 120 RTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKS 179
R +++ N + KI+ +V+ L I +S GF R C E N+
Sbjct: 323 RPLSVGNPRDKIVQEVMRMILDTIFDKKMSTHSHGF---RKNMSCQTAIWEVRNMFG--- 376
Query: 180 FGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGF 239
G N +++D+ K FDT++ D ++ LK + ++ F + + +L + +I G H
Sbjct: 377 -GSNWFIEVDLKKCFDTISHDLIIKELKRYISDKGFIDLVYKLLRAG--YIDEKGTYH-- 431
Query: 240 FNCNRGVRQGDPLSPLLFCIV 260
G+ QG +SP+L IV
Sbjct: 432 -KPMLGLPQGSLISPILCNIV 451
>PRL_MELGA (P17572) Prolactin precursor (PRL)
Length = 229
Score = 38.9 bits (89), Expect = 0.035
Identities = 34/151 (22%), Positives = 67/151 (43%), Gaps = 21/151 (13%)
Query: 363 LGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSW 422
L P K + + IH + + + + L W L+ +A +Q +K ++L + I
Sbjct: 93 LTTPEDKEQTQQIHHEELLNLILGVLRSWNDPLIHLASEVQRIKEAPDTILWKAVEIEEQ 152
Query: 423 PIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKIC 482
++L+ MEK I I SGD A ++ + ++ G+ SL +E + L
Sbjct: 153 NKRLLEGMEK-IVGRIHSGD---------AGNEVFSQWD----GLPSLQLADEDSRLFAF 198
Query: 483 WNLM-------QSDEQWANIIRSRVIRDHRC 506
+NL+ + + +++ R+I D+ C
Sbjct: 199 YNLLHCLRRDSHKIDNYLKVLKCRLIHDNNC 229
>PRL_CHICK (P14676) Prolactin precursor (PRL)
Length = 229
Score = 38.1 bits (87), Expect = 0.061
Identities = 35/151 (23%), Positives = 66/151 (43%), Gaps = 21/151 (13%)
Query: 363 LGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSW 422
L P K + + IH + + + V L W L+ +A +Q +K ++L + I
Sbjct: 93 LTTPEDKEQAQQIHHEDLLNLVVGVLRSWNDPLIHLASEVQRIKEAPDTILWKAVEIEEQ 152
Query: 423 PIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKIC 482
++L+ MEK I + SGD A +I + ++ G+ SL +E + L
Sbjct: 153 NKRLLEGMEK-IVGRVHSGD---------AGNEIYSHWD----GLPSLQLADEDSRLFAF 198
Query: 483 WNLM-------QSDEQWANIIRSRVIRDHRC 506
+NL+ + + +++ R+I D C
Sbjct: 199 YNLLHCLRRDSHKIDNYLKVLKCRLIHDSNC 229
>AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein
Length = 789
Score = 37.4 bits (85), Expect = 0.10
Identities = 39/160 (24%), Positives = 70/160 (43%), Gaps = 17/160 (10%)
Query: 107 IPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIA 166
IPKT +R +++ N + KI+ + + L I + S GF R C+
Sbjct: 283 IPKTSGG-----FRPLSVGNPREKIVQESMRMMLEIIYNNSFSYYSHGF---RPNLSCLT 334
Query: 167 LTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSS 226
+ N + N +K+D+ K FDT+ + L+ VL ++ F + + +L +
Sbjct: 335 AIIQCKNYMQYC----NWFIKVDLNKCFDTIPHNMLINVLNERIKDKGFMDLLYKLLRAG 390
Query: 227 KMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSR 266
++ N H N G+ QG +SP+L I + L +
Sbjct: 391 --YVDKNNNYH---NTTLGIPQGSVVSPILCNIFLDKLDK 425
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 37.0 bits (84), Expect = 0.13
Identities = 22/83 (26%), Positives = 38/83 (45%)
Query: 188 IDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVR 247
+D +KAFD ++ D LL L + N+ W+ L + + + GV
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 248 QGDPLSPLLFCIVEEVLSRSISI 270
QG +SP+LF I +S ++ +
Sbjct: 83 QGSVISPVLFGIFVNEISANLPV 105
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,179,625
Number of Sequences: 164201
Number of extensions: 2657310
Number of successful extensions: 6908
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 6859
Number of HSP's gapped (non-prelim): 41
length of query: 555
length of database: 59,974,054
effective HSP length: 115
effective length of query: 440
effective length of database: 41,090,939
effective search space: 18080013160
effective search space used: 18080013160
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC149134.1


