

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
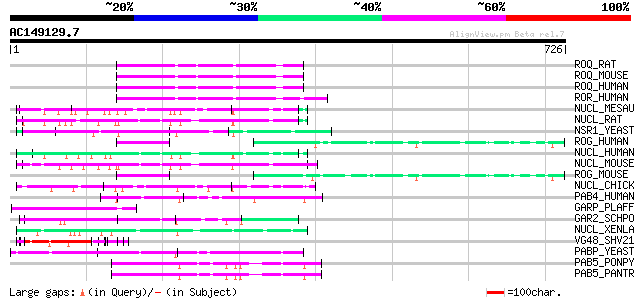
Query= AC149129.7 + phase: 0
(726 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ROQ_RAT (Q7TP47) Heterogeneous nuclear ribonucleoprotein Q (hnRN... 96 3e-19
ROQ_MOUSE (Q7TMK9) Heterogeneous nuclear ribonucleoprotein Q (hn... 96 3e-19
ROQ_HUMAN (O60506) Heterogeneous nuclear ribonucleoprotein Q (hn... 94 1e-18
ROR_HUMAN (O43390) Heterogeneous nuclear ribonucleoprotein R (hn... 90 2e-17
NUCL_MESAU (P08199) Nucleolin (Protein C23) 80 2e-14
NUCL_RAT (P13383) Nucleolin (Protein C23) 76 4e-13
NSR1_YEAST (P27476) Nuclear localization sequence binding protei... 76 4e-13
ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hn... 74 1e-12
NUCL_HUMAN (P19338) Nucleolin (Protein C23) 71 9e-12
NUCL_MOUSE (P09405) Nucleolin (Protein C23) 71 1e-11
ROG_MOUSE (O35479) Heterogeneous nuclear ribonucleoprotein G (hn... 70 3e-11
NUCL_CHICK (P15771) Nucleolin (Protein C23) 69 4e-11
PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4 (Poly(A)-bin... 69 6e-11
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 69 6e-11
GAR2_SCHPO (P41891) Protein gar2 68 7e-11
NUCL_XENLA (P20397) Nucleolin (Protein C23) 68 1e-10
VG48_SHV21 (Q01033) Hypothetical gene 48 protein 66 3e-10
PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic a... 65 5e-10
PAB5_PONPY (P60050) Polyadenylate-binding protein 5 (Poly(A)-bin... 65 5e-10
PAB5_PANTR (P60049) Polyadenylate-binding protein 5 (Poly(A)-bin... 65 5e-10
>ROQ_RAT (Q7TP47) Heterogeneous nuclear ribonucleoprotein Q (hnRNP
Q) (hnRNP-Q) (Synaptotagmin binding, cytoplasmic RNA
interacting protein) (Liver regeneration-related protein
LRRG077) (Ab2-339)
Length = 533
Score = 95.9 bits (237), Expect = 3e-19
Identities = 69/247 (27%), Positives = 123/247 (48%), Gaps = 15/247 (6%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
E+FVG + ++ E +L +F K G I ++R+ ++P T N+G+A + F T E + A+
Sbjct: 73 EIFVGKIPRDLFEDELVPLFEKAGPIWDLRLMMDPLTGLNRGYAFVTFCTKEAAQEAVKL 132
Query: 200 LKNPVI-NGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDD 258
N I +GK G+ I+ ++ L++ +I KS KE + E+ E D+ L
Sbjct: 133 YNNHEIRSGKHIGVCISVA--NNRLFVGSIPKSKTKEQILEEFSKV-TEGLTDVILYHQP 189
Query: 259 NNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV-VFGVDKPAEVSFANSFIDLGDDIMAQV 317
+++ N G FLE+ H + A +RL V V+G V +A+ D ++MA+V
Sbjct: 190 DDKKKNRGFCFLEYEDHKTAAQARRRLMSGKVKVWG--NVGTVEWADPIEDPDPEVMAKV 247
Query: 318 KTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAE 377
K +F+ L + E+ + ++G +E+V+ + K+Y F+ F AV+ E
Sbjct: 248 KVLFVRNLANTVTEEILEKSFSQFGKLERVK--------KLKDYAFIHFDERDGAVKAME 299
Query: 378 SITSAGL 384
+ L
Sbjct: 300 EMNGKDL 306
>ROQ_MOUSE (Q7TMK9) Heterogeneous nuclear ribonucleoprotein Q (hnRNP
Q) (hnRNP-Q) (Synaptotagmin binding, cytoplasmic RNA
interacting protein) (Glycine-and tyrosine-rich RNA
binding protein) (GRY-RBP) (NS1-associated protein 1)
(pp68)
Length = 623
Score = 95.9 bits (237), Expect = 3e-19
Identities = 69/247 (27%), Positives = 123/247 (48%), Gaps = 15/247 (6%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
E+FVG + ++ E +L +F K G I ++R+ ++P T N+G+A + F T E + A+
Sbjct: 163 EIFVGKIPRDLFEDELVPLFEKAGPIWDLRLMMDPLTGLNRGYAFVTFCTKEAAQEAVKL 222
Query: 200 LKNPVI-NGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDD 258
N I +GK G+ I+ ++ L++ +I KS KE + E+ E D+ L
Sbjct: 223 YNNHEIRSGKHIGVCISVA--NNRLFVGSIPKSKTKEQILEEFSKV-TEGLTDVILYHQP 279
Query: 259 NNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV-VFGVDKPAEVSFANSFIDLGDDIMAQV 317
+++ N G FLE+ H + A +RL V V+G V +A+ D ++MA+V
Sbjct: 280 DDKKKNRGFCFLEYEDHKTAAQARRRLMSGKVKVWG--NVGTVEWADPIEDPDPEVMAKV 337
Query: 318 KTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAE 377
K +F+ L + E+ + ++G +E+V+ + K+Y F+ F AV+ E
Sbjct: 338 KVLFVRNLANTVTEEILEKSFSQFGKLERVK--------KLKDYAFIHFDERDGAVKAME 389
Query: 378 SITSAGL 384
+ L
Sbjct: 390 EMNGKDL 396
>ROQ_HUMAN (O60506) Heterogeneous nuclear ribonucleoprotein Q (hnRNP
Q) (hnRNP-Q) (Synaptotagmin binding, cytoplasmic RNA
interacting protein) (Glycine-and tyrosine-rich RNA
binding protein) (GRY-RBP) (NS1-associated protein 1)
Length = 623
Score = 94.0 bits (232), Expect = 1e-18
Identities = 68/247 (27%), Positives = 122/247 (48%), Gaps = 15/247 (6%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
E+FVG + ++ E +L +F K G I ++R+ ++P T N+G+A + F T E + A+
Sbjct: 163 EIFVGKIPRDLFEDELVPLFEKAGPIWDLRLMMDPLTGLNRGYAFVTFCTKEAAQEAVKL 222
Query: 200 LKNPVI-NGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDD 258
N I +GK G+ I+ ++ L++ +I KS KE + E+ E D+ L
Sbjct: 223 YNNHEIRSGKHIGVCISVA--NNRLFVGSIPKSKTKEQILEEFSKV-TEGLTDVILYHQP 279
Query: 259 NNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV-VFGVDKPAEVSFANSFIDLGDDIMAQV 317
+++ N FLE+ H + A +RL V V+G V +A+ D ++MA+V
Sbjct: 280 DDKKKNRSFCFLEYEDHKTAAQARRRLMSGKVKVWG--NVGTVEWADPIEDPDPEVMAKV 337
Query: 318 KTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAE 377
K +F+ L + E+ + ++G +E+V+ + K+Y F+ F AV+ E
Sbjct: 338 KVLFVRNLANTVTEEILEKAFSQFGKLERVK--------KLKDYAFIHFDERDGAVKAME 389
Query: 378 SITSAGL 384
+ L
Sbjct: 390 EMNGKDL 396
>ROR_HUMAN (O43390) Heterogeneous nuclear ribonucleoprotein R (hnRNP
R)
Length = 633
Score = 89.7 bits (221), Expect = 2e-17
Identities = 72/278 (25%), Positives = 136/278 (48%), Gaps = 19/278 (6%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
EVFVG + ++ E +L +F K G I ++R+ ++P + +N+G+A + F E + A+
Sbjct: 166 EVFVGKIPRDLYEDELVPLFEKAGPIWDLRLMMDPLSGQNRGYAFITFCGKEAAQEAVKL 225
Query: 200 LKNPVIN-GKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDD 258
+ I GK G+ I+ ++ L++ +I K+ KE + E+ E D+ L
Sbjct: 226 CDSYEIRPGKHLGVCISVA--NNRLFVGSIPKNKTKENILEEFSKV-TEGLVDVILYHQP 282
Query: 259 NNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV-VFGVDKPAEVSFANSFIDLGDDIMAQV 317
+++ N G FLE+ H + A +RL V V+G V +A+ + ++MA+V
Sbjct: 283 DDKKKNRGFCFLEYEDHKSAAQARRRLMSGKVKVWG--NVVTVEWADPVEEPDPEVMAKV 340
Query: 318 KTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAE 377
K +F+ L + E+ + ++G +E+V+ + K+Y FV F AAV+ +
Sbjct: 341 KVLFVRNLATTVTEEILEKSFSEFGKLERVK--------KLKDYAFVHFEDRGAAVKAMD 392
Query: 378 SITSAGLGEGNKKAKVRARLSRPLPKVRGKHVPHRDIS 415
+ G+ + ++ L++P K R + R S
Sbjct: 393 EMN----GKEIEGEEIEIVLAKPPDKKRKERQAARQAS 426
>NUCL_MESAU (P08199) Nucleolin (Protein C23)
Length = 713
Score = 80.1 bits (196), Expect = 2e-14
Identities = 96/427 (22%), Positives = 179/427 (41%), Gaps = 74/427 (17%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAG------ 62
A NG+ K+ + ++D+ D +D+ + + EE +D+++ ++ +G+ A
Sbjct: 132 AKNGKNAKKEDSDEDEDDDDDEDDSDEDEEDEE---EDEFEPPVVKGKQGKVAAAAPASE 188
Query: 63 --DEVEEEPEENVDEEEGDTGDEE----------------VEEVEYVYEEVEGDDDDVAG 104
DE E+E EE DEEE D +EE V V+ E DDD+
Sbjct: 189 DEDEEEDEEEEEEDEEEEDDSEEEEAMEITPAKGKKAPAKVVPVKAKNVAEEDDDDEEED 248
Query: 105 DDEHASEENVHEHLADVKEEEQVEV-------IKETQKQKEL-----------------E 140
+DE EE + + +EEE+ V KE KQKE+
Sbjct: 249 EDEEEDEEEEEDEEEEEEEEEEEPVKPAPGKRKKEMTKQKEVPEAKKQKVEGSESTTPFN 308
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
+F+G L+ + +L+ S+ ++ + V+ +T N+ F + FE+ E +++AL EL
Sbjct: 309 LFIGNLNPNKSVAELKVAISEPFAKNDLAV-VDVRTGTNRKFGYVDFESAEDLEKAL-EL 366
Query: 201 KNPVINGKQCGITITTCQDSD------TLYLDNICKSWKKEALKEKLKHYGVESFKDLTL 254
+ G + + +DS TL N+ + ++ LK E F+D
Sbjct: 367 TGLKVFGNEIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELK--------EVFEDALE 418
Query: 255 LEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV---VFGVDKPAEVSFANSFIDLGD 311
+ + +G + G A++EF S +D++ + Q ++ + E
Sbjct: 419 IRLVSQDGKSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEKGQRQERTGKNS 478
Query: 312 DIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAA 371
+ KT+ + L S E+ ++ + +K +++ +N G + K Y F+ F +
Sbjct: 479 TWSGESKTLVLSNLSYSATEETLQEVFEK---ATFIKVPQNQQG-KSKGYAFIEFASFED 534
Query: 372 AVECAES 378
A E S
Sbjct: 535 AKEALNS 541
Score = 59.3 bits (142), Expect = 3e-08
Identities = 106/470 (22%), Positives = 178/470 (37%), Gaps = 124/470 (26%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVG--------DDDYDERGIEQDEGQEAGDE 64
EE +E D+ E++E ++ + P + V +DD DE E+DE +E +E
Sbjct: 200 EEDEEEEDDSEEEEAMEITPAKGKKAPAKVVPVKAKNVAEEDDDDE---EEDEDEEEDEE 256
Query: 65 VEEEPEENVDEEEGD-------------TGDEEVEEVEYVYEEVEGDDD----------- 100
EE+ EE +EEE + T +EV E + ++VEG +
Sbjct: 257 EEEDEEEEEEEEEEEPVKPAPGKRKKEMTKQKEVPEAKK--QKVEGSESTTPFNLFIGNL 314
Query: 101 ----DVAGDDEHASEENVHEHLA--DVK----------EEEQVEVIKETQKQKELEVFVG 144
VA SE LA DV+ + E E +++ + L+VF
Sbjct: 315 NPNKSVAELKVAISEPFAKNDLAVVDVRTGTNRKFGYVDFESAEDLEKALELTGLKVFGN 374
Query: 145 GLDKEA------------------------TEHDLRKVFSKVGEITEVRMTVNPQTKRNK 180
+ E TE +L++VF EI V Q ++K
Sbjct: 375 EIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELKEVFEDALEIRLVS-----QDGKSK 429
Query: 181 GFALLRFETVEHVKRALAELKNPVINGK---------------QCGITITTCQDSDTLYL 225
G A + F++ ++ L E + I+G+ + G T +S TL L
Sbjct: 430 GIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEKGQRQERTGKNSTWSGESKTLVL 489
Query: 226 DNICKSWKKEALKEKLKHYGVESFKDLTLLE-DDNNEGTNCGCAFLEFSSHSDSKDAYKR 284
N+ S +E L+E F+ T ++ N +G + G AF+EF+S D+K+A
Sbjct: 490 SNLSYSATEETLQEV--------FEKATFIKVPQNQQGKSKGYAFIEFASFEDAKEALNS 541
Query: 285 LQKTDVV-----FGVDKPAEVSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLK 339
K ++ + P A S KT+F+ L E+ L +
Sbjct: 542 CNKMEIEGRTIRLELQGPRGSPNARS---------QPSKTLFVKGLSEDTTEE---TLKE 589
Query: 340 KYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESITSAGLGEGNK 389
+ + + + K +GFV F + A E++ + +GNK
Sbjct: 590 SFEGSVRARIVTDRETGSSKGFGFVDFNSEEDAKAAKEAMEDGEI-DGNK 638
Score = 52.0 bits (123), Expect = 6e-06
Identities = 52/220 (23%), Positives = 97/220 (43%), Gaps = 21/220 (9%)
Query: 82 DEEVEEVEYVYEEVEGDDDDVAGDDEHASEE--NVHEHLADVKEEEQVEVIKETQKQKEL 139
D + + + Y+ + E D + + + A + +V + K + Q K + E
Sbjct: 425 DGKSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEKGQRQERTGKNSTWSGES 484
Query: 140 EVFV-GGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALA 198
+ V L ATE L++VF K T +++ N Q K +KG+A + F + E K AL
Sbjct: 485 KTLVLSNLSYSATEETLQEVFEKA---TFIKVPQNQQGK-SKGYAFIEFASFEDAKEALN 540
Query: 199 ELKNPVINGKQCGITI--------TTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFK 250
I G+ + + Q S TL++ + + +E LKE + G +
Sbjct: 541 SCNKMEIEGRTIRLELQGPRGSPNARSQPSKTLFVKGLSEDTTEETLKESFE--GSVRAR 598
Query: 251 DLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV 290
+T D G++ G F++F+S D+K A + ++ ++
Sbjct: 599 IVT----DRETGSSKGFGFVDFNSEEDAKAAKEAMEDGEI 634
>NUCL_RAT (P13383) Nucleolin (Protein C23)
Length = 712
Score = 75.9 bits (185), Expect = 4e-13
Identities = 100/430 (23%), Positives = 176/430 (40%), Gaps = 78/430 (18%)
Query: 9 ADNGEEVK-----ESIDEYEKDEHLDFEDNYHEYDPEEYVG------------------D 45
A NG+ K E DE ++D+ + ED E++P G +
Sbjct: 133 AKNGKNAKKEDSDEDEDEEDEDDSDEDEDEEDEFEPPVVKGVKPAKAAPAAPASEDEDEE 192
Query: 46 DDYDERGIEQDEGQEAGDEVEEEPEE-------------------NVDEEEGDTGDEEVE 86
DD DE + DE +E D+ EEE E +V EEE D D+E E
Sbjct: 193 DDDDEDDDDDDEEEEEEDDSEEEVMEITPAKGKKTPAKVVPVKAKSVAEEEEDDEDDEDE 252
Query: 87 EVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEE-EQVEVIKETQKQK-------- 137
E +E E D++D +DE EE V K+E + + E +KQK
Sbjct: 253 E----EDEDEEDEEDDEDEDEEEEEEPVKAAPGKRKKEMTKQKEAPEAKKQKIEGSEPTT 308
Query: 138 ELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRAL 197
+F+G L+ + +L+ S++ ++ V+ +T N+ F + FE+ E +++AL
Sbjct: 309 PFNLFIGNLNPNKSVAELKVAISELFAKNDL-AAVDVRTGTNRKFGYVDFESAEDLEKAL 367
Query: 198 AELKNPVINGKQCGITITTCQDSD------TLYLDNICKSWKKEALKEKLKHYGVESFKD 251
EL + G + + +DS TL N+ + ++ LK E F+D
Sbjct: 368 -ELTGLKVFGNEIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELK--------EVFED 418
Query: 252 LTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV---VFGVDKPAEVSFANSFID 308
+ + +G + G A++EF S +D++ + Q ++ + E
Sbjct: 419 AVEIRLVSQDGRSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEKGQRQERTG 478
Query: 309 LGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGT 368
+ KT+ + L S E+ ++ + +K +++ +N P + K Y F+ F +
Sbjct: 479 KNSTWSGESKTLVLSNLSYSATEETLQEVFEK---ATFIKVPQN-PHGKSKGYAFIEFAS 534
Query: 369 HAAAVECAES 378
A E S
Sbjct: 535 FEDAKEALNS 544
Score = 59.7 bits (143), Expect = 3e-08
Identities = 94/432 (21%), Positives = 163/432 (36%), Gaps = 86/432 (19%)
Query: 17 ESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGD------EVEEEPE 70
+S+ E E+D+ D ED + D E+ D+D DE E+ G + +E PE
Sbjct: 237 KSVAEEEEDDE-DDEDEEEDEDEEDEEDDEDEDEEEEEEPVKAAPGKRKKEMTKQKEAPE 295
Query: 71 ENVDEEEGD-------------TGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENV--- 114
+ EG ++ V E++ E+ +D A D +
Sbjct: 296 AKKQKIEGSEPTTPFNLFIGNLNPNKSVAELKVAISELFAKNDLAAVDVRTGTNRKFGYV 355
Query: 115 -HEHLADVKEEEQVEVIKETQKQKELE---------------VFVGGLDKEATEHDLRKV 158
E D+++ ++ +K + +LE + L TE +L++V
Sbjct: 356 DFESAEDLEKALELTGLKVFGNEIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELKEV 415
Query: 159 FSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGK---------- 208
F EI V Q R+KG A + F++ ++ L E + I+G+
Sbjct: 416 FEDAVEIRLVS-----QDGRSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEK 470
Query: 209 -----QCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLE-DDNNEG 262
+ G T +S TL L N+ S +E L+E F+ T ++ N G
Sbjct: 471 GQRQERTGKNSTWSGESKTLVLSNLSYSATEETLQEV--------FEKATFIKVPQNPHG 522
Query: 263 TNCGCAFLEFSSHSDSKDAYKRLQKTDVV-----FGVDKPAEVSFANSFIDLGDDIMAQV 317
+ G AF+EF+S D+K+A K ++ + P A S
Sbjct: 523 KSKGYAFIEFASFEDAKEALNSCNKMEIEGRTIRLELQGPRGSPNARS---------QPS 573
Query: 318 KTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAE 377
KT+F+ L E+ L + + + + + K +GFV F + A E
Sbjct: 574 KTLFVKGLSEDTTEE---TLKESFEGSVRARIVTDRETGSSKGFGFVDFNSEEDAKAAKE 630
Query: 378 SITSAGLGEGNK 389
++ + +GNK
Sbjct: 631 AMEDGEI-DGNK 641
>NSR1_YEAST (P27476) Nuclear localization sequence binding protein
(P67)
Length = 414
Score = 75.9 bits (185), Expect = 4e-13
Identities = 63/292 (21%), Positives = 123/292 (41%), Gaps = 28/292 (9%)
Query: 17 ESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEE 76
ES E E + + + + D E +++E +++ + + +EE
Sbjct: 44 ESESESESESESSSSSSSSDSESSSSSSSDSESEAETKKEESKDSSSSSSDSSSDEEEEE 103
Query: 77 EGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQ 136
E + +E + + D + E ++++ D +EEE E + QK
Sbjct: 104 EKEETKKEESKESSSSDSSSSSSSDSESEKEESNDKKRKSE--DAEEEEDEESSNKKQKN 161
Query: 137 KELE----VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEH 192
+E E +FVG L + L+K F +G + R+ T R++G+ + FE +
Sbjct: 162 EETEEPATIFVGRLSWSIDDEWLKKEFEHIGGVIGARVIYERGTDRSRGYGYVDFENKSY 221
Query: 193 VKRALAELKNPVINGKQCGITITT-----------------CQDSDTLYLDNICKSWKKE 235
++A+ E++ I+G+ ++T + SDTL+L N+ + ++
Sbjct: 222 AEKAIQEMQGKEIDGRPINCDMSTSKPAGNNDRAKKFGDTPSEPSDTLFLGNLSFNADRD 281
Query: 236 ALKEKL-KHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQ 286
A+ E KH V S + T E + + G +++FS+ D+K A LQ
Sbjct: 282 AIFELFAKHGEVVSVRIPTHPETEQPK----GFGYVQFSNMEDAKKALDALQ 329
Score = 51.2 bits (121), Expect = 9e-06
Identities = 48/246 (19%), Positives = 99/246 (39%), Gaps = 50/246 (20%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEE 68
+ + EE +E +E +K+E + + D D E+ D+ +++ D EEE
Sbjct: 95 SSSDEEEEEEKEETKKEESKESSSS----DSSSSSSSDSESEKEESNDKKRKSEDAEEEE 150
Query: 69 PEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEH-------------ASEENVH 115
EE+ ++++ + EE + DD+ + + EH ++ +
Sbjct: 151 DEESSNKKQKNEETEEPATIFVGRLSWSIDDEWLKKEFEHIGGVIGARVIYERGTDRSRG 210
Query: 116 EHLADVKEEEQVEVIKETQKQKELE---------------------------------VF 142
D + + E + + KE++ +F
Sbjct: 211 YGYVDFENKSYAEKAIQEMQGKEIDGRPINCDMSTSKPAGNNDRAKKFGDTPSEPSDTLF 270
Query: 143 VGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKN 202
+G L A + ++F+K GE+ VR+ +P+T++ KGF ++F +E K+AL L+
Sbjct: 271 LGNLSFNADRDAIFELFAKHGEVVSVRIPTHPETEQPKGFGYVQFSNMEDAKKALDALQG 330
Query: 203 PVINGK 208
I+ +
Sbjct: 331 EYIDNR 336
Score = 45.8 bits (107), Expect = 4e-04
Identities = 75/370 (20%), Positives = 128/370 (34%), Gaps = 59/370 (15%)
Query: 60 EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLA 119
E+ E E + + E + E E ++ E D + D + EE E
Sbjct: 48 ESESESESSSSSSSSDSESSSSSSSDSESEAETKKEESKDSSSSSSDSSSDEEEEEE--- 104
Query: 120 DVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRN 179
KEE + E KE+ + E E + +K S+ E E + N + K
Sbjct: 105 --KEETKKEESKESSSSDSSSSSSSDSESEKEESNDKKRKSEDAEEEEDEESSNKKQKNE 162
Query: 180 KGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKE 239
+ ++ T+++ + S E LK+
Sbjct: 163 E------------------------------------TEEPATIFVGRLSWSIDDEWLKK 186
Query: 240 KLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVV-------F 292
+ +H G ++ + + + G +++F + S ++ A + +Q ++
Sbjct: 187 EFEHIG--GVIGARVIYERGTDRSR-GYGYVDFENKSYAEKAIQEMQGKEIDGRPINCDM 243
Query: 293 GVDKPA-EVSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAK 351
KPA A F GD T+F+ L + D D + L K+G V V +
Sbjct: 244 STSKPAGNNDRAKKF---GDTPSEPSDTLFLGNLSFNADRDAIFELFAKHGEVVSVRIPT 300
Query: 352 NMPGARRKNYGFVTFGTHAAAVECAESITSAGLGEGNKKAKVRARLSRPLPKVRGKHVPH 411
+ + K +G+V F +E A+ A GE VR S P P G
Sbjct: 301 HPETEQPKGFGYVQFSN----MEDAKKALDALQGEYIDNRPVRLDFSSPRPNNDGGRGGS 356
Query: 412 RDISGRKIGR 421
R GR GR
Sbjct: 357 RGFGGRGGGR 366
>ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hnRNP
G) (RNA binding motif protein, X chromosome)
(Glycoprotein p43)
Length = 391
Score = 73.9 bits (180), Expect = 1e-12
Identities = 122/425 (28%), Positives = 157/425 (36%), Gaps = 62/425 (14%)
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
+FI L +E + A+ KYG + +V L K+ + + + FVTF + A A + A +
Sbjct: 10 LFIGGLNTETNEKALEAVFGKYGRIVEVLLMKDRETNKSRGFAFVTFESPADAKDAARDM 69
Query: 380 TSAGLGEGNKKAKVRARLS--------RPLPKVRGKHVPHRDISGRKIGRLERPSRSRPA 431
L K + + S P P+ RG P R + G GR P
Sbjct: 70 NGKSLDGKAIKVEQATKPSFESGRRGPPPPPRSRG---PPRGLRG---GRGGSGGTRGPP 123
Query: 432 PRSRPAPQSRPAHVFRRLGSR--IPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAA 489
R + F SR +P R R+ P P R+ P S P RS +
Sbjct: 124 SRGGHMDDGGYSMNFNMSSSRGPLPVKRGPPPRSGGPP---PKRSAP-SGPVRSSSGMGG 179
Query: 490 AYPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYR---DYPARDSGYPDLHRDT 546
P S + YG P P SR +S R Y Y +RD YP
Sbjct: 180 RAPVSRGRDSYGGPPRREPLPSRRDV-----YLSPRDDGYSTKDSYSSRD--YPSSRDTR 232
Query: 547 SRTAPRRGYLDDGYDRRLERPPSPSPRLSYRE--GRPRDY-DALSGSKRPYAAIDDISPR 603
P R Y Y R PS S R+ GR RDY D SG + D
Sbjct: 233 DYAPPPRDYTYRDYGHSSSRDDYPSRGYSDRDGYGRDRDYSDHPSG-----GSYRDSYES 287
Query: 604 YADAGARQSRSRLDYDYGGSASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQD 663
Y ++ + YGGS SRY D S GY GSR S S+ S YSS
Sbjct: 288 YGNSRSAPPTRGPPPSYGGS-SRY-----DDYSSSRDGYGGSRDSYSSSRS-DLYSS--- 337
Query: 664 PSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGSSYSSMYSSRGM---NGSSSYMSGSGG 720
GR +G + G+ S + RG SYSS SSRG G S GG
Sbjct: 338 ---GRDRVGRQERGLPPS------MERGYPPPRDSYSS--SSRGAPRGGGRGGSRSDRGG 386
Query: 721 GSRSY 725
G Y
Sbjct: 387 GRSRY 391
Score = 51.6 bits (122), Expect = 7e-06
Identities = 24/69 (34%), Positives = 40/69 (57%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++F+GGL+ E E L VF K G I EV + + +T +++GFA + FE+ K A +
Sbjct: 9 KLFIGGLNTETNEKALEAVFGKYGRIVEVLLMKDRETNKSRGFAFVTFESPADAKDAARD 68
Query: 200 LKNPVINGK 208
+ ++GK
Sbjct: 69 MNGKSLDGK 77
>NUCL_HUMAN (P19338) Nucleolin (Protein C23)
Length = 706
Score = 71.2 bits (173), Expect = 9e-12
Identities = 92/426 (21%), Positives = 172/426 (39%), Gaps = 73/426 (17%)
Query: 9 ADNGEEVK--ESIDEYEKDEHLDFEDNYHEYDPEEYV---------------GDDDYDER 51
A NG+ K +S +E + D D ED+ E + E+ + +DD D+
Sbjct: 133 AKNGKNAKKEDSDEEEDDDSEEDEEDDEDEDEDEDEIEPAAMKAAAAAPASEDEDDEDDE 192
Query: 52 GIEQDEGQEAGDEVEEEPEEN----------VDEEEGDTGDEEVEEVEYVYEEVEGDDDD 101
E D+ E D EE E V + + ++E EE + E+ + D+DD
Sbjct: 193 DDEDDDDDEEDDSEEEAMETTPAKGKKAAKVVPVKAKNVAEDEDEEEDDEDEDDDDDEDD 252
Query: 102 VAGDDEHASEENVHEHLADVKE----------------EEQVEVIKETQKQKELEVFVGG 145
DDE EE E VKE E + + ++ T+ +FVG
Sbjct: 253 EDDDDEDDEEEEEEEEEEPVKEAPGKRKKEMAKQKAAPEAKKQKVEGTEPTTAFNLFVGN 312
Query: 146 LDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVI 205
L+ + +L+ S V ++ + V+ + + F + FE+ E +++AL EL +
Sbjct: 313 LNFNKSAPELKTGISDVFAKNDLAV-VDVRIGMTRKFGYVDFESAEDLEKAL-ELTGLKV 370
Query: 206 NGKQCGITITTCQDSD------TLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDDN 259
G + + +DS TL N+ ++ LK E F+D + +
Sbjct: 371 FGNEIKLEKPKGKDSKKERDARTLLAKNLPYKVTQDELK--------EVFEDAAEIRLVS 422
Query: 260 NEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV-------VFGVDKPAEVSFANSFIDLGDD 312
+G + G A++EF + +D++ ++ Q T++ + +K +
Sbjct: 423 KDGKSKGIAYIEFKTEADAEKTFEEKQGTEIDGRSISLYYTGEKGQNQDYRGG---KNST 479
Query: 313 IMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAA 372
+ KT+ + L S E+ ++ + +K +++ +N G + K Y F+ F + A
Sbjct: 480 WSGESKTLVLSNLSYSATEETLQEVFEK---ATFIKVPQNQNG-KSKGYAFIEFASFEDA 535
Query: 373 VECAES 378
E S
Sbjct: 536 KEALNS 541
Score = 63.2 bits (152), Expect = 2e-09
Identities = 94/435 (21%), Positives = 162/435 (36%), Gaps = 108/435 (24%)
Query: 31 EDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEV-- 88
ED E D E+ DDD D+ E D+ ++ +E EEE EE V E G E ++
Sbjct: 233 EDEDEEEDDEDEDDDDDEDD---EDDDDEDDEEEEEEEEEEPVKEAPGKRKKEMAKQKAA 289
Query: 89 -EYVYEEVEGDDDDVAGD------DEHASEENVHEHLADVKEEEQVEVI----------- 130
E ++VEG + A + + + S + ++DV + + V+
Sbjct: 290 PEAKKQKVEGTEPTTAFNLFVGNLNFNKSAPELKTGISDVFAKNDLAVVDVRIGMTRKFG 349
Query: 131 ---------------------------------KETQKQKELEVFVG-GLDKEATEHDLR 156
K+++K+++ + L + T+ +L+
Sbjct: 350 YVDFESAEDLEKALELTGLKVFGNEIKLEKPKGKDSKKERDARTLLAKNLPYKVTQDELK 409
Query: 157 KVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQC------ 210
+VF EI V + ++KG A + F+T ++ E + I+G+
Sbjct: 410 EVFEDAAEIRLVS-----KDGKSKGIAYIEFKTEADAEKTFEEKQGTEIDGRSISLYYTG 464
Query: 211 ----------GITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLE-DDN 259
G T +S TL L N+ S +E L+E F+ T ++ N
Sbjct: 465 EKGQNQDYRGGKNSTWSGESKTLVLSNLSYSATEETLQEV--------FEKATFIKVPQN 516
Query: 260 NEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVV-----FGVDKPAEVSFANSFIDLGDDIM 314
G + G AF+EF+S D+K+A K ++ + P A S
Sbjct: 517 QNGKSKGYAFIEFASFEDAKEALNSCNKREIEGRAIRLELQGPRGSPNARS--------- 567
Query: 315 AQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVE 374
KT+F+ L E+ L + + + + + K +GFV F + A E
Sbjct: 568 QPSKTLFVKGLSEDTTEE---TLKESFDGSVRARIVTDRETGSSKGFGFVDFNSEEDAKE 624
Query: 375 CAESITSAGLGEGNK 389
E G +GNK
Sbjct: 625 AMED----GEIDGNK 635
>NUCL_MOUSE (P09405) Nucleolin (Protein C23)
Length = 706
Score = 70.9 bits (172), Expect = 1e-11
Identities = 95/452 (21%), Positives = 184/452 (40%), Gaps = 76/452 (16%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEE 68
A NG+ K+ + ++DE D +D+ + D EE +D+++ ++ + +A
Sbjct: 133 AKNGKNAKKEDSDEDEDEE-DEDDSDEDEDDEE---EDEFEPPIVKGVKPAKAAPAAPAS 188
Query: 69 PEENVDEEEGDTGDEEVEEVEYVYEEV-------------------------EGDDDDVA 103
+E DE+E D D++ EE + EEV E DD++
Sbjct: 189 EDEEDDEDEDDEEDDDEEEEDDSEEEVMEITTAKGKKTPAKVVPMKAKSVAEEEDDEEED 248
Query: 104 GDDEHASEENVHEHLADVKEEEQVEV-------IKETQKQKE-----------------L 139
DDE +E + D +EEE+ V KE KQKE
Sbjct: 249 EDDEDEDDEEEDDEDDDEEEEEEEPVKAAPGKRKKEMTKQKEAPEAKKQKVEGSEPTTPF 308
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+F+G L+ + ++L+ S++ ++ + V+ +T N+ F + FE+ E +++AL E
Sbjct: 309 NLFIGNLNPNKSVNELKFAISELFAKNDLAV-VDVRTGTNRKFGYVDFESAEDLEKAL-E 366
Query: 200 LKNPVINGKQCGITITTCQDSD------TLYLDNICKSWKKEALKEKLKHYGVESFKDLT 253
L + G + + +DS TL N+ + ++ LK E F+D
Sbjct: 367 LTGLKVFGNEIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELK--------EVFEDAM 418
Query: 254 LLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV---VFGVDKPAEVSFANSFIDLG 310
+ + +G + G A++EF S +D++ + Q ++ + E
Sbjct: 419 EIRLVSQDGKSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTGEKGQRQERTGKT 478
Query: 311 DDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHA 370
+ KT+ + L S ++ + + +K +++ +N P + K Y F+ F +
Sbjct: 479 STWSGESKTLVLSNLSYSATKETLEEVFEK---ATFIKVPQN-PHGKPKGYAFIEFASFE 534
Query: 371 AAVECAESITSAGLGEGNKKAKVRARLSRPLP 402
A E S + + +++ SR P
Sbjct: 535 DAKEALNSCNKMEIEGRTIRLELQGSNSRSQP 566
Score = 65.9 bits (159), Expect = 4e-10
Identities = 99/429 (23%), Positives = 168/429 (39%), Gaps = 85/429 (19%)
Query: 17 ESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGD--------EVEEE 68
+S+ E E DE D ED+ E D EE DDD +E E++E +A + +E
Sbjct: 236 KSVAEEEDDEEED-EDDEDEDDEEEDDEDDDEEE---EEEEPVKAAPGKRKKEMTKQKEA 291
Query: 69 PEENVDEEEGD-------------TGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENV- 114
PE + EG ++ V E+++ E+ +D D +
Sbjct: 292 PEAKKQKVEGSEPTTPFNLFIGNLNPNKSVNELKFAISELFAKNDLAVVDVRTGTNRKFG 351
Query: 115 ---HEHLADVKEEEQVEVIKETQKQKELE---------------VFVGGLDKEATEHDLR 156
E D+++ ++ +K + +LE + L TE +L+
Sbjct: 352 YVDFESAEDLEKALELTGLKVFGNEIKLEKPKGRDSKKVRAARTLLAKNLSFNITEDELK 411
Query: 157 KVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGK-------- 208
+VF EI V Q ++KG A + F++ ++ L E + I+G+
Sbjct: 412 EVFEDAMEIRLVS-----QDGKSKGIAYIEFKSEADAEKNLEEKQGAEIDGRSVSLYYTG 466
Query: 209 -------QCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLE-DDNN 260
+ G T T +S TL L N+ S KE L+E F+ T ++ N
Sbjct: 467 EKGQRQERTGKTSTWSGESKTLVLSNLSYSATKETLEEV--------FEKATFIKVPQNP 518
Query: 261 EGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFIDLGDDIMAQVKTV 320
G G AF+EF+S D+K+A K ++ G E+ +NS KT+
Sbjct: 519 HGKPKGYAFIEFASFEDAKEALNSCNKMEIE-GRTIRLELQGSNS-------RSQPSKTL 570
Query: 321 FIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESIT 380
F+ L E+ L + + + + + K +GFV F + A E++
Sbjct: 571 FVKGLSEDTTEE---TLKESFEGSVRARIVTDRETGSSKGFGFVDFNSEEDAKAAKEAME 627
Query: 381 SAGLGEGNK 389
+ +GNK
Sbjct: 628 DGEI-DGNK 635
>ROG_MOUSE (O35479) Heterogeneous nuclear ribonucleoprotein G (hnRNP
G) (RNA binding motif protein, X chromosome)
Length = 388
Score = 69.7 bits (169), Expect = 3e-11
Identities = 122/427 (28%), Positives = 156/427 (35%), Gaps = 69/427 (16%)
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
+FI L +E + A+ KYG + ++ L K+ + + + FVT + A A + A +
Sbjct: 10 LFIGGLNTETNEKALEAVFGKYGRIVEIILMKDRETNKSRGFAFVTLESPADAKDAARDM 69
Query: 380 TSAGLGEGNKKAKVR----------ARLSRPLPKVRGKHVPHRDISGRKIGRLERPSRSR 429
L K KV R P P+ RG P R + G G PSR
Sbjct: 70 NGKSL--DGKAIKVEQATKPFFESGRRGPPPPPRSRG---PPRGLRGGSGGTRGPPSRGG 124
Query: 430 PAPRSRPAPQSRPAHVFRRLGSR--IPPARPSSARNRRPVTSIPVRARPASPPARSYRRL 487
+ F SR +P R R+ P P R+ P S P RS +
Sbjct: 125 YMDDGGYSMN------FNMSSSRGPLPVKRGPPPRSGGPP---PKRSTP-SGPVRSSSGM 174
Query: 488 AAAYPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYR---DYPARDSGYPDLHR 544
P S + YG P P SR +S R Y Y +RD YP
Sbjct: 175 GGRTPVSRGRDSYGGPPRREPLPSRRDV-----YLSPRDDGYSTKDSYSSRD--YPSSRD 227
Query: 545 DTSRTAPRRGYLDDGYDRRLERPPSPSPRLSYRE--GRPRDY-DALSGSKRPYAAIDDIS 601
T P R Y Y R PS R+ GR RDY D SG + D
Sbjct: 228 TRDYTPPPRDYTYRDYCHSSSRDDYPSRGYGDRDGYGRDRDYSDHPSG-----GSYRDSY 282
Query: 602 PRYADAGARQSRSRLDYDYGGSASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSR 661
Y ++ + YGGS SRY D S GY GSR S S+ S YSS
Sbjct: 283 ESYGNSRSAPPTRGPPPSYGGS-SRY-----DDYSSSRDGYGGSRDSYSSSRS-DLYSS- 334
Query: 662 QDPSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGSSYSSMYSSRGM---NGSSSYMSGS 718
GR +G + G+ S + RG SYSS SSRG G S
Sbjct: 335 -----GRDRVGRQERGLPPS------MERGYPPPRDSYSS--SSRGAPRGGGRGGSRSDR 381
Query: 719 GGGSRSY 725
GGG Y
Sbjct: 382 GGGRSRY 388
Score = 48.5 bits (114), Expect = 6e-05
Identities = 22/69 (31%), Positives = 39/69 (55%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++F+GGL+ E E L VF K G I E+ + + +T +++GFA + E+ K A +
Sbjct: 9 KLFIGGLNTETNEKALEAVFGKYGRIVEIILMKDRETNKSRGFAFVTLESPADAKDAARD 68
Query: 200 LKNPVINGK 208
+ ++GK
Sbjct: 69 MNGKSLDGK 77
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 68.9 bits (167), Expect = 4e-11
Identities = 114/473 (24%), Positives = 190/473 (40%), Gaps = 123/473 (26%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGI--------------- 53
A NG+ K+ E+ E D +D E D +E D+ +E +
Sbjct: 105 AKNGKNAKK-----EESEEEDEDDEDDEEDEDEEEESDEEEEPAVPVKPAAKKSAAAVPA 159
Query: 54 ----------EQDEGQEAGDEVEEEPEENVDEEEGDTG---------------------- 81
E +E +E DE E+E ++ ++E DT
Sbjct: 160 KKPAVVPAKQESEEEEEEDDEEEDEEDDESEDEAMDTTPAPVKKPTPAKATPAKAKAESE 219
Query: 82 DEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQK---- 137
DEE EE E E+ E +DD+ ++E E+ V E K+E + E +K+K
Sbjct: 220 DEEDEEDED--EDEEDEDDEEEDEEESEDEKPVKEAPGKRKKEMANKSAPEAKKKKTETP 277
Query: 138 --ELEVFVGGL----DKEATEHDLRKVFSKVG-EITEVRMTVNPQTKRNKGFALLRFETV 190
+FV L D E +++ F K +++EVR+ +K F + F +
Sbjct: 278 ASAFSLFVKNLTPTKDYEELRTAIKEFFGKKNLQVSEVRI------GSSKRFGYVDFLSA 331
Query: 191 EHVKRALAELKNPVINGKQ-CGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESF 249
E + +AL +NGK+ G+ I + K+ KE+LKE K +
Sbjct: 332 EDMDKALQ------LNGKKLMGLEI------------KLEKAKSKESLKENKKERDARTL 373
Query: 250 --KDL--TLLEDD---------------NNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDV 290
K+L + ED+ N EG++ G A++EF + ++++ A + Q T+V
Sbjct: 374 FVKNLPYRVTEDEMKNVFENALEVRLVLNKEGSSKGMAYIEFKTEAEAEKALEEKQGTEV 433
Query: 291 ---VFGVDKPAEVSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKV 347
+D E S S G+ + KT+ ++ L + E+ ++ L KK +
Sbjct: 434 DGRAMVIDYTGEKSQQESQKGGGE---RESKTLIVNNLSYAASEETLQELFKK---ATSI 487
Query: 348 ELAKNMPGARRKNYGFVTFGTHAAAVECAESITSAGLGEGNKKAKVRARLSRP 400
++ +N G R K Y FV F T A E S + + EG +R S P
Sbjct: 488 KMPQNNQG-RPKGYAFVEFPTAEDAKEALNSCNNTEI-EGR---AIRLEFSSP 535
Score = 57.0 bits (136), Expect = 2e-07
Identities = 48/182 (26%), Positives = 85/182 (46%), Gaps = 24/182 (13%)
Query: 123 EEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGF 182
E+ Q E K +++ + V L A+E L+++F K T ++M N Q R KG+
Sbjct: 445 EKSQQESQKGGGERESKTLIVNNLSYAASEETLQELFKKA---TSIKMPQNNQG-RPKGY 500
Query: 183 ALLRFETVEHVKRALAELKNPVINGKQCGITITTC--------------QDSDTLYLDNI 228
A + F T E K AL N I G+ + ++ Q S TL++ +
Sbjct: 501 AFVEFPTAEDAKEALNSCNNTEIEGRAIRLEFSSPSWQKGNMNARGGFNQQSKTLFVRGL 560
Query: 229 CKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKT 288
+ +E L+E + G S + +T D + G++ G F++FSS D+K A + ++
Sbjct: 561 SEDTTEETLRESFE--GSISARIVT----DRDTGSSKGFGFVDFSSPEDAKAAKEAMEDG 614
Query: 289 DV 290
++
Sbjct: 615 EI 616
Score = 36.6 bits (83), Expect = 0.24
Identities = 23/74 (31%), Positives = 39/74 (52%), Gaps = 3/74 (4%)
Query: 136 QKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKR 195
Q+ +FV GL ++ TE LR+ F G I+ R+ + T +KGF + F + E K
Sbjct: 550 QQSKTLFVRGLSEDTTEETLRESFE--GSIS-ARIVTDRDTGSSKGFGFVDFSSPEDAKA 606
Query: 196 ALAELKNPVINGKQ 209
A +++ I+G +
Sbjct: 607 AKEAMEDGEIDGNK 620
>PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4
(Poly(A)-binding protein 4) (PABP 4) (Inducible
poly(A)-binding protein) (iPABP) (Activated-platelet
protein-1) (APP-1)
Length = 644
Score = 68.6 bits (166), Expect = 6e-11
Identities = 64/285 (22%), Positives = 129/285 (44%), Gaps = 25/285 (8%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++VG L + TE L + FS G + +R+ + T+R+ G+A + F+ +RAL +
Sbjct: 13 LYVGDLHSDVTEAMLYEKFSPAGPVLSIRVCRDMITRRSLGYAYVNFQQPADAERALDTM 72
Query: 201 KNPVINGKQCGITITTCQDS------DTLYLDNICKSWKKEALKEKLKHYGVESFKDLTL 254
VI GK I + S +++ N+ KS +AL + +G + +
Sbjct: 73 NFDVIKGKPIRIMWSQRDPSLRKSGVGNVFIKNLDKSIDNKALYDTFSAFG--NILSCKV 130
Query: 255 LEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFIDLGDDIM 314
+ D+N + G AF+ F + + A K ++K + + D+ V S + ++
Sbjct: 131 VCDENG---SKGYAFVHFET---QEAADKAIEKMNGMLLNDRKVFVGRFKSRKEREAELG 184
Query: 315 AQVK---TVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAA 371
A+ K V+I D++ ++ L ++G V++ ++ P + K +GFV++ H
Sbjct: 185 AKAKEFTNVYIKNFGEEVDDESLKELFSQFGKTLSVKVMRD-PNGKSKGFGFVSYEKHED 243
Query: 372 AVECAESITSAGL-------GEGNKKAKVRARLSRPLPKVRGKHV 409
A + E + + G KK + +A L R +++ + +
Sbjct: 244 ANKAVEEMNGKEISGKIIFVGRAQKKVERQAELKRKFEQLKQERI 288
Score = 44.7 bits (104), Expect = 9e-04
Identities = 27/117 (23%), Positives = 55/117 (46%), Gaps = 10/117 (8%)
Query: 119 ADVKEEEQVEVIKETQKQKE--------LEVFVGGLDKEATEHDLRKVFSKVGEITEVRM 170
A K E Q E+ ++ ++ K+ + +++ LD + LRK FS G IT ++
Sbjct: 266 AQKKVERQAELKRKFEQLKQERISRYQGVNLYIKNLDDTIDDEKLRKEFSPFGSITSAKV 325
Query: 171 TVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDN 227
+ + R+KGF + F + E +A+ E+ ++ K + + ++ +L N
Sbjct: 326 ML--EDGRSKGFGFVCFSSPEEATKAVTEMNGRIVGSKPLYVALAQRKEERKAHLTN 380
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 68.6 bits (166), Expect = 6e-11
Identities = 49/167 (29%), Positives = 89/167 (52%), Gaps = 10/167 (5%)
Query: 3 LNIYLIADNGEEVKESIDEYE--KDEHLDFEDNYHEYDPE--EYVGDDDYDERGIEQDEG 58
+N+ DN ++ I+E E K +H+D E++ E E E + DE +E+DE
Sbjct: 518 VNVVPRRDNHKKKMAKIEEAELQKQKHVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEE 577
Query: 59 QEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHL 118
+E +E EEE EE +EEE + +EE E+ + E+ +D+D A +DE +EE+ E
Sbjct: 578 EEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEED 637
Query: 119 ADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEI 165
D +++++ E E +++E E ++E +E +++ K +I
Sbjct: 638 DDEEDDDEDEDEDEEDEEEEEE------EEEESEKKIKRNLRKNAKI 678
Score = 33.9 bits (76), Expect = 1.6
Identities = 33/142 (23%), Positives = 61/142 (42%), Gaps = 4/142 (2%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
+E K+ + E +E + D E E + + E + G+ EEE +E
Sbjct: 328 KEKKKKKHDKENEETMQQPDQTSEETNNEIMVPLPSPLTDVTTPEEHKEGEHKEEEHKEG 387
Query: 73 VDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKE 132
+ +EG+ +EE +E E+ EE + + G + ++ +H KE+ + V+K
Sbjct: 388 -EHKEGEHKEEEHKEEEHKKEEHKSKEHKSKGKKDKGKKDK-GKHKKAKKEKVKKHVVKN 445
Query: 133 T--QKQKELEVFVGGLDKEATE 152
+ K+ + DKEA E
Sbjct: 446 VIEDEDKDGVEIINLEDKEACE 467
Score = 32.0 bits (71), Expect = 5.9
Identities = 47/234 (20%), Positives = 86/234 (36%), Gaps = 46/234 (19%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDE-------GQEAGDEV 65
++ KE ++ +K E D ++ H+ + + D++ E E GQ
Sbjct: 130 KDKKEKKEKKDKKEKKDKKEKKHKKEKKH-----KKDKKKKENSEVMSLYKTGQHKPKNA 184
Query: 66 EEEPEENVDEEEGDTGDEEVE-----EVEYVYEEVEG----------DDDDVAGDDEHAS 110
E EEN+DEE + + Y Y E G +D D E+ S
Sbjct: 185 TEHGEENLDEEMVSEINNNAQGGLLLSSPYQYREQGGCGIISSVHETSNDTKDNDKENIS 244
Query: 111 EENVHEHLAD-----------------VKEEEQVEVIKETQKQKELEVFVGGLDKEATEH 153
E+ +H + +KE+E++E K+ Q++KE + K+ +
Sbjct: 245 EDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEEKEKKKQEKERKKQEKKE 304
Query: 154 --DLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVI 205
K K +I + R + K+ K ET++ + E N ++
Sbjct: 305 RKQKEKEMKKQKKIEKERKKKEEKEKKKKKHDKENEETMQQPDQTSEETNNEIM 358
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 68.2 bits (165), Expect = 7e-11
Identities = 71/319 (22%), Positives = 130/319 (40%), Gaps = 36/319 (11%)
Query: 13 EEVKESIDEYEKD-EHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPE- 70
EE KES E E + E+ + + ++ D E + E + + E EEE E
Sbjct: 131 EEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEV 190
Query: 71 --ENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLAD-VKEEEQV 127
+ +++EG + E GD D + + +S E+ + A+ EE
Sbjct: 191 VEKTEEKKEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAEPASEERPA 250
Query: 128 EVIKETQKQKEL-EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLR 186
++ K +Q E VFVG L + L + F + G I R+ ++ Q+ R+KG+ +
Sbjct: 251 KITKPSQDSNETCTVFVGRLSWNVDDQWLGQEFEEYGTIVGARVIMDGQSGRSKGYGYVD 310
Query: 187 FETVEHVKRALAELKNPVINGKQCGITITT---------------------CQDSDTLYL 225
FET E K A+A I+G+ + ++ + SDT+++
Sbjct: 311 FETPEAAKAAVAANGTKEIDGRMVNLDLSNPRPANPQPYAQQRAGNFGDQLSEPSDTVFV 370
Query: 226 DNICKSWKKEALKEKLKHYG-VESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKR 284
N+ + ++ L G ++S + L D G G ++ FS D+ K+
Sbjct: 371 GNLSFNATEDDLSTAFGGCGDIQSIR----LPTDPQSGRLKGFGYVTFS----DIDSAKK 422
Query: 285 LQKTDVVFGVDKPAEVSFA 303
+ + F +P + F+
Sbjct: 423 CVEMNGHFIAGRPCRLDFS 441
Score = 53.5 bits (127), Expect = 2e-06
Identities = 27/76 (35%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
VFVG L ATE DL F G+I +R+ +PQ+ R KGF + F ++ K+ + E+
Sbjct: 368 VFVGNLSFNATEDDLSTAFGGCGDIQSIRLPTDPQSGRLKGFGYVTFSDIDSAKKCV-EM 426
Query: 201 KNPVINGKQCGITITT 216
I G+ C + +T
Sbjct: 427 NGHFIAGRPCRLDFST 442
Score = 52.8 bits (125), Expect = 3e-06
Identities = 72/371 (19%), Positives = 138/371 (36%), Gaps = 39/371 (10%)
Query: 20 DEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEV----EEEPEENVDE 75
+E + + + E E E + E+ +EV EE+ E + +
Sbjct: 81 EESSSESESESSSSESESSSSESESSSSESESSSSESSSSESEEEVIVKTEEKKESSSES 140
Query: 76 EEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQK 135
+EE E V + E+ E D E +S E+ E + EEE+ EV+++T++
Sbjct: 141 SSSSESEEEEEAVVKIEEKKESSSDS---SSESSSSESESESSSSESEEEE-EVVEKTEE 196
Query: 136 QKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKR 195
+KE + +++ S +E + + KR K
Sbjct: 197 KKEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKR---------------KA 241
Query: 196 ALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLL 255
A + P K + TC T+++ + + + L ++ + YG + +
Sbjct: 242 EPASEERPAKITKPSQDSNETC----TVFVGRLSWNVDDQWLGQEFEEYGTIVGARVIM- 296
Query: 256 EDDNNEGTNCGCAFLEFSSHSDSKDAY-----KRLQKTDVVFGVDKPAEVS---FANSFI 307
D G + G +++F + +K A K + V + P + +A
Sbjct: 297 --DGQSGRSKGYGYVDFETPEAAKAAVAANGTKEIDGRMVNLDLSNPRPANPQPYAQQRA 354
Query: 308 -DLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTF 366
+ GD + TVF+ L + ED + G ++ + L + R K +G+VTF
Sbjct: 355 GNFGDQLSEPSDTVFVGNLSFNATEDDLSTAFGGCGDIQSIRLPTDPQSGRLKGFGYVTF 414
Query: 367 GTHAAAVECAE 377
+A +C E
Sbjct: 415 SDIDSAKKCVE 425
Score = 34.3 bits (77), Expect = 1.2
Identities = 45/263 (17%), Positives = 104/263 (39%), Gaps = 12/263 (4%)
Query: 110 SEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVR 169
S+++V + K+EE + E E + ++E + S E +E
Sbjct: 67 SKKSVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSESSSSE-SEEE 125
Query: 170 MTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNIC 229
+ V + K+ E + A+ +++ + + S++ +
Sbjct: 126 VIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSS---SESSSSESESESSSS 182
Query: 230 KSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTD 289
+S ++E + EK + S + + E ++ + G + S S+S ++ +K +
Sbjct: 183 ESEEEEEVVEKTEEKKEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAE 242
Query: 290 VVFGVDKPAEVSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVEL 349
++PA+++ + + + TVF+ L + D+ ++ ++YG + +
Sbjct: 243 PA-SEERPAKITKPSQDSN-------ETCTVFVGRLSWNVDDQWLGQEFEEYGTIVGARV 294
Query: 350 AKNMPGARRKNYGFVTFGTHAAA 372
+ R K YG+V F T AA
Sbjct: 295 IMDGQSGRSKGYGYVDFETPEAA 317
>NUCL_XENLA (P20397) Nucleolin (Protein C23)
Length = 650
Score = 67.8 bits (164), Expect = 1e-10
Identities = 100/442 (22%), Positives = 172/442 (38%), Gaps = 81/442 (18%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYH--------EYDPEEYVGDDDYDERGIEQDEGQE 60
A +E +E DE +DE ++ E D EE +DD DE + +E Q+
Sbjct: 148 AGKKQESEEEDDEESEDEPMEVAPALKGKKTAQAAEEDDEE---EDDDDEEDDDDEEEQQ 204
Query: 61 AGDEVEEEPEENVDEEEG---DTGDE-------------EVEEVE-----------YVYE 93
+ ++E + + E + DT E E +E++ +
Sbjct: 205 GSAKRKKEMPKTIPEAKKTKTDTASEGLSIFIGNLNSTKEFDELKDALREFFSKKNLTIQ 264
Query: 94 EVEGDDDDVAGDDEHASEENVHEHL---------ADVKEEEQVEVIK----ETQKQKELE 140
++ + G + +SEE V + L +VK E+ + K E +K+++
Sbjct: 265 DIRIGNSKKFGYVDFSSEEEVEKALKLTGKKILGTEVKIEKAMAFDKNKTAENKKERDSR 324
Query: 141 -VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+FV + T +L+++F +I + NKG A + F + +AL E
Sbjct: 325 TLFVKNIPYSTTVEELQEIFENAKDIR----IPTGKDGSNKGIAYVEFSNEDEANKALEE 380
Query: 200 LKNPVINGKQCGITITTCQ------------DSDTLYLDNICKSWKKEALKEKLKHYGVE 247
+ I G+ + T + DS L ++N+ S +++L+E
Sbjct: 381 KQGAEIEGRSIFVDFTGEKSQNSGNKKGPEGDSKVLVVNNLSYSATEDSLREV------- 433
Query: 248 SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFI 307
F+ T + N+G G AF+EFSS D+KDA T++ G E S
Sbjct: 434 -FEKATSIRIPQNQGRAKGFAFIEFSSAEDAKDAMDSCNNTEIE-GRSIRLEFSQGGGPQ 491
Query: 308 DLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFG 367
G AQ KT+F+ L E+ L + + + + K +GFV F
Sbjct: 492 GGGRGGSAQSKTLFVRGLSEDTTEE---TLKEAFDGSVNARIVTDRDTGASKGFGFVDFS 548
Query: 368 THAAAVECAESITSAGLGEGNK 389
T A + A+ G +GNK
Sbjct: 549 T-AEDAKAAKEAMEDGEIDGNK 569
>VG48_SHV21 (Q01033) Hypothetical gene 48 protein
Length = 797
Score = 66.2 bits (160), Expect = 3e-10
Identities = 50/135 (37%), Positives = 70/135 (51%), Gaps = 6/135 (4%)
Query: 10 DNGEEVKESIDEYEK--DEHLDFEDNYHEYDPEEYVGD--DDYDERGIEQDEGQEAGDEV 65
D G+E E DE E DE + ED E D E GD D+ ++ G E DEG++ GDE
Sbjct: 454 DEGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEG 513
Query: 66 EEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEE 125
+E +E + +EGD GDE E E+ E EG+D+ G+DE E+ E D E+E
Sbjct: 514 DEGKDEGDEGDEGDEGDE--GEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDE 571
Query: 126 QVEVIKETQKQKELE 140
+ E + + E E
Sbjct: 572 GEDEGDEGEDEGEDE 586
Score = 62.0 bits (149), Expect = 5e-09
Identities = 47/119 (39%), Positives = 67/119 (55%), Gaps = 6/119 (5%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIE-QDEGQEAGDEVEEE 68
D G+E E DE+E DE + ED E + E G+D+ ++ G E +DEG++ GDE E+E
Sbjct: 524 DEGDEGDEGEDEWE-DEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDE 582
Query: 69 PEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD--DEHASEENVHEHLADVKEEE 125
E+ DE E D G++E +E E E+ EGD+ D D DE E + E D E+E
Sbjct: 583 GEDEGDEGE-DEGEDEGDEGEDEGED-EGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDE 639
Score = 60.8 bits (146), Expect = 1e-08
Identities = 52/134 (38%), Positives = 71/134 (52%), Gaps = 6/134 (4%)
Query: 10 DNGEEVKESIDEYEK-DEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEE 68
D G+E E DE ++ DE D D E E GD+ DE G E DEG E GDE E+E
Sbjct: 478 DEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEGDEGKDE-GDEGDEGDE-GDEGEDE 535
Query: 69 PEENVD--EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQ 126
E+ D E+EGD G++E +E E E+ EGD+ + G+DE E+ E D E+E
Sbjct: 536 WEDEGDEGEDEGDEGEDEGDEGEDEGED-EGDEGEDEGEDEGDEGEDEGEDEGDEGEDEG 594
Query: 127 VEVIKETQKQKELE 140
+ E + + E E
Sbjct: 595 EDEGDEGEDEGEDE 608
Score = 60.8 bits (146), Expect = 1e-08
Identities = 48/142 (33%), Positives = 75/142 (52%), Gaps = 4/142 (2%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP 69
D GE+ + ++ +DE + ED + E G+D+ DE E DEG++ GDE E+E
Sbjct: 581 DEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEG 640
Query: 70 EENVDEEEGDTGDEEVEEVEYVYEE-VEGDDDDVAGDDEHASEENVHEHLADVKE-EEQV 127
+E E+EGD G++E +E E +E EG+D+ G+DE E+ + + E +E
Sbjct: 641 DEG--EDEGDEGEDEGDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGDEGDEGD 698
Query: 128 EVIKETQKQKELEVFVGGLDKE 149
E E + + E E G DKE
Sbjct: 699 EGEDEGEDEGEDEGDEGTKDKE 720
Score = 60.5 bits (145), Expect = 2e-08
Identities = 43/106 (40%), Positives = 58/106 (54%), Gaps = 5/106 (4%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP-EE 71
EE + +E EKDE + ED E D E G+D+ ++ G E DEG E DE E+E EE
Sbjct: 417 EEDEGEDEEDEKDEKEEGED---EGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEE 473
Query: 72 NVDEEEGDTGDEEVEEVEYVYE-EVEGDDDDVAGDDEHASEENVHE 116
+ E+EGD GDE +E + E E EGD+ D D+ +E E
Sbjct: 474 DEGEDEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEGDEGKDE 519
Score = 59.7 bits (143), Expect = 3e-08
Identities = 46/117 (39%), Positives = 64/117 (54%), Gaps = 7/117 (5%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP 69
D GE+ E + E DE + ED + D EE G+D+ DE +DEG E GDE E+E
Sbjct: 443 DEGEDEGEDEGD-EGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDE-GDEGEDEG 500
Query: 70 EENVD-EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEE 125
+E + ++EGD GDE +E + E EGD+ D G+DE E + E D E+E
Sbjct: 501 DEGDEGKDEGDEGDEGKDEGD---EGDEGDEGD-EGEDEWEDEGDEGEDEGDEGEDE 553
Score = 59.7 bits (143), Expect = 3e-08
Identities = 42/116 (36%), Positives = 66/116 (56%), Gaps = 8/116 (6%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP 69
D GE+ + ++ +DE + ED E + E G+D+ ++ G E DEG++ GDE E+E
Sbjct: 570 DEGEDEGDEGEDEGEDEGDEGED---EGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEG 626
Query: 70 EENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEE 125
+E E+EGD G++E +E E E EG+D+ G+DE E + E D E+E
Sbjct: 627 DEG--EDEGDEGEDEGDEGE--DEGDEGEDEGDEGEDE-GDEGDEGEDEGDEGEDE 677
Score = 59.3 bits (142), Expect = 3e-08
Identities = 40/121 (33%), Positives = 67/121 (55%), Gaps = 8/121 (6%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIE-QDEGQEAGDEVEEE 68
D G+E ++ DE E + + ++ E + E G+D+ ++ G E +DEG++ GDE E+E
Sbjct: 545 DEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDE 604
Query: 69 PEENVDE-----EEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKE 123
E+ DE +EGD G++E +E E E EG+D+ G+DE E+ + D +
Sbjct: 605 GEDEGDEGDEGEDEGDEGEDEGDEGED--EGDEGEDEGDEGEDEGDEGEDEGDEGEDEGD 662
Query: 124 E 124
E
Sbjct: 663 E 663
Score = 59.3 bits (142), Expect = 3e-08
Identities = 53/149 (35%), Positives = 72/149 (47%), Gaps = 11/149 (7%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDE---RGIEQDEGQEAGDEVE 66
D G+E ++ DE E DE + ED E + E G+D+ DE G E DEG++ GDE E
Sbjct: 617 DEGDEGEDEGDEGE-DEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGEDEGDEGE 675
Query: 67 EEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQ 126
+E +E E+EGD GDE E E E EG+D+ G+DE +E + + + Q
Sbjct: 676 DEGDEG--EDEGDEGDEGDEGDEGDEGEDEGEDE---GEDE--GDEGTKDKEGNANKVVQ 728
Query: 127 VEVIKETQKQKELEVFVGGLDKEATEHDL 155
QK V ATEH L
Sbjct: 729 NPFDYYNWLQKSTLSIVHNKSIHATEHML 757
Score = 59.3 bits (142), Expect = 3e-08
Identities = 36/90 (40%), Positives = 58/90 (64%), Gaps = 5/90 (5%)
Query: 20 DEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEE--E 77
D+ +KDE+ + E+ + D EE G+D+ DE+ E++EG++ GD+ E+E E+ ++E E
Sbjct: 398 DDRDKDEY-ELENEEYNRDEEEDEGEDEEDEKD-EKEEGEDEGDDGEDEGEDEGEDEGDE 455
Query: 78 GDTGDE-EVEEVEYVYEEVEGDDDDVAGDD 106
GD GDE E E + EE EG+D+ GD+
Sbjct: 456 GDEGDEGEDEGEDEDDEEDEGEDEGDEGDE 485
Score = 58.2 bits (139), Expect = 8e-08
Identities = 40/120 (33%), Positives = 67/120 (55%), Gaps = 3/120 (2%)
Query: 10 DNGEEVKESIDEYEK-DEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEE 68
D G+E E DE ++ DE + ++ E++ E G+D+ DE E DEG++ G++ +E
Sbjct: 508 DEGDEGDEGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDE 567
Query: 69 PEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVE 128
E+ E+EGD G++E E+ E E EG+D+ G+DE E + + D +E + E
Sbjct: 568 GEDE-GEDEGDEGEDEGED-EGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDE 625
Score = 56.2 bits (134), Expect = 3e-07
Identities = 44/119 (36%), Positives = 62/119 (51%), Gaps = 7/119 (5%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP 69
+ GE+ + ++ +DE D D E D E G+D+ DE +DEG E GDE E+E
Sbjct: 432 EEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGDE-GDEGEDEG 490
Query: 70 EENVD-EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGD--DEHASEENVHEHLADVKEEE 125
+E + E+EGD GDE +E + E EG D+ GD DE E+ E D E+E
Sbjct: 491 DEGDEGEDEGDEGDEGKDEGD---EGDEGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDE 546
Score = 53.5 bits (127), Expect = 2e-06
Identities = 36/110 (32%), Positives = 61/110 (54%), Gaps = 5/110 (4%)
Query: 15 VKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVD 74
V + +E E + +D D+ + D +EY +++ R E+DEG+ DE +E+ E+
Sbjct: 378 VDKQANEKEYKKIIDKSDDRDDRDKDEYELENEEYNRDEEEDEGE---DEEDEKDEKEEG 434
Query: 75 EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEE 124
E+EGD G++E E+ E E EGD+ D G+DE E++ + D +E
Sbjct: 435 EDEGDDGEDEGED-EGEDEGDEGDEGD-EGEDEGEDEDDEEDEGEDEGDE 482
Score = 42.4 bits (98), Expect = 0.004
Identities = 37/128 (28%), Positives = 62/128 (47%), Gaps = 11/128 (8%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVG-DDDYDERGIEQDEGQEAGDEVEEEPEE 71
++ ES++ + L + +E + ++ + DD D+R DE E E EE
Sbjct: 361 DDESESVNSVALRQVLTVDKQANEKEYKKIIDKSDDRDDRD---------KDEYELENEE 411
Query: 72 -NVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVI 130
N DEEE + DEE E+ E E EGDD + G+DE E + + + ++E + E
Sbjct: 412 YNRDEEEDEGEDEEDEKDEKEEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDD 471
Query: 131 KETQKQKE 138
+E + + E
Sbjct: 472 EEDEGEDE 479
Score = 32.0 bits (71), Expect = 5.9
Identities = 29/119 (24%), Positives = 49/119 (40%), Gaps = 8/119 (6%)
Query: 25 DEHLDFEDNYHEYDPEEY---VGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTG 81
D L Y ++D +E + DD+ D ++G + E E +V + T
Sbjct: 324 DHKLSPVSEYEDFDEDEVELCISDDEVDS-----EDGNLCVLDDESESVNSVALRQVLTV 378
Query: 82 DEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELE 140
D++ E EY + DD D DE+ E + + E E E K+ +++ E E
Sbjct: 379 DKQANEKEYKKIIDKSDDRDDRDKDEYELENEEYNRDEEEDEGEDEEDEKDEKEEGEDE 437
>PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic and
nuclear (Poly(A)-binding protein) (PABP) (ARS consensus
binding protein ACBP-67) (Polyadenylate tail-binding
protein)
Length = 576
Score = 65.5 bits (158), Expect = 5e-10
Identities = 58/278 (20%), Positives = 121/278 (42%), Gaps = 15/278 (5%)
Query: 116 EHLADVKEEEQVEVIKETQ--KQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVN 173
E+L +++Q E+Q + ++VG L+ +E L +FS +G ++ +R+ +
Sbjct: 12 ENLNIQDDQKQAATGSESQSVENSSASLYVGDLEPSVSEAHLYDIFSPIGSVSSIRVCRD 71
Query: 174 PQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITIT------TCQDSDTLYLDN 227
TK + G+A + F E ++A+ +L I G+ C I + + S +++ N
Sbjct: 72 AITKTSLGYAYVNFNDHEAGRKAIEQLNYTPIKGRLCRIMWSQRDPSLRKKGSGNIFIKN 131
Query: 228 ICKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQK 287
+ +AL + +G + D+N G + G F+ F +K+A L
Sbjct: 132 LHPDIDNKALYDTFSVFG--DILSSKIATDEN--GKSKGFGFVHFEEEGAAKEAIDALNG 187
Query: 288 TDVVFGVDKPAEVSFANSFIDLG-DDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEK 346
++ G + + D ++ A +++ + ++ + L K+G +
Sbjct: 188 M-LLNGQEIYVAPHLSRKERDSQLEETKAHYTNLYVKNINSETTDEQFQELFAKFGPIVS 246
Query: 347 VELAKNMPGARRKNYGFVTFGTHAAAVECAESITSAGL 384
L K+ G + K +GFV + H AV+ E++ + L
Sbjct: 247 ASLEKDADG-KLKGFGFVNYEKHEDAVKAVEALNDSEL 283
Score = 52.0 bits (123), Expect = 6e-06
Identities = 49/222 (22%), Positives = 101/222 (45%), Gaps = 9/222 (4%)
Query: 2 ALNIYLIADNGEE--VKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQ 59
ALN L+ NG+E V + E+D L+ ++ + + + DE+ Q+
Sbjct: 184 ALNGMLL--NGQEIYVAPHLSRKERDSQLEETKAHYTNLYVKNINSETTDEQF--QELFA 239
Query: 60 EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEG-DDDDVAGDDEHASE-ENVHEH 117
+ G V E++ D + G E+ E + VE +D ++ G+ + + +E
Sbjct: 240 KFGPIVSASLEKDADGKLKGFGFVNYEKHEDAVKAVEALNDSELNGEKLYVGRAQKKNER 299
Query: 118 LADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTK 177
+ +K++ + +++ K + + +FV LD + L + F+ G IT ++ + +
Sbjct: 300 MHVLKKQYEAYRLEKMAKYQGVNLFVKNLDDSVDDEKLEEEFAPYGTITSAKV-MRTENG 358
Query: 178 RNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQD 219
++KGF + F T E +A+ E ++ GK + I +D
Sbjct: 359 KSKGFGFVCFSTPEEATKAITEKNQQIVAGKPLYVAIAQRKD 400
>PAB5_PONPY (P60050) Polyadenylate-binding protein 5
(Poly(A)-binding protein 5) (PABP 5)
Length = 382
Score = 65.5 bits (158), Expect = 5e-10
Identities = 74/304 (24%), Positives = 126/304 (41%), Gaps = 52/304 (17%)
Query: 134 QKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHV 193
+K + ++VG LD + TE L K F G + R+ +P T+ G+ + F
Sbjct: 13 KKYLKAALYVGDLDPDVTEDMLYKKFRPAGPLRFTRICRDPVTRSPLGYGYVNFRFPADA 72
Query: 194 KRALAELKNPVINGKQCGITITTCQDS------DTLYLDNICKSWKKEALKEKLKHYGVE 247
+ AL + +INGK + + D +++ N+ KS AL +G
Sbjct: 73 EWALNTMNFDLINGKPFRLMWSQPDDRLRKSGVGNIFIKNLDKSIDNRALFYLFSAFG-- 130
Query: 248 SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAY-----KRLQKTDVVFG-----VDKP 297
+ ++ DDN G A++ F S + + A RL V G ++
Sbjct: 131 NILSCKVVCDDNGSK---GYAYVHFDSLAAANRAIWHMNGVRLNNRQVYVGRFKFPEERA 187
Query: 298 AEV------SFANSFI-DLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELA 350
AEV +F N F+ ++GDDI D++ ++ L +YG E V++
Sbjct: 188 AEVRTRDRATFTNVFVKNIGDDI----------------DDEKLKELFCEYGPTESVKVI 231
Query: 351 KNMPGARRKNYGFVTFGTHAAAVECAESITSAGL-------GEGNKKAKVRARLSRPLPK 403
++ G + K +GFV + TH AA + + + G KK + A L R +
Sbjct: 232 RDASG-KSKGFGFVRYETHEAAQKAVLDLHGKSIDGKVLYVGRAQKKIERLAELRRRFER 290
Query: 404 VRGK 407
+R K
Sbjct: 291 LRLK 294
Score = 44.3 bits (103), Expect = 0.001
Identities = 23/99 (23%), Positives = 50/99 (50%), Gaps = 2/99 (2%)
Query: 116 EHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ 175
E LA+++ + +KE + + +++ LD+ + L++ FS G I+ R V +
Sbjct: 279 ERLAELRRRFERLRLKEKSRPPGVPIYIKNLDETINDEKLKEEFSSFGSIS--RAKVMME 336
Query: 176 TKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITI 214
+ KGF ++ F + E +A+ E+ ++ K +T+
Sbjct: 337 VGQGKGFGVVCFSSFEEATKAVDEMNGRIVGSKPLHVTL 375
Score = 41.6 bits (96), Expect = 0.007
Identities = 33/148 (22%), Positives = 69/148 (46%), Gaps = 12/148 (8%)
Query: 124 EEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFA 183
EE+ ++ + VFV + + + L+++F + G V++ + + ++KGF
Sbjct: 184 EERAAEVRTRDRATFTNVFVKNIGDDIDDEKLKELFCEYGPTESVKV-IRDASGKSKGFG 242
Query: 184 LLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKH 243
+R+ET E ++A+ +L I+GK + L + + +++ LKEK +
Sbjct: 243 FVRYETHEAAQKAVLDLHGKSIDGK---VLYVGRAQKKIERLAELRRRFERLRLKEKSRP 299
Query: 244 YGV--------ESFKDLTLLEDDNNEGT 263
GV E+ D L E+ ++ G+
Sbjct: 300 PGVPIYIKNLDETINDEKLKEEFSSFGS 327
>PAB5_PANTR (P60049) Polyadenylate-binding protein 5
(Poly(A)-binding protein 5) (PABP 5)
Length = 382
Score = 65.5 bits (158), Expect = 5e-10
Identities = 74/304 (24%), Positives = 126/304 (41%), Gaps = 52/304 (17%)
Query: 134 QKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHV 193
+K + ++VG LD + TE L K F G + R+ +P T+ G+ + F
Sbjct: 13 KKYLKAALYVGDLDPDVTEDMLYKKFRPAGPLRFTRICRDPVTRSPLGYGYVNFRFPADA 72
Query: 194 KRALAELKNPVINGKQCGITITTCQDS------DTLYLDNICKSWKKEALKEKLKHYGVE 247
+ AL + +INGK + + D +++ N+ KS AL +G
Sbjct: 73 EWALNTMNFDLINGKPFRLMWSQPDDRLRKSGVGNIFIKNLDKSIDNRALFYLFSAFG-- 130
Query: 248 SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAY-----KRLQKTDVVFG-----VDKP 297
+ ++ DDN G A++ F S + + A RL V G ++
Sbjct: 131 NILSCKVVCDDNGSK---GYAYVHFDSLAAANRAIWHMNGVRLNNRQVYVGRFKFPEERA 187
Query: 298 AEV------SFANSFI-DLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELA 350
AEV +F N F+ ++GDDI D++ ++ L +YG E V++
Sbjct: 188 AEVRTRDRATFTNVFVKNIGDDI----------------DDEKLKELFCEYGPTESVKVI 231
Query: 351 KNMPGARRKNYGFVTFGTHAAAVECAESITSAGL-------GEGNKKAKVRARLSRPLPK 403
++ G + K +GFV + TH AA + + + G KK + A L R +
Sbjct: 232 RDASG-KSKGFGFVRYETHEAAQKAVLDLHGKSIDGKVLYVGRAQKKIERLAELRRRFER 290
Query: 404 VRGK 407
+R K
Sbjct: 291 LRLK 294
Score = 44.3 bits (103), Expect = 0.001
Identities = 23/99 (23%), Positives = 50/99 (50%), Gaps = 2/99 (2%)
Query: 116 EHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ 175
E LA+++ + +KE + + +++ LD+ + L++ FS G I+ R V +
Sbjct: 279 ERLAELRRRFERLRLKEKSRPPGVPIYIKNLDETINDEKLKEEFSSFGSIS--RAKVMME 336
Query: 176 TKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITI 214
+ KGF ++ F + E +A+ E+ ++ K +T+
Sbjct: 337 VGQGKGFGVVCFSSFEEATKAVDEMNGRIVGSKPLHVTL 375
Score = 41.6 bits (96), Expect = 0.007
Identities = 33/148 (22%), Positives = 69/148 (46%), Gaps = 12/148 (8%)
Query: 124 EEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFA 183
EE+ ++ + VFV + + + L+++F + G V++ + + ++KGF
Sbjct: 184 EERAAEVRTRDRATFTNVFVKNIGDDIDDEKLKELFCEYGPTESVKV-IRDASGKSKGFG 242
Query: 184 LLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKH 243
+R+ET E ++A+ +L I+GK + L + + +++ LKEK +
Sbjct: 243 FVRYETHEAAQKAVLDLHGKSIDGK---VLYVGRAQKKIERLAELRRRFERLRLKEKSRP 299
Query: 244 YGV--------ESFKDLTLLEDDNNEGT 263
GV E+ D L E+ ++ G+
Sbjct: 300 PGVPIYIKNLDETINDEKLKEEFSSFGS 327
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 95,416,382
Number of Sequences: 164201
Number of extensions: 4810862
Number of successful extensions: 37216
Number of sequences better than 10.0: 1317
Number of HSP's better than 10.0 without gapping: 590
Number of HSP's successfully gapped in prelim test: 785
Number of HSP's that attempted gapping in prelim test: 20336
Number of HSP's gapped (non-prelim): 7145
length of query: 726
length of database: 59,974,054
effective HSP length: 118
effective length of query: 608
effective length of database: 40,598,336
effective search space: 24683788288
effective search space used: 24683788288
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149129.7


