

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149129.9 + phase: 0
(388 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
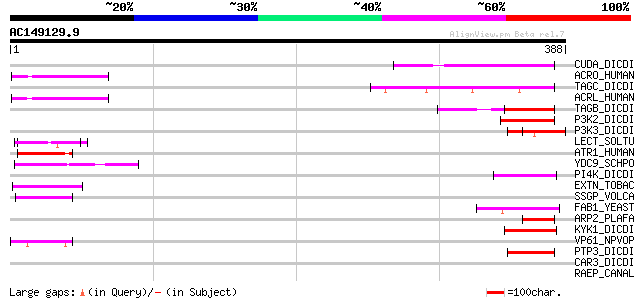
Score E
Sequences producing significant alignments: (bits) Value
CUDA_DICDI (O00841) Putative transcriptional regulator cudA 55 3e-07
ACRO_HUMAN (P10323) Acrosin precursor (EC 3.4.21.10) 50 1e-05
TAGC_DICDI (Q23868) Prestalk-specific protein tagC precursor (EC... 48 5e-05
ACRL_HUMAN (P58840) Hypothetical acrosin-like protease (EC 3.4.2... 47 6e-05
TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC... 47 8e-05
P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.13... 47 1e-04
P3K3_DICDI (P54675) Phosphatidylinositol 3-kinase 3 (EC 2.7.1.13... 46 1e-04
LECT_SOLTU (Q9S8M0) Chitin-binding lectin 1 precursor (PL-I) 46 1e-04
ATR1_HUMAN (Q9H6X2) Anthrax toxin receptor 1 precursor (Tumor en... 46 2e-04
YDC9_SCHPO (Q10172) Hypothetical protein C25G10.09c in chromosome I 45 2e-04
PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67) ... 45 2e-04
EXTN_TOBAC (P13983) Extensin precursor (Cell wall hydroxyproline... 45 2e-04
SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor ... 45 3e-04
FAB1_YEAST (P34756) 1-phosphatidylinositol-3-phosphate 5-kinase ... 45 4e-04
ARP2_PLAFA (P13824) Clustered-asparagine-rich protein (Fragment) 45 4e-04
KYK1_DICDI (P18160) Non-receptor tyrosine kinase spore lysis A (... 44 5e-04
VP61_NPVOP (O10270) 61 kDa protein homolog 44 7e-04
PTP3_DICDI (P54637) Protein-tyrosine phosphatase 3 (EC 3.1.3.48)... 44 7e-04
CAR3_DICDI (P35352) Cyclic AMP receptor 3 44 0.001
RAEP_CANAL (O93831) Rab proteins geranylgeranyltransferase compo... 43 0.001
>CUDA_DICDI (O00841) Putative transcriptional regulator cudA
Length = 791
Score = 55.1 bits (131), Expect = 3e-07
Identities = 29/113 (25%), Positives = 54/113 (47%), Gaps = 7/113 (6%)
Query: 269 VLGKRERERDVERERERDPIGEMVNAIKVLRDGFVRMEQMKMEMAREIETMRMEMEMKRT 328
V+ K ++ + ++R+R P +++ + R+E + E R ++ + +
Sbjct: 309 VVSKPDQVKKKAKKRKRAPTDSLMDTLN-------RIEHQQKEQQRLLKKLCYHDKENNI 361
Query: 329 EMILESQQRIVEAFAKAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+++ QQ+ + + N + N NNN+NN NNN NNNNNNNN
Sbjct: 362 IQLIQQQQQQQQLLNNVTNNINNNNNINNNNNNNNNNNNNNNNNNNNNNNNNN 414
Score = 41.2 bits (95), Expect = 0.005
Identities = 18/26 (69%), Positives = 20/26 (76%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIAS 384
N NNN+NN NNN +NNNNNNN AS
Sbjct: 89 NANNNNNNNNSNNNNSNNNNNNNNAS 114
Score = 40.4 bits (93), Expect = 0.008
Identities = 16/23 (69%), Positives = 19/23 (82%)
Query: 362 NNNDNNIINNNGNNNNNNNNIAS 384
NNN+NN NNN NNNNNNNN ++
Sbjct: 93 NNNNNNSNNNNSNNNNNNNNASN 115
Score = 37.4 bits (85), Expect = 0.066
Identities = 16/30 (53%), Positives = 20/30 (66%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N NN++NN NNN NNNN +NN+ S S
Sbjct: 94 NNNNNSNNNNSNNNNNNNNASNNLTSNKSS 123
Score = 36.2 bits (82), Expect = 0.15
Identities = 15/23 (65%), Positives = 18/23 (78%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N IN N+NN NN+ NNN+NNNN
Sbjct: 86 NCINANNNNNNNNSNNNNSNNNN 108
Score = 33.9 bits (76), Expect = 0.72
Identities = 15/30 (50%), Positives = 20/30 (66%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N N+N+NN NNN NNN +NN ++ S S
Sbjct: 95 NNNNSNNNNSNNNNNNNNASNNLTSNKSSS 124
Score = 33.9 bits (76), Expect = 0.72
Identities = 15/32 (46%), Positives = 19/32 (58%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
N G+T++ N N NNN NNN+NNNN
Sbjct: 72 NSPNGTTNGSTMSPNCINANNNNNNNNSNNNN 103
Score = 31.2 bits (69), Expect = 4.7
Identities = 16/42 (38%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Query: 347 SEKNKKRKLRKGNT-INNNDNNIINNNGNNNNNNNNIASPSE 387
+ +N + ++ NT NNN+NNIINN N+ N N I + +
Sbjct: 442 NNENTEHIVKIENTECNNNNNNIINNTENDENINKPILNSKD 483
>ACRO_HUMAN (P10323) Acrosin precursor (EC 3.4.21.10)
Length = 421
Score = 50.1 bits (118), Expect = 1e-05
Identities = 27/68 (39%), Positives = 35/68 (50%), Gaps = 2/68 (2%)
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSYRDK 61
A PPP SP P PP A PLP P PPP P+S+ +LP + LI+ + K
Sbjct: 341 AAQPPPPPSPPP--PPPPPASPLPPPPPPPPPTPSSTTKLPQGLSFAKRLQQLIEVLKGK 398
Query: 62 WYSLGRTN 69
YS G+ +
Sbjct: 399 TYSDGKNH 406
Score = 40.0 bits (92), Expect = 0.010
Identities = 21/47 (44%), Positives = 24/47 (50%), Gaps = 2/47 (4%)
Query: 7 PPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSA 53
PP P P PP+A P P P PPPPP +S PPP P S+
Sbjct: 330 PPPRPLP--PRPPAAQPPPPPSPPPPPPPPASPLPPPPPPPPPTPSS 374
>TAGC_DICDI (Q23868) Prestalk-specific protein tagC precursor (EC
3.4.21.-)
Length = 1743
Score = 47.8 bits (112), Expect = 5e-05
Identities = 44/144 (30%), Positives = 66/144 (45%), Gaps = 15/144 (10%)
Query: 253 SGSRINGGT--RLVKERVVLGKRERERDVERERERDPIGE--MVNAIKVLRDGF-VRMEQ 307
+GS ++GG R+ R + KR+ E E DP E + +IKVL G V M
Sbjct: 1586 AGSLLSGGQKKRIAIARAICAKRKIMLLDEITAELDPESEEAITQSIKVLTQGHTVVMVA 1645
Query: 308 MKMEMAREIETMRME-----MEMKRTEMILESQQRIVEAFAKAISEKNKKRKL-----RK 357
K+ R+ + + + +E + ++E + + F E+ L +
Sbjct: 1646 HKVAAVRDCDKIFVLDKGYLVEEGTHDELMERKGKYHRMFNNEKDEEELLNNLGLPSNNE 1705
Query: 358 GNTINNNDNNIINNNGNNNNNNNN 381
N NNN+NN NNN NNNNNNNN
Sbjct: 1706 TNNENNNENNNNNNNNNNNNNNNN 1729
Score = 35.0 bits (79), Expect = 0.33
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 10/46 (21%)
Query: 346 ISEKNKKRKLRKGNT-INNNDNN---------IINNNGNNNNNNNN 381
I + N K ++ IN+NDNN I+NNN NNNNNN +
Sbjct: 60 IIDTNNKPSIKNNQIFINDNDNNENISPILLRILNNNNNNNNNNGD 105
>ACRL_HUMAN (P58840) Hypothetical acrosin-like protease (EC
3.4.21.-) (Fragment)
Length = 232
Score = 47.4 bits (111), Expect = 6e-05
Identities = 26/68 (38%), Positives = 34/68 (49%), Gaps = 3/68 (4%)
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSYRDK 61
A PPPS P P PP PLP P PPP P+S+ +LP + LI+ + K
Sbjct: 153 AQPRPPPSPPP---PPPPPPSPLPPPPPPPPPTPSSTTKLPQGLSFAKRLQQLIEVLKGK 209
Query: 62 WYSLGRTN 69
YS G+ +
Sbjct: 210 TYSDGKNH 217
Score = 39.7 bits (91), Expect = 0.013
Identities = 21/47 (44%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Query: 7 PPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSA 53
PP P P PP+A P P P PPPPP S PPP P S+
Sbjct: 141 PPPRPLP--PRPPAAQPRPPPSPPPPPPPPPSPLPPPPPPPPPTPSS 185
>TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC
3.4.21.-)
Length = 1905
Score = 47.0 bits (110), Expect = 8e-05
Identities = 20/35 (57%), Positives = 22/35 (62%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N KL N NNN+NN NNN NNNNNNNN
Sbjct: 98 NNNNNNNKLNNNNNNNNNNNNNNNNNNNNNNNNNN 132
Score = 44.7 bits (104), Expect = 4e-04
Identities = 26/83 (31%), Positives = 42/83 (50%), Gaps = 10/83 (12%)
Query: 300 DGFVRMEQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAK-AISEKNKKRKLRKG 358
D + +Q + + + +I + ++ +K +I++ K +I+ N K
Sbjct: 56 DSIYKNKQQQQQFSNKIYSNEKKILLKN---------KIIDTTIKPSININNNNNNNNKL 106
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N NNN+NN NNN NNNNNNNN
Sbjct: 107 NNNNNNNNNNNNNNNNNNNNNNN 129
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/35 (54%), Positives = 21/35 (59%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N K N NNN+NN NNN NNNNNNNN
Sbjct: 100 NNNNNKLNNNNNNNNNNNNNNNNNNNNNNNNNNNN 134
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 21/35 (59%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N + N NNN+NN NNN NNNNNNNN
Sbjct: 99 NNNNNNKLNNNNNNNNNNNNNNNNNNNNNNNNNNN 133
Score = 37.7 bits (86), Expect = 0.050
Identities = 44/184 (23%), Positives = 73/184 (38%), Gaps = 29/184 (15%)
Query: 221 GSGSGFRIRIPTGVSVAQPGSKFYP---------KMNNNSESGSRINGGT--RLVKERVV 269
G G+ I PT + + Y K + ++GG R+ R +
Sbjct: 1614 GENIGYAIDNPTQEDIIEAAKLAYAHEFINDLPKKYDTQIGEAGNLSGGQKKRIAVARAI 1673
Query: 270 LGKRERERDVERERERDPIGE--MVNAIKVLRDGF-VRMEQMKMEMAREIETMRMEMEMK 326
KR+ E E DP E + +IKVL G V M K+ R+ + +
Sbjct: 1674 CAKRKIMLLDEITAELDPESEEAITQSIKVLTQGHTVVMVAHKVAAVRDCDKI------- 1726
Query: 327 RTEMILESQQRIVEA-FAKAISEKNKKRKL----RKGNTINNNDNNIINNNGNNNNNNNN 381
+LE + E + ++ K K ++ + + NN+NN NNN NNNN ++
Sbjct: 1727 ---FVLEKGYLVEEGTHDELMANKGKYYRMFSEDKDDTPLQNNNNNKNNNNNNNNNEPSS 1783
Query: 382 IASP 385
++P
Sbjct: 1784 SSTP 1787
Score = 35.4 bits (80), Expect = 0.25
Identities = 16/33 (48%), Positives = 18/33 (54%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
N N NNN+NN NNN NNNN N+I
Sbjct: 107 NNNNNNNNNNNNNNNNNNNNNNNNNNNNYYNSI 139
>P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K)
Length = 1858
Score = 46.6 bits (109), Expect = 1e-04
Identities = 21/38 (55%), Positives = 24/38 (62%)
Query: 344 KAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
K + NK+ K N NNN+NN NNN NNNNNNNN
Sbjct: 1000 KENKDSNKENKDSSSNNNNNNNNNNNNNNNNNNNNNNN 1037
Score = 45.1 bits (105), Expect = 3e-04
Identities = 20/38 (52%), Positives = 24/38 (62%)
Query: 344 KAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
K +++NK N NNN+NN NNN NNNNNNNN
Sbjct: 1003 KDSNKENKDSSSNNNNNNNNNNNNNNNNNNNNNNNNNN 1040
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/38 (50%), Positives = 21/38 (55%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIAS 384
+ N N NNN+NN NNN NNNNNNNN S
Sbjct: 192 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTTS 229
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/35 (54%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
S N N NNN+NN NNN NNNNNNNN
Sbjct: 184 SNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 218
Score = 43.1 bits (100), Expect = 0.001
Identities = 21/55 (38%), Positives = 33/55 (59%), Gaps = 2/55 (3%)
Query: 329 EMILESQQRIVEAFA--KAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ I + QQ+ ++ K +++NK ++ +NN+NN NNN NNNNNNNN
Sbjct: 979 QSIQQQQQQQIQTVINIKETNKENKDSNKENKDSSSNNNNNNNNNNNNNNNNNNN 1033
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 189 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 223
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 187 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 221
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 186 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 220
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 183 TSNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 217
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 191 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 225
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 190 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 224
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 185 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 219
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 188 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 222
Score = 41.2 bits (95), Expect = 0.005
Identities = 23/81 (28%), Positives = 40/81 (48%), Gaps = 3/81 (3%)
Query: 306 EQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEKNKKRKLRKGNTINNND 365
+ ++ + ++I+T+ + +K T + + + + + N N NNN+
Sbjct: 979 QSIQQQQQQQIQTV---INIKETNKENKDSNKENKDSSSNNNNNNNNNNNNNNNNNNNNN 1035
Query: 366 NNIINNNGNNNNNNNNIASPS 386
NN NNNGNNN NN+N S S
Sbjct: 1036 NNNNNNNGNNNGNNSNNNSNS 1056
Score = 41.2 bits (95), Expect = 0.005
Identities = 18/35 (51%), Positives = 19/35 (53%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
S N N NNN+NN NNN NNNN NNN
Sbjct: 1012 SSSNNNNNNNNNNNNNNNNNNNNNNNNNNNNGNNN 1046
Score = 40.4 bits (93), Expect = 0.008
Identities = 17/38 (44%), Positives = 21/38 (54%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIAS 384
+ N N NNN+NN NNN NNNNNNN ++
Sbjct: 193 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTTST 230
Score = 40.0 bits (92), Expect = 0.010
Identities = 20/45 (44%), Positives = 22/45 (48%)
Query: 344 KAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
K S N N NNN+NN NNN NNN NNN S + S
Sbjct: 1010 KDSSSNNNNNNNNNNNNNNNNNNNNNNNNNNNNGNNNGNNSNNNS 1054
Score = 39.3 bits (90), Expect = 0.017
Identities = 16/22 (72%), Positives = 18/22 (81%)
Query: 360 TINNNDNNIINNNGNNNNNNNN 381
T +NN+NN NNN NNNNNNNN
Sbjct: 182 TTSNNNNNNNNNNNNNNNNNNN 203
Score = 37.7 bits (86), Expect = 0.050
Identities = 19/42 (45%), Positives = 24/42 (56%), Gaps = 2/42 (4%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNN--NNNNNNIASPS 386
+ N N NNN+NN NNNGNN NN+N+NI+ S
Sbjct: 1021 NNNNNNNNNNNNNNNNNNNNNNGNNNGNNSNNNSNSNISRGS 1062
Score = 37.0 bits (84), Expect = 0.086
Identities = 16/42 (38%), Positives = 21/42 (49%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
+ N N NNN+NN NNN NNNNN + + + S
Sbjct: 195 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTTSTTTTTTS 236
Score = 31.6 bits (70), Expect = 3.6
Identities = 28/131 (21%), Positives = 57/131 (43%), Gaps = 17/131 (12%)
Query: 111 RSEIQKLRSLPVPRSRSSSSWVLFKAMDSMEKGPSPPSPPHKPENPNHNANHNHIHSHNL 170
RS + S P+ + +SSS + + PSP + + N N+N N+N+ +++N
Sbjct: 149 RSRSGSIGSKPICNNLTSSS----SSSSTTATTPSPTTTSNNNNNNNNNNNNNNNNNNNN 204
Query: 171 INHRETLENDDYDDDDLYEE-------------LRSAAGGGSGSGNTRSLDKLYRNGVSG 217
N+ N++ ++++ L S++ S S ++ S D+ + N +
Sbjct: 205 NNNNNNNNNNNNNNNNNNNNNNTTSTTTTTTSILISSSPPPSSSSSSSSNDEQFNNNNNN 264
Query: 218 GFGGSGSGFRI 228
SG R+
Sbjct: 265 NNSNSGGSSRM 275
>P3K3_DICDI (P54675) Phosphatidylinositol 3-kinase 3 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K) (Fragment)
Length = 1585
Score = 46.2 bits (108), Expect = 1e-04
Identities = 20/30 (66%), Positives = 22/30 (72%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N NNN+NN NNN NNNNNNNN+ PS S
Sbjct: 62 NNNNNNNNNNNNNNNNNNNNNNNVIIPSAS 91
Score = 43.9 bits (102), Expect = 7e-04
Identities = 22/43 (51%), Positives = 27/43 (62%), Gaps = 3/43 (6%)
Query: 349 KNKKRKLRKGNTINNND---NNIINNNGNNNNNNNNIASPSES 388
K K ++K N NNN+ NN NNN +N+NNNNN SPS S
Sbjct: 191 KKKLENIKKNNNNNNNNGNGNNNSNNNNSNSNNNNNGISPSSS 233
Score = 43.1 bits (100), Expect = 0.001
Identities = 19/36 (52%), Positives = 21/36 (57%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
I +N N NNN+NN NNN NNNNNNNN
Sbjct: 343 IFNENNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 378
Score = 43.1 bits (100), Expect = 0.001
Identities = 18/23 (78%), Positives = 19/23 (82%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N INNN+NN NNN NNNNNNNN
Sbjct: 55 NLINNNNNNNNNNNNNNNNNNNN 77
Score = 38.1 bits (87), Expect = 0.038
Identities = 16/33 (48%), Positives = 18/33 (54%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
N N NNN+NN NNN NNNNNN +
Sbjct: 349 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNEEL 381
Score = 37.7 bits (86), Expect = 0.050
Identities = 15/25 (60%), Positives = 18/25 (72%)
Query: 357 KGNTINNNDNNIINNNGNNNNNNNN 381
+ + N N+NN NNN NNNNNNNN
Sbjct: 340 QSDIFNENNNNNNNNNNNNNNNNNN 364
Score = 37.0 bits (84), Expect = 0.086
Identities = 19/46 (41%), Positives = 23/46 (49%), Gaps = 4/46 (8%)
Query: 340 EAFAKAISEKNKKRKLRKGNTINNNDNNIINNNGNNN----NNNNN 381
+ F + + N N NNN+NN NNN NNN NNNNN
Sbjct: 342 DIFNENNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNEELINNNNN 387
Score = 36.6 bits (83), Expect = 0.11
Identities = 18/39 (46%), Positives = 20/39 (51%), Gaps = 4/39 (10%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNG----NNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 352 NNNNNNNNNNNNNNNNNNNNNNNNNNNEELINNNNNNNN 390
Score = 35.8 bits (81), Expect = 0.19
Identities = 19/53 (35%), Positives = 24/53 (44%), Gaps = 11/53 (20%)
Query: 347 SEKNKKRKLRKGNTINNNDNNII-----------NNNGNNNNNNNNIASPSES 388
+ N N NNN+NN+I N+N N+NNNNN SP S
Sbjct: 64 NNNNNNNNNNNNNNNNNNNNNVIIPSASTENKEENDNNNSNNNNNINLSPDSS 116
Score = 35.8 bits (81), Expect = 0.19
Identities = 14/34 (41%), Positives = 18/34 (52%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNN 380
+ N N NNN+ +INNN NNNN+ N
Sbjct: 360 NNNNNNNNNNNNNNNNNNNEELINNNNNNNNDEN 393
Score = 35.0 bits (79), Expect = 0.33
Identities = 18/34 (52%), Positives = 19/34 (54%), Gaps = 3/34 (8%)
Query: 351 KKRKLRKGN---TINNNDNNIINNNGNNNNNNNN 381
K+ L N TI N I NNN NNNNNNNN
Sbjct: 37 KENSLNNSNIYLTIPTTQNLINNNNNNNNNNNNN 70
Score = 34.7 bits (78), Expect = 0.42
Identities = 17/39 (43%), Positives = 20/39 (50%), Gaps = 4/39 (10%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNN----GNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNN+
Sbjct: 353 NNNNNNNNNNNNNNNNNNNNNNNNNNEELINNNNNNNND 391
Score = 34.3 bits (77), Expect = 0.55
Identities = 16/42 (38%), Positives = 19/42 (45%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
+ N N NNN+ I NNN NNN+ N I ES
Sbjct: 361 NNNNNNNNNNNNNNNNNNEELINNNNNNNNDENYKIEETEES 402
Score = 34.3 bits (77), Expect = 0.55
Identities = 14/35 (40%), Positives = 18/35 (51%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+N + NN NNNNN+ N
Sbjct: 359 NNNNNNNNNNNNNNNNNNNNEELINNNNNNNNDEN 393
Score = 33.5 bits (75), Expect = 0.95
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 14/55 (25%)
Query: 348 EKNKKRKLRKGNTINNNDNNIINNNGNNN--------------NNNNNIASPSES 388
+KN GN NN++NN N+N NNN NNNNN ++ + S
Sbjct: 198 KKNNNNNNNNGNGNNNSNNNNSNSNNNNNGISPSSSPPSHLNGNNNNNNSNNTNS 252
Score = 30.8 bits (68), Expect = 6.1
Identities = 18/49 (36%), Positives = 21/49 (42%), Gaps = 11/49 (22%)
Query: 344 KAISEKNKKRKLRKGNTINNNDNNIINNNG-----------NNNNNNNN 381
K + N N+ NNN N+ NNNG N NNNNNN
Sbjct: 198 KKNNNNNNNNGNGNNNSNNNNSNSNNNNNGISPSSSPPSHLNGNNNNNN 246
Score = 30.8 bits (68), Expect = 6.1
Identities = 13/30 (43%), Positives = 19/30 (63%)
Query: 349 KNKKRKLRKGNTINNNDNNIINNNGNNNNN 378
K K+++ K NNDNN +NN N+NN+
Sbjct: 1547 KEKEKEKEKEKEKENNDNNDKDNNNNSNND 1576
Score = 30.4 bits (67), Expect = 8.0
Identities = 17/75 (22%), Positives = 36/75 (47%), Gaps = 8/75 (10%)
Query: 307 QMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEKNKKRKLRKGNTINNNDN 366
Q K++++R ++ R++ R+ + + + K+ K K ++ +N
Sbjct: 1510 QSKLDLSRS--------DLSRSDSSRSDSSRLDLSRSDKKNNKDNKEKEKEKEKEKEKEN 1561
Query: 367 NIINNNGNNNNNNNN 381
N N+ NNNN+NN+
Sbjct: 1562 NDNNDKDNNNNSNND 1576
>LECT_SOLTU (Q9S8M0) Chitin-binding lectin 1 precursor (PL-I)
Length = 323
Score = 46.2 bits (108), Expect = 1e-04
Identities = 25/52 (48%), Positives = 26/52 (49%), Gaps = 3/52 (5%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPP---PPPTSSRRLPPPCWTPEETSAL 54
PPPS P S PSPP P P P+PP PPP S PPP P AL
Sbjct: 157 PPPSPPPPSPPSPPPPSPPPPPPPSPPPPSPPPPSPSPPPPPASPPPPPPAL 208
Score = 44.3 bits (103), Expect = 5e-04
Identities = 23/46 (50%), Positives = 25/46 (54%), Gaps = 1/46 (2%)
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPE 49
S PPPS P PSPP P P S P+PPPPP S PP P+
Sbjct: 168 SPPPPSPPPPPPPSPPPPSPPPPS-PSPPPPPASPPPPPPALPYPQ 212
Score = 41.2 bits (95), Expect = 0.005
Identities = 21/43 (48%), Positives = 23/43 (52%), Gaps = 2/43 (4%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPP P+ PSPPS P P S P PPPP PPP +P
Sbjct: 154 PPPPPPSPPPPSPPS--PPPPSPPPPPPPSPPPPSPPPPSPSP 194
Score = 37.0 bits (84), Expect = 0.086
Identities = 22/48 (45%), Positives = 22/48 (45%), Gaps = 5/48 (10%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETS 52
SPPP P PSPP P P S P P PPP PPP P S
Sbjct: 151 SPPPPPPP---PSPPP--PSPPSPPPPSPPPPPPPSPPPPSPPPPSPS 193
Score = 30.4 bits (67), Expect = 8.0
Identities = 15/31 (48%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Query: 15 IPSPPSAVPLPL-SLPAPPPPPTSSRRLPPP 44
+PSPP P P P+PP PP S PPP
Sbjct: 149 LPSPPPPPPPPSPPPPSPPSPPPPSPPPPPP 179
>ATR1_HUMAN (Q9H6X2) Anthrax toxin receptor 1 precursor (Tumor
endothelial marker 8)
Length = 564
Score = 45.8 bits (107), Expect = 2e-04
Identities = 22/39 (56%), Positives = 24/39 (61%), Gaps = 2/39 (5%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
PPPS+PT IPSPPS +P P P P P SR PPP
Sbjct: 524 PPPSAPTPPIPSPPSTLPPPPQAPPPNRAPPPSR--PPP 560
Score = 37.4 bits (85), Expect = 0.066
Identities = 19/42 (45%), Positives = 20/42 (47%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
TSSPPP+ P P P P S P PP P S PPP
Sbjct: 503 TSSPPPAPIYTPPPPAPHCPPPPPSAPTPPIPSPPSTLPPPP 544
Score = 31.6 bits (70), Expect = 3.6
Identities = 14/34 (41%), Positives = 18/34 (52%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSS 38
+PP SP ++P PP A P + P PPP S
Sbjct: 530 TPPIPSPPSTLPPPPQAPPPNRAPPPSRPPPRPS 563
>YDC9_SCHPO (Q10172) Hypothetical protein C25G10.09c in chromosome I
Length = 1794
Score = 45.4 bits (106), Expect = 2e-04
Identities = 29/87 (33%), Positives = 41/87 (46%), Gaps = 7/87 (8%)
Query: 4 SSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSALIDSYRDKWY 63
S+P P P S+P PPSA P+P P+ PPPP + P P + SAL+
Sbjct: 1703 SAPTPPPPPMSVPPPPSAPPMPAGPPSAPPPPLPASS-APSVPNPGDRSALLQQIHT--- 1758
Query: 64 SLGRTNLKATHWQEVADAVAVRCPNSS 90
T LK T + + +A R ++S
Sbjct: 1759 ---GTRLKKTVTTDKSKPIAGRVLDAS 1782
Score = 36.2 bits (82), Expect = 0.15
Identities = 19/42 (45%), Positives = 23/42 (54%), Gaps = 4/42 (9%)
Query: 8 PSSPTESIP-SPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
P+ P + P P SA P +S P PPPPP S +PPP P
Sbjct: 1683 PAHPVSTPPVRPQSAAPPQMSAPTPPPPPMS---VPPPPSAP 1721
Score = 32.7 bits (73), Expect = 1.6
Identities = 18/46 (39%), Positives = 22/46 (47%), Gaps = 6/46 (13%)
Query: 5 SPPPSSPTESIP---SPPSAVPLPLSLPAP---PPPPTSSRRLPPP 44
S PP P + P S P+ P P+S+P P PP P PPP
Sbjct: 1688 STPPVRPQSAAPPQMSAPTPPPPPMSVPPPPSAPPMPAGPPSAPPP 1733
>PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67)
(PI4-kinase) (PtdIns-4-kinase) (PI4K-alpha)
Length = 1093
Score = 45.4 bits (106), Expect = 2e-04
Identities = 20/44 (45%), Positives = 26/44 (58%)
Query: 339 VEAFAKAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
+E + E+ K ++ NNN+NN NNN NNNNNNNNI
Sbjct: 255 IETLNSTLCEETKTSPIKDDMENNNNNNNNNNNNNNNNNNNNNI 298
Score = 43.1 bits (100), Expect = 0.001
Identities = 18/30 (60%), Positives = 24/30 (80%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N+ NNN+NNI NNN NN++NNNN P+E+
Sbjct: 184 NSNNNNNNNINNNNSNNDDNNNNEILPNEN 213
Score = 40.0 bits (92), Expect = 0.010
Identities = 20/38 (52%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNN--NNNNI 382
+ N N NNN+NNI NNN NNNN NNNNI
Sbjct: 277 NNNNNNNNNNNNNNNNNNNNNINNNNINNNNINNNNNI 314
Score = 38.1 bits (87), Expect = 0.038
Identities = 17/36 (47%), Positives = 19/36 (52%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN INNN NNNN NN
Sbjct: 275 MENNNNNNNNNNNNNNNNNNNNNINNNNINNNNINN 310
Score = 37.0 bits (84), Expect = 0.086
Identities = 20/44 (45%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Query: 340 EAFAKAISEKNKKRKLRKGNTINNNDNNIINNNGNNNN-NNNNI 382
E I + + N NNN+NN NNN NNNN NNNNI
Sbjct: 265 ETKTSPIKDDMENNNNNNNNNNNNNNNNNNNNNINNNNINNNNI 308
Score = 32.7 bits (73), Expect = 1.6
Identities = 17/34 (50%), Positives = 18/34 (52%), Gaps = 2/34 (5%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNN 380
+ N N INN NNI NNN NNNNN N
Sbjct: 284 NNNNNNNNNNNNNNINN--NNINNNNINNNNNIN 315
>EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein)
Length = 620
Score = 45.4 bits (106), Expect = 2e-04
Identities = 21/49 (42%), Positives = 28/49 (56%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEET 51
T SPPP + + PSP + P P P+PPP PT + PPP ++P T
Sbjct: 281 TYSPPPPTYSPPPPSPIYSPPPPAYSPSPPPTPTPTFSPPPPAYSPPPT 329
Score = 42.4 bits (98), Expect = 0.002
Identities = 24/53 (45%), Positives = 28/53 (52%), Gaps = 4/53 (7%)
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTPEETSAL 54
A S PPP PT S P P + P P PPPPPT S PPP ++P S +
Sbjct: 439 AYSPPPP--PTYSPPPPTYSPPPPAYAQPPPPPPTYSP--PPPAYSPPPPSPI 487
Score = 42.0 bits (97), Expect = 0.003
Identities = 24/49 (48%), Positives = 27/49 (54%), Gaps = 7/49 (14%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAP---PPPPTSSRRLPPPCWTP 48
T SPPP SP S P PP+ P P P P PPPP S PPP ++P
Sbjct: 288 TYSPPPPSPIYS-PPPPAYSPSPPPTPTPTFSPPPPAYS---PPPTYSP 332
Score = 40.4 bits (93), Expect = 0.008
Identities = 23/49 (46%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Query: 3 TSSPPP---SSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
T SPPP S P P PP+ +PLP S PPPP S PPP ++P
Sbjct: 316 TFSPPPPAYSPPPTYSPPPPTYLPLPSSPIYSPPPPVYSPP-PPPSYSP 363
Score = 39.3 bits (90), Expect = 0.017
Identities = 21/44 (47%), Positives = 25/44 (56%), Gaps = 5/44 (11%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLP-PPCWTP 48
PPPSSP P P + P P +PPPPP S LP PP ++P
Sbjct: 372 PPPSSP----PPPSFSPPPPTYEQSPPPPPAYSPPLPAPPTYSP 411
Score = 38.9 bits (89), Expect = 0.023
Identities = 23/46 (50%), Positives = 25/46 (54%), Gaps = 3/46 (6%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
T SPPP PT S P P A P PL PPPP S PPP ++P
Sbjct: 408 TYSPPP--PTYSPPPPTYAQPPPLPPTYSPPPPAYSPP-PPPTYSP 450
Score = 38.9 bits (89), Expect = 0.023
Identities = 21/47 (44%), Positives = 25/47 (52%), Gaps = 2/47 (4%)
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
A S P P+ PT S P P + P P PP PPT S PPP ++P
Sbjct: 398 AYSPPLPAPPTYSPPPPTYSPPPPTYAQPPPLPPTYSP--PPPAYSP 442
Score = 38.1 bits (87), Expect = 0.038
Identities = 22/44 (50%), Positives = 26/44 (59%), Gaps = 6/44 (13%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
SPPP P S P PPS P P + PPPPP+S PPP ++P
Sbjct: 347 SPPP--PVYSPPPPPSYSPPPPTY-LPPPPPSSP---PPPSFSP 384
Score = 37.7 bits (86), Expect = 0.050
Identities = 20/49 (40%), Positives = 25/49 (50%), Gaps = 5/49 (10%)
Query: 4 SSPPPSS----PTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
S PPP S P +P PP + P P S +PPPP PPP ++P
Sbjct: 354 SPPPPPSYSPPPPTYLPPPPPSSPPPPSF-SPPPPTYEQSPPPPPAYSP 401
Score = 36.6 bits (83), Expect = 0.11
Identities = 19/46 (41%), Positives = 23/46 (49%), Gaps = 3/46 (6%)
Query: 2 ATSSPPPSSPTESIPSPPSAVPLPLSLPAPPPP---PTSSRRLPPP 44
A + PPP PT S P P + P P + +PPPP P PPP
Sbjct: 461 AYAQPPPPPPTYSPPPPAYSPPPPSPIYSPPPPQVQPLPPTFSPPP 506
Score = 36.6 bits (83), Expect = 0.11
Identities = 20/41 (48%), Positives = 24/41 (57%), Gaps = 4/41 (9%)
Query: 5 SPPPSSPTESIPSPPS-AVPLPLSLPAPPPPPTSSRRLPPP 44
SPPP + +S P PP+ + PLP PPPPT S PPP
Sbjct: 383 SPPPPTYEQSPPPPPAYSPPLPAPPTYSPPPPTYS---PPP 420
Score = 36.6 bits (83), Expect = 0.11
Identities = 21/46 (45%), Positives = 23/46 (49%), Gaps = 2/46 (4%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
T SPPP PT S P P A P P PPPP S P P ++P
Sbjct: 447 TYSPPP--PTYSPPPPAYAQPPPPPPTYSPPPPAYSPPPPSPIYSP 490
Score = 35.8 bits (81), Expect = 0.19
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 3/42 (7%)
Query: 3 TSSPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
T SPPP + + P PP+ P P + PPP P S PPP
Sbjct: 454 TYSPPPPAYAQPPPPPPTYSPPPPAYSPPPPSPIYS---PPP 492
Score = 35.4 bits (80), Expect = 0.25
Identities = 25/76 (32%), Positives = 26/76 (33%), Gaps = 33/76 (43%)
Query: 5 SPPPSSPTESIPSPPSAVPLP----------LSLPAPP---------------------- 32
SPPP SP S P PP PLP + LP PP
Sbjct: 480 SPPPPSPIYS-PPPPQVQPLPPTFSPPPPRRIHLPPPPHRQPRPPTPTYGQPPSPPTFSP 538
Query: 33 PPPTSSRRLPPPCWTP 48
PPP PPP W P
Sbjct: 539 PPPRQIHSPPPPHWQP 554
Score = 33.9 bits (76), Expect = 0.72
Identities = 23/54 (42%), Positives = 25/54 (45%), Gaps = 8/54 (14%)
Query: 3 TSSPPPSSPTESIPSPPSAVP-----LPLSLPAPPP---PPTSSRRLPPPCWTP 48
TS PPPSS S PS P+ P +P APP PP S LPP P
Sbjct: 39 TSQPPPSSIGLSPPSAPTTTPPSRGHVPSPRHAPPRHAYPPPSHGHLPPSVGGP 92
Score = 33.5 bits (75), Expect = 0.95
Identities = 24/59 (40%), Positives = 28/59 (46%), Gaps = 17/59 (28%)
Query: 5 SPPPSS----PTESIPSPPSAVPLPLSL---PAPPP--------PPTSSRRLPPPCWTP 48
SPPP + P S P PPS P P + P PPP PPT S PPP ++P
Sbjct: 362 SPPPPTYLPPPPPSSPPPPSFSPPPPTYEQSPPPPPAYSPPLPAPPTYSP--PPPTYSP 418
Score = 32.3 bits (72), Expect = 2.1
Identities = 18/48 (37%), Positives = 21/48 (43%), Gaps = 2/48 (4%)
Query: 3 TSSPPPSSPTESIPSPPS--AVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
T PPS PT S P P + P P P PP P PP ++P
Sbjct: 560 TYGQPPSPPTFSAPPPRQIHSPPPPHRQPRPPTPTYGQPPSPPTTYSP 607
Score = 31.6 bits (70), Expect = 3.6
Identities = 18/42 (42%), Positives = 21/42 (49%), Gaps = 2/42 (4%)
Query: 8 PSSPTESIPSPPSAVPLPLSLPA-PPPPPTSSRRLPPPCWTP 48
P PT S P PP+ P P PPPPT S P P ++P
Sbjct: 260 PQPPTYS-PPPPAYAQSPQPSPTYSPPPPTYSPPPPSPIYSP 300
Score = 30.4 bits (67), Expect = 8.0
Identities = 17/42 (40%), Positives = 20/42 (47%), Gaps = 1/42 (2%)
Query: 6 PPPSSPTESIP-SPPSAVPLPLSLPAPPPPPTSSRRLPPPCW 46
P P +PT P SPP+ P P PPPP R P P +
Sbjct: 520 PRPPTPTYGQPPSPPTFSPPPPRQIHSPPPPHWQPRTPTPTY 561
>SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor
(SSG 185)
Length = 485
Score = 45.1 bits (105), Expect = 3e-04
Identities = 20/40 (50%), Positives = 22/40 (55%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
SPPP SP+ P PP P P P+PPPPP PPP
Sbjct: 252 SPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPP 291
Score = 43.1 bits (100), Expect = 0.001
Identities = 20/43 (46%), Positives = 22/43 (50%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPP P PSPP P P P PPPPP+ S PP +P
Sbjct: 266 PPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSP 308
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 2 ATSSPP--PSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
A+S PP P SP P PPS P P P PPPPP PPP
Sbjct: 237 ASSRPPSPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPP 281
Score = 41.6 bits (96), Expect = 0.003
Identities = 21/40 (52%), Positives = 21/40 (52%), Gaps = 3/40 (7%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
SP P SP PSPP P P P PPPPP S PPP
Sbjct: 247 SPRPPSPPPPSPSPPPPPPPP---PPPPPPPPPSPPPPPP 283
Score = 41.2 bits (95), Expect = 0.005
Identities = 22/46 (47%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Query: 4 SSPPPS-SPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
S PPPS SP P PP P P P PPPPP PPP +P
Sbjct: 252 SPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSP 297
Score = 40.4 bits (93), Expect = 0.008
Identities = 20/43 (46%), Positives = 21/43 (48%), Gaps = 6/43 (13%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
PPP P P PPS P P P PPPPP PPP +P
Sbjct: 263 PPPPPPPPPPPPPPSPPPPPPPPPPPPPPP------PPPSPSP 299
Score = 40.0 bits (92), Expect = 0.010
Identities = 19/40 (47%), Positives = 20/40 (49%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPP 44
SPPPS S P P + P P P PPPPP PPP
Sbjct: 243 SPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPPP 282
Score = 39.7 bits (91), Expect = 0.013
Identities = 19/40 (47%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPP-PPPTSSRRLPPP 44
PPPS P P PP P P P+PP PP+ S +PPP
Sbjct: 274 PPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPP 313
Score = 35.0 bits (79), Expect = 0.33
Identities = 17/33 (51%), Positives = 17/33 (51%), Gaps = 1/33 (3%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSS 38
PPP P PSPP P P S P PPPP S
Sbjct: 287 PPPPPPPPPSPSPPRKPPSP-SPPVPPPPSPPS 318
Score = 34.7 bits (78), Expect = 0.42
Identities = 20/42 (47%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Query: 4 SSPPPSSPTESIPS-PPSAVPLPLSLPAPPPPPTSSRRLPPP 44
+SP P SP + S PPS P P P+PPPP S PPP
Sbjct: 226 NSPLPPSPQPTASSRPPSPPPSPRP-PSPPPPSPSPPPPPPP 266
Score = 32.7 bits (73), Expect = 1.6
Identities = 21/44 (47%), Positives = 22/44 (49%), Gaps = 6/44 (13%)
Query: 6 PPPSSPTESI--PSPPSAVPLPLSLPAP---PPPPTSSRRLPPP 44
PP PT S PSPP + P P S P P PPPP PPP
Sbjct: 230 PPSPQPTASSRPPSPPPS-PRPPSPPPPSPSPPPPPPPPPPPPP 272
Score = 32.7 bits (73), Expect = 1.6
Identities = 15/38 (39%), Positives = 15/38 (39%)
Query: 6 PPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPP 43
PPP P P PP P P P P PP PP
Sbjct: 280 PPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPP 317
Score = 30.4 bits (67), Expect = 8.0
Identities = 16/42 (38%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Query: 7 PPSSPTESIPSPPSAVPLPLSLPAP----PPPPTSSRRLPPP 44
P +SP P P ++ P P+P PPPP+ S PPP
Sbjct: 224 PNNSPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPPPP 265
Score = 30.4 bits (67), Expect = 8.0
Identities = 17/44 (38%), Positives = 20/44 (44%), Gaps = 1/44 (2%)
Query: 5 SPPPSSPTESIPSPPSAVPLPLSLPAPPPPPTSSRRLPPPCWTP 48
+P +P S P PPS P S P PPP PPP +P
Sbjct: 218 NPIGPAPNNS-PLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSP 260
>FAB1_YEAST (P34756) 1-phosphatidylinositol-3-phosphate 5-kinase
FAB1 (EC 2.7.1.150) (Phosphatidylinositol-3-phosphate
5-kinase) (Type III PIP kinase)
Length = 2278
Score = 44.7 bits (104), Expect = 4e-04
Identities = 25/62 (40%), Positives = 35/62 (56%), Gaps = 4/62 (6%)
Query: 327 RTEMILESQQRIVEAFA----KAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
++ IL+ RI+ +A K N K ++ +T N N++N NNN NNNNNNNN
Sbjct: 532 QSSSILDPANRIIGNYAHRNYKFKFNYNSKGPSQQNDTANGNNDNNNNNNNNNNNNNNNS 591
Query: 383 AS 384
AS
Sbjct: 592 AS 593
Score = 32.3 bits (72), Expect = 2.1
Identities = 16/30 (53%), Positives = 21/30 (69%), Gaps = 1/30 (3%)
Query: 360 TINNNDNNIINN-NGNNNNNNNNIASPSES 388
TINN +N NN N NN N+N+NI +P+ S
Sbjct: 473 TINNLNNTTSNNSNYNNTNSNSNINNPAHS 502
Score = 31.2 bits (69), Expect = 4.7
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 6/46 (13%)
Query: 343 AKAISEKNKKRKLRKGNTINNNDNNIINNNGNNN----NNNNNIAS 384
+K S++N N NNN+NN NNN NN+ +NNNI S
Sbjct: 560 SKGPSQQNDTANGNNDNNNNNNNNN--NNNNNNSASGIADNNNIPS 603
>ARP2_PLAFA (P13824) Clustered-asparagine-rich protein (Fragment)
Length = 451
Score = 44.7 bits (104), Expect = 4e-04
Identities = 18/23 (78%), Positives = 20/23 (86%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N +NNN+NNI NNN NNNNNNNN
Sbjct: 261 NHLNNNNNNINNNNNNNNNNNNN 283
Score = 33.9 bits (76), Expect = 0.72
Identities = 17/33 (51%), Positives = 18/33 (54%)
Query: 349 KNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
KN R G N+ NN NN NNNNNNNN
Sbjct: 247 KNVNDMYRDGEMSPNHLNNNNNNINNNNNNNNN 279
Score = 33.1 bits (74), Expect = 1.2
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Query: 347 SEKNKKRKLRKGNTINNN-DNNIINNNGNNNNNNNNI 382
+ N + + N +N N N +NNN NNNN+NNN+
Sbjct: 175 NNNNNNQTNTQNNFMNRNMKNKNMNNNNNNNNSNNNM 211
Score = 32.3 bits (72), Expect = 2.1
Identities = 17/30 (56%), Positives = 21/30 (69%), Gaps = 6/30 (20%)
Query: 359 NTINNNDNNIINNNGNNNN-----NNNNIA 383
N INNN+NN NNN NNNN NN+++A
Sbjct: 268 NNINNNNNNN-NNNNNNNNVMFRQNNSHLA 296
Score = 31.6 bits (70), Expect = 3.6
Identities = 16/35 (45%), Positives = 19/35 (53%), Gaps = 2/35 (5%)
Query: 349 KNKKRKLRKGNTINNNDNN--IINNNGNNNNNNNN 381
+N K K N NNN NN ++N N NNN NN
Sbjct: 191 RNMKNKNMNNNNNNNNSNNNMMMNMNFNNNQQMNN 225
Score = 31.6 bits (70), Expect = 3.6
Identities = 14/24 (58%), Positives = 17/24 (70%), Gaps = 1/24 (4%)
Query: 357 KGNTINNNDNNIINNNGNNNNNNN 380
+GN N N+ N NNN +NNNNNN
Sbjct: 158 QGNN-NMNNYNFYNNNSSNNNNNN 180
>KYK1_DICDI (P18160) Non-receptor tyrosine kinase spore lysis A (EC
2.7.1.112) (Tyrosine-protein kinase 1)
Length = 1584
Score = 44.3 bits (103), Expect = 5e-04
Identities = 19/36 (52%), Positives = 23/36 (63%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
S N N+ +N++NN INNN NNNNNNNNI
Sbjct: 498 SNSNNNNNNNNSNSNSNSNNNNINNNNNNNNNNNNI 533
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/35 (54%), Positives = 21/35 (59%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
S N + N NNN+NN NNN NNNNNNNN
Sbjct: 438 SSINNNEDISSNNNNNNNNNNNNNNNNNNNNNNNN 472
Score = 42.0 bits (97), Expect = 0.003
Identities = 20/39 (51%), Positives = 23/39 (58%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
N+ N NNN+NN NNN NNNNNNNN + S S
Sbjct: 443 NEDISSNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNS 481
Score = 42.0 bits (97), Expect = 0.003
Identities = 18/30 (60%), Positives = 22/30 (73%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N NNN+NN NNN NNNNNNNN ++ S +
Sbjct: 455 NNNNNNNNNNNNNNNNNNNNNNNNSNSSNT 484
Score = 41.6 bits (96), Expect = 0.003
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 3/45 (6%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNG---NNNNNNNNIASPSES 388
+ N NT NNN NN NNN NNNNNNNN S S S
Sbjct: 471 NNNNNNNNSNSSNTNNNNINNTTNNNNSNSNNNNNNNNSNSNSNS 515
Score = 41.6 bits (96), Expect = 0.003
Identities = 19/30 (63%), Positives = 21/30 (69%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N NNN+NN NNN NNNNNNNN S S +
Sbjct: 454 NNNNNNNNNNNNNNNNNNNNNNNNNSNSSN 483
Score = 41.2 bits (95), Expect = 0.005
Identities = 18/36 (50%), Positives = 21/36 (58%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
IS N N NNN+NN NNN NNNNN+N+
Sbjct: 446 ISSNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNS 481
Score = 40.4 bits (93), Expect = 0.008
Identities = 17/39 (43%), Positives = 23/39 (58%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
N N NNN+NN NNN NNNNNN+N ++ + +
Sbjct: 449 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNSSNTNNN 487
Score = 38.5 bits (88), Expect = 0.029
Identities = 20/42 (47%), Positives = 21/42 (49%), Gaps = 6/42 (14%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNN------NNNNI 382
S N N NNN+NN NNN NNNN NNNNI
Sbjct: 448 SNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNSSNTNNNNI 489
Score = 35.8 bits (81), Expect = 0.19
Identities = 15/36 (41%), Positives = 22/36 (60%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
+ N N+ +NN+NN N+N N+N+NNNNI
Sbjct: 485 NNNNINNTTNNNNSNSNNNNNNNNSNSNSNSNNNNI 520
Score = 35.0 bits (79), Expect = 0.33
Identities = 15/35 (42%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N +N+++N NNN NNNNNNNN
Sbjct: 494 NNNNSNSNNNNNNNNSNSNSNSNNNNINNNNNNNN 528
Score = 35.0 bits (79), Expect = 0.33
Identities = 15/35 (42%), Positives = 18/35 (50%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N N+N N+ NNN NNNNNNN
Sbjct: 493 TNNNNSNSNNNNNNNNSNSNSNSNNNNINNNNNNN 527
Score = 35.0 bits (79), Expect = 0.33
Identities = 14/35 (40%), Positives = 19/35 (54%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N+ N N+NNI N NNN+N+NN
Sbjct: 468 NNNNNNNNNNNSNSSNTNNNNINNTTNNNNSNSNN 502
Score = 34.7 bits (78), Expect = 0.42
Identities = 15/32 (46%), Positives = 19/32 (58%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
N N+ NNN+NN N+N N+NNNN N
Sbjct: 490 NNTTNNNNSNSNNNNNNNNSNSNSNSNNNNIN 521
Score = 34.3 bits (77), Expect = 0.55
Identities = 15/21 (71%), Positives = 17/21 (80%), Gaps = 2/21 (9%)
Query: 362 NNNDNNIINNNGNNNNNNNNI 382
N N+NN N+N NNNNNNNNI
Sbjct: 403 NGNNNN--NSNNNNNNNNNNI 421
Score = 32.7 bits (73), Expect = 1.6
Identities = 14/36 (38%), Positives = 19/36 (51%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
I+ N NNN++N +N+ NNN NNNN
Sbjct: 489 INNTTNNNNSNSNNNNNNNNSNSNSNSNNNNINNNN 524
Score = 32.3 bits (72), Expect = 2.1
Identities = 13/35 (37%), Positives = 20/35 (57%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N N+N+NN NN+ +N+N+NNN
Sbjct: 484 TNNNNINNTTNNNNSNSNNNNNNNNSNSNSNSNNN 518
Score = 32.0 bits (71), Expect = 2.8
Identities = 13/24 (54%), Positives = 17/24 (70%)
Query: 361 INNNDNNIINNNGNNNNNNNNIAS 384
+ N +NN +NN NNNNNNN I +
Sbjct: 401 VPNGNNNNNSNNNNNNNNNNIIGN 424
Score = 31.2 bits (69), Expect = 4.7
Identities = 15/40 (37%), Positives = 18/40 (44%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPS 386
+ N + NNN+ N NNN NNNNN PS
Sbjct: 501 NNNNNNNNSNSNSNSNNNNINNNNNNNNNNNNIYLTKKPS 540
Score = 31.2 bits (69), Expect = 4.7
Identities = 13/17 (76%), Positives = 13/17 (76%)
Query: 366 NNIINNNGNNNNNNNNI 382
NN N N NNNNNNNNI
Sbjct: 1195 NNGNNGNNNNNNNNNNI 1211
Score = 31.2 bits (69), Expect = 4.7
Identities = 23/120 (19%), Positives = 51/120 (42%), Gaps = 8/120 (6%)
Query: 155 NPNHNANHNHIHSHNLINHRETLENDDYDDDDLYEELRSAAGGGSGSGNTRSLDKLYRNG 214
N N+N N+N+ +++N N+ + ++ + ++ S + + + N+ S N
Sbjct: 460 NNNNNNNNNNNNNNNNNNNSNSSNTNNNNINNTTNNNNSNSNNNNNNNNSNSNSNSNNNN 519
Query: 215 VSGGFGGSGSGFRIRIPTGVSVAQPGSKFYPKM-------NNNSESGSRINGGTRLVKER 267
++ + + I + S+ + NNNS SGS I + ++K+R
Sbjct: 520 INNNNNNNNNNNNIYLTKKPSIGSTDESSTGSLGGNNSSGNNNSSSGS-IGNNSSIIKQR 578
>VP61_NPVOP (O10270) 61 kDa protein homolog
Length = 474
Score = 43.9 bits (102), Expect = 7e-04
Identities = 23/54 (42%), Positives = 28/54 (51%), Gaps = 10/54 (18%)
Query: 1 MATSSPPPSSP------TESIPSPPSAVPLPLSLPAPPPPPTS----SRRLPPP 44
+ TS PPP P T S+P PP V L S+P PPPPP + +PPP
Sbjct: 277 VTTSMPPPPPPFPSADVTTSMPPPPPMVDLATSMPPPPPPPPPMVDLATSMPPP 330
Score = 33.9 bits (76), Expect = 0.72
Identities = 18/46 (39%), Positives = 23/46 (49%), Gaps = 7/46 (15%)
Query: 6 PPPSSP----TESIPSPPSAVP---LPLSLPAPPPPPTSSRRLPPP 44
PPP P T S+P PP P + S+P PPP + +PPP
Sbjct: 268 PPPPFPSADVTTSMPPPPPPFPSADVTTSMPPPPPMVDLATSMPPP 313
>PTP3_DICDI (P54637) Protein-tyrosine phosphatase 3 (EC 3.1.3.48)
(Protein-tyrosine-phosphate phosphohydrolase 3)
Length = 989
Score = 43.9 bits (102), Expect = 7e-04
Identities = 19/33 (57%), Positives = 20/33 (60%)
Query: 349 KNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
KN N NNN+NN NNN NNNNNNNN
Sbjct: 136 KNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 168
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/36 (52%), Positives = 20/36 (54%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
I N N NNN+NN NNN NNNNNNNN
Sbjct: 135 IKNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 170
Score = 43.1 bits (100), Expect = 0.001
Identities = 19/36 (52%), Positives = 21/36 (57%)
Query: 346 ISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
I + N N NNN+NN NNN NNNNNNNN
Sbjct: 134 IIKNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 169
Score = 43.1 bits (100), Expect = 0.001
Identities = 20/42 (47%), Positives = 23/42 (54%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
+ N N NNN+NN NNN NNNNNNNN + S S
Sbjct: 148 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNS 189
Score = 42.7 bits (99), Expect = 0.002
Identities = 20/41 (48%), Positives = 22/41 (52%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSE 387
+ N N NNN+NN NNN NNNNNNNN S E
Sbjct: 152 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNSNIE 192
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 137 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 171
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 138 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 172
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 141 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 175
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 145 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 179
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 143 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 177
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 140 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 174
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 139 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 173
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 144 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 178
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 146 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 180
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 142 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 176
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 20/35 (56%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
+ N N NNN+NN NNN NNNNNNNN
Sbjct: 147 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN 181
Score = 40.0 bits (92), Expect = 0.010
Identities = 18/28 (64%), Positives = 19/28 (67%)
Query: 354 KLRKGNTINNNDNNIINNNGNNNNNNNN 381
KL I NN+NN NNN NNNNNNNN
Sbjct: 128 KLSNTMIIKNNNNNNNNNNNNNNNNNNN 155
Score = 36.2 bits (82), Expect = 0.15
Identities = 15/23 (65%), Positives = 16/23 (69%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N+INN NN N NNNNNNNN
Sbjct: 96 NSINNKINNNTTTNNNNNNNNNN 118
Score = 36.2 bits (82), Expect = 0.15
Identities = 16/20 (80%), Positives = 16/20 (80%), Gaps = 2/20 (10%)
Query: 362 NNNDNNIINNNGNNNNNNNN 381
NNN NN NNN NNNNNNNN
Sbjct: 943 NNNKNN--NNNSNNNNNNNN 960
Score = 35.4 bits (80), Expect = 0.25
Identities = 19/47 (40%), Positives = 21/47 (44%), Gaps = 19/47 (40%)
Query: 354 KLRKGNTINNNDNN-------------------IINNNGNNNNNNNN 381
K+ T NNN+NN II NN NNNNNNNN
Sbjct: 101 KINNNTTTNNNNNNNNNNDDKFDTNALKLSNTMIIKNNNNNNNNNNN 147
Score = 34.3 bits (77), Expect = 0.55
Identities = 17/40 (42%), Positives = 21/40 (52%), Gaps = 2/40 (5%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPS 386
+ N N NNN+NN NNN NN+N+N I PS
Sbjct: 160 NNNNNNNNNNNNNNNNNNNNN--NNNNNNSNSNIEINVPS 197
Score = 33.5 bits (75), Expect = 0.95
Identities = 13/20 (65%), Positives = 15/20 (75%)
Query: 362 NNNDNNIINNNGNNNNNNNN 381
N N+NN NNN NNNNN N+
Sbjct: 945 NKNNNNNSNNNNNNNNNKNS 964
Score = 33.1 bits (74), Expect = 1.2
Identities = 13/20 (65%), Positives = 15/20 (75%)
Query: 362 NNNDNNIINNNGNNNNNNNN 381
NN +NN +NN NNNNNN N
Sbjct: 944 NNKNNNNNSNNNNNNNNNKN 963
Score = 32.7 bits (73), Expect = 1.6
Identities = 13/23 (56%), Positives = 17/23 (73%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N NNN+N+ NNN NNN N++N
Sbjct: 944 NNKNNNNNSNNNNNNNNNKNSDN 966
Score = 32.7 bits (73), Expect = 1.6
Identities = 14/32 (43%), Positives = 20/32 (61%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
N ++ + + N N NNI N++ N NNNNNN
Sbjct: 40 NNTPEMIQSQSENTNTNNINNSSSNINNNNNN 71
Score = 32.7 bits (73), Expect = 1.6
Identities = 13/23 (56%), Positives = 16/23 (69%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N NN+NN NNN NNNN N++
Sbjct: 943 NNNKNNNNNSNNNNNNNNNKNSD 965
Score = 32.3 bits (72), Expect = 2.1
Identities = 13/23 (56%), Positives = 17/23 (73%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N NNN++N NNN NN N++NN
Sbjct: 945 NKNNNNNSNNNNNNNNNKNSDNN 967
Score = 32.0 bits (71), Expect = 2.8
Identities = 13/27 (48%), Positives = 16/27 (59%)
Query: 362 NNNDNNIINNNGNNNNNNNNIASPSES 388
NNN NN NNN N N++NN E+
Sbjct: 949 NNNSNNNNNNNNNKNSDNNGTKDKDEN 975
Score = 31.2 bits (69), Expect = 4.7
Identities = 14/26 (53%), Positives = 19/26 (72%), Gaps = 2/26 (7%)
Query: 358 GNTINNND--NNIINNNGNNNNNNNN 381
G++ N++ NN NNN N+NNNNNN
Sbjct: 933 GSSSTNSECSNNNKNNNNNSNNNNNN 958
>CAR3_DICDI (P35352) Cyclic AMP receptor 3
Length = 490
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/35 (54%), Positives = 23/35 (65%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
S+++ K N NNN+NN NNN NNNNNNNN
Sbjct: 388 SQQDDKDSPNSNNNNNNNNNNNNNNNNNNNNNNNN 422
Score = 41.6 bits (96), Expect = 0.003
Identities = 19/37 (51%), Positives = 22/37 (59%)
Query: 345 AISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNN 381
A + +K N NNN+NN NNN NNNNNNNN
Sbjct: 387 ASQQDDKDSPNSNNNNNNNNNNNNNNNNNNNNNNNNN 423
Score = 41.6 bits (96), Expect = 0.003
Identities = 24/79 (30%), Positives = 35/79 (43%), Gaps = 3/79 (3%)
Query: 302 FVRMEQMKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEKNKKRKLRKGNTI 361
FV + A IE+ ++ E ++ + + A +K+ N
Sbjct: 349 FVNNDSSNYYTASMIESFSVQNENSKS---INGADNFKQNGASQQDDKDSPNSNNNNNNN 405
Query: 362 NNNDNNIINNNGNNNNNNN 380
NNN+NN NNN NNNNNNN
Sbjct: 406 NNNNNNNNNNNNNNNNNNN 424
Score = 40.8 bits (94), Expect = 0.006
Identities = 17/23 (73%), Positives = 18/23 (77%)
Query: 359 NTINNNDNNIINNNGNNNNNNNN 381
N NNN+NN NNN NNNNNNNN
Sbjct: 402 NNNNNNNNNNNNNNNNNNNNNNN 424
Score = 38.1 bits (87), Expect = 0.038
Identities = 17/39 (43%), Positives = 20/39 (50%)
Query: 350 NKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
N N NNN+NN NNN NNNN NN P ++
Sbjct: 397 NSNNNNNNNNNNNNNNNNNNNNNNNNNNYNNKDIEPIDN 435
Score = 35.4 bits (80), Expect = 0.25
Identities = 16/36 (44%), Positives = 19/36 (52%)
Query: 347 SEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNI 382
S + N NNN+NN NNN NNN NN +I
Sbjct: 395 SPNSNNNNNNNNNNNNNNNNNNNNNNNNNNYNNKDI 430
Score = 35.0 bits (79), Expect = 0.33
Identities = 14/20 (70%), Positives = 16/20 (80%)
Query: 362 NNNDNNIINNNGNNNNNNNN 381
NNN NN NN+ NNNNNNN+
Sbjct: 328 NNNHNNNHNNHNNNNNNNNS 347
Score = 34.7 bits (78), Expect = 0.42
Identities = 22/59 (37%), Positives = 29/59 (48%), Gaps = 18/59 (30%)
Query: 332 LESQQRIVEAFAKAISEKNKKRKLR---------KGNTINNNDNNIINNNGNNNNNNNN 381
+E+QQR+ EKNK + N ++N NN NNN NN+NNNNN
Sbjct: 294 VETQQRL---------EKNKNNNNHSPVGLSNNAQNNNHHHNHNNNHNNNHNNHNNNNN 343
Score = 34.3 bits (77), Expect = 0.55
Identities = 29/121 (23%), Positives = 44/121 (35%), Gaps = 28/121 (23%)
Query: 145 SPPSPPHKPENPNHNANHNHIHSHNLINHRETLENDDYD----DDDLY---EELRSAAGG 197
SP + +N NH+ NHN+ H++N NH N++ D D Y + S +
Sbjct: 310 SPVGLSNNAQNNNHHHNHNNNHNNNHNNHNNNNNNNNSDFVNNDSSNYYTASMIESFSVQ 369
Query: 198 GSGSGNTRSLDKLYRNGVSGGFGGSGSGFRIRIPTGVSVAQPGSKFYPKMNNNSESGSRI 257
S + D +NG S Q K P NNN+ + +
Sbjct: 370 NENSKSINGADNFKQNGAS---------------------QQDDKDSPNSNNNNNNNNNN 408
Query: 258 N 258
N
Sbjct: 409 N 409
Score = 33.5 bits (75), Expect = 0.95
Identities = 14/30 (46%), Positives = 20/30 (66%)
Query: 359 NTINNNDNNIINNNGNNNNNNNNIASPSES 388
N NN++NN N+N NNNNNN++ + S
Sbjct: 326 NHNNNHNNNHNNHNNNNNNNNSDFVNNDSS 355
Score = 31.2 bits (69), Expect = 4.7
Identities = 15/40 (37%), Positives = 22/40 (54%), Gaps = 2/40 (5%)
Query: 349 KNKKRKLRKGNTINNNDNNIINNNGNNNNNNNNIASPSES 388
+N N NNN NN +NN NNNNN++ + + S +
Sbjct: 319 QNNNHHHNHNNNHNNNHNN--HNNNNNNNNSDFVNNDSSN 356
Score = 30.4 bits (67), Expect = 8.0
Identities = 15/49 (30%), Positives = 26/49 (52%), Gaps = 7/49 (14%)
Query: 338 IVEAFAKAISEKNKKR-----KLRKGNTINNNDNNIINNNGNNNNNNNN 381
++E+F+ + +N K ++ +D + N+N NNNNNNNN
Sbjct: 362 MIESFS--VQNENSKSINGADNFKQNGASQQDDKDSPNSNNNNNNNNNN 408
>RAEP_CANAL (O93831) Rab proteins geranylgeranyltransferase
component A (Rab escort protein) (REP)
Length = 640
Score = 43.1 bits (100), Expect = 0.001
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 2/61 (3%)
Query: 321 MEMEMKRTEMILESQQRIVEAFAKAISEKNKKRKLRKGNTINNNDNNIINNNGNNNNNNN 380
+E ++ R ++ LES +E S + + N NNN+NN NNN NNNNNNN
Sbjct: 381 VEQDLSRAKVDLESAFEKMETSLLRESSEEIVNDILGDNNNNNNNNN--NNNNNNNNNNN 438
Query: 381 N 381
N
Sbjct: 439 N 439
Score = 30.4 bits (67), Expect = 8.0
Identities = 20/73 (27%), Positives = 32/73 (43%), Gaps = 13/73 (17%)
Query: 308 MKMEMAREIETMRMEMEMKRTEMILESQQRIVEAFAKAISEKNKKRKLRKGNTINNNDNN 367
++ +++R + E T ++ ES + IV + N N NNN+NN
Sbjct: 381 VEQDLSRAKVDLESAFEKMETSLLRESSEEIVNDILGDNNNNNN-------NNNNNNNNN 433
Query: 368 IINNNGNNNNNNN 380
NNNNNN+
Sbjct: 434 ------NNNNNND 440
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.129 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,662,466
Number of Sequences: 164201
Number of extensions: 2552731
Number of successful extensions: 37304
Number of sequences better than 10.0: 838
Number of HSP's better than 10.0 without gapping: 398
Number of HSP's successfully gapped in prelim test: 465
Number of HSP's that attempted gapping in prelim test: 22535
Number of HSP's gapped (non-prelim): 7525
length of query: 388
length of database: 59,974,054
effective HSP length: 112
effective length of query: 276
effective length of database: 41,583,542
effective search space: 11477057592
effective search space used: 11477057592
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149129.9


