

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149127.7 - phase: 0
(360 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
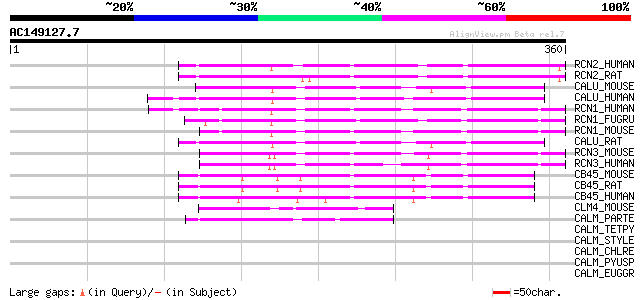
Sequences producing significant alignments: (bits) Value
RCN2_HUMAN (Q14257) Reticulocalbin 2 precursor (Calcium-binding ... 69 2e-11
RCN2_RAT (Q62703) Reticulocalbin 2 precursor (Calcium-binding pr... 68 3e-11
CALU_MOUSE (O35887) Calumenin precursor (Crocalbin) 66 2e-10
CALU_HUMAN (O43852) Calumenin precursor (Crocalbin) (IEF SSP 9302) 64 5e-10
RCN1_HUMAN (Q15293) Reticulocalbin 1 precursor 60 7e-09
RCN1_FUGRU (O93434) Reticulocalbin 1 precursor 60 9e-09
RCN1_MOUSE (Q05186) Reticulocalbin 1 precursor 57 7e-08
CALU_RAT (O35783) Calumenin precursor (Crocalbin) (CBP-50) 57 9e-08
RCN3_MOUSE (Q8BH97) Reticulocalbin 3 precursor 54 5e-07
RCN3_HUMAN (Q96D15) Reticulocalbin 3 precursor (EF-hand calcium ... 53 1e-06
CB45_MOUSE (Q61112) 45 kDa calcium-binding protein precursor (Ca... 50 7e-06
CB45_RAT (Q91ZS3) 45 kDa calcium-binding protein precursor (Cab4... 47 7e-05
CB45_HUMAN (Q9BRK5) 45 kDa calcium-binding protein precursor (Ca... 47 7e-05
CLM4_MOUSE (Q9JM83) Calmodulin 4 (Calcium-binding protein Dd112) 45 3e-04
CALM_PARTE (P07463) Calmodulin (CaM) 44 5e-04
CALM_TETPY (P02598) Calmodulin (CaM) 43 0.001
CALM_STYLE (P27166) Calmodulin (CaM) 43 0.001
CALM_CHLRE (P04352) Calmodulin (CaM) 43 0.001
CALM_PYUSP (P11121) Calmodulin (CaM) 42 0.002
CALM_EUGGR (P11118) Calmodulin (CaM) 42 0.002
>RCN2_HUMAN (Q14257) Reticulocalbin 2 precursor (Calcium-binding
protein ERC-55) (E6-binding protein) (E6BP)
Length = 317
Score = 68.6 bits (166), Expect = 2e-11
Identities = 68/270 (25%), Positives = 116/270 (42%), Gaps = 36/270 (13%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREY- 168
RL + +D D DGF+ +EL SW+ + + DKN D +++ EY
Sbjct: 65 RLQAIIKKIDLD-SDGFLTESELSSWIQMSFKHYAMQEAKQQFVEYDKNSDDTVTWDEYN 123
Query: 169 ---------------LPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDF 213
L D E+ K ++ ++F+ A+ D L+ E F
Sbjct: 124 IQMYDRVIDFDENTALDDAEEESFRKLHLKD------KKRFEKANQDSGPGLSLEEFIAF 177
Query: 214 LHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPT 273
HPE+ M ++++ + D DG ++ +F + + D E I
Sbjct: 178 EHPEEVDY--MTEFVIQEALEEHDKNGDGFVSLEEFLGDYRWDPTANEDPEW-----ILV 230
Query: 274 AKDKFA-ELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDE 332
KD+F + D + D L P+EL +P+V P A+ +L++E D N D+KL+ +E
Sbjct: 231 EKDRFVNDYDKDNDGRLDPQEL---LPWVVPNNQGIAQEEALHLIDEMDLNGDKKLSEEE 287
Query: 333 MLDHEFAFFNTVHADGHVEIDDD--DHDEL 360
+L++ F + D ++ DD HDEL
Sbjct: 288 ILENPDLFLTSEATDYGRQLHDDYFYHDEL 317
>RCN2_RAT (Q62703) Reticulocalbin 2 precursor (Calcium-binding
protein ERC-55) (Taipoxin-associated calcium-binding
protein-49) (TCBP-49)
Length = 318
Score = 68.2 bits (165), Expect = 3e-11
Identities = 69/264 (26%), Positives = 118/264 (44%), Gaps = 24/264 (9%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYL 169
RL + +D D DGF+ NEL W+ + + DKN D +++ EY
Sbjct: 66 RLQSIIKKIDSD-SDGFLTENELSQWIQMSFKHYAMQEAKQQFVEYDKNSDGTVTWDEYN 124
Query: 170 PDLSEKDIE-KKNMAHGEAG-------WLMEK--FDVADYDHNGLLNFTELRDFLHPEDS 219
+ ++ I+ +N A + L +K F+ A+ D LN E F HPE+
Sbjct: 125 VQMYDRVIDFDENTALDDTEEESFRQLHLKDKKRFEKANQDSGPGLNLEEFIAFEHPEEV 184
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFA 279
M ++++ + D DG ++ +F + + D E I KD+F
Sbjct: 185 DY--MTEFVIQEALEEHDKNGDGFVSLEEFLGDYRRDPTANEDPEW-----ILVEKDRFV 237
Query: 280 -ELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEF 338
+ D + D L P+EL + +V P A+ +L++E D N D+KL+ +E+L+++
Sbjct: 238 NDYDKDSDGRLDPQEL---LSWVVPNNQGIAQEEALHLIDEMDLNSDKKLSEEEILENQD 294
Query: 339 AFFNTVHADGHVEIDDD--DHDEL 360
F + D ++ DD HDEL
Sbjct: 295 LFLTSEATDYGRQLHDDYFYHDEL 318
>CALU_MOUSE (O35887) Calumenin precursor (Crocalbin)
Length = 315
Score = 65.9 bits (159), Expect = 2e-10
Identities = 61/242 (25%), Positives = 105/242 (43%), Gaps = 34/242 (14%)
Query: 121 DPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYL----------P 170
D KDGFV +EL+ W+ + + + + D N D +S+ EY P
Sbjct: 82 DDKDGFVTVDELKGWIKFAQKRWIHEDVERQWKGHDLNEDGLVSWEEYKNATYGYVLDDP 141
Query: 171 DLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
D + K+ M E +F +AD D + + E FLHPE+ + + +V
Sbjct: 142 DPDDGFNYKQMMVRDE-----RRFKMADKDGDLIATKEEFTAFLHPEEYDYMKDI--VVQ 194
Query: 231 DKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIP----TAKDKFAEL-DVNK 285
+ D DG I+ ++ ++Y +G D P T +++F E D N+
Sbjct: 195 ETMEDIDKNADGFIDLEEYIGDMY---------SHDGNADEPEWVKTEREQFVEFRDKNR 245
Query: 286 DQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVH 345
D + EE ++ P + +A+ +L+ E+D N+D KLT +E++D F +
Sbjct: 246 DGKMDKEETKD---WILPSDYDHAEAEARHLVYESDQNKDGKLTKEEIVDKYDLFVGSQA 302
Query: 346 AD 347
D
Sbjct: 303 TD 304
Score = 31.2 bits (69), Expect = 4.3
Identities = 47/217 (21%), Positives = 83/217 (37%), Gaps = 48/217 (22%)
Query: 163 LSFREYLPDLS-------EKDIEKKNMAHGEAGWLMEKFDVA---DYDHNGLLNFTELRD 212
+ R++L LS K EKK+ H E + + A DYDH+ L E +
Sbjct: 1 MDLRQFLMCLSLCTAFALSKPTEKKDRVHHEPQLSDKVHNDAQNFDYDHDAFLGAEEAKS 60
Query: 213 F--LHPEDSQNK-------------------EMLKWM-----------VNDKFNMDDYEH 240
F L PE+S+ + E+ W+ V ++ D
Sbjct: 61 FDQLTPEESKERLGKIVSKIDDDKDGFVTVDELKGWIKFAQKRWIHEDVERQWKGHDLNE 120
Query: 241 DGKINFNQFEDNV--YVTYESYVDFETNGEGDIPTAKDKFAELDVNKDQFLSPEELFPII 298
DG +++ ++++ YV + D N + + + +F D + D + EE
Sbjct: 121 DGLVSWEEYKNATYGYVLDDPDPDDGFNYKQMMVRDERRFKMADKDGDLIATKEE---FT 177
Query: 299 PYVYPGELAYAK-YYTSYLMNEADDNEDRKLTLDEML 334
+++P E Y K M + D N D + L+E +
Sbjct: 178 AFLHPEEYDYMKDIVVQETMEDIDKNADGFIDLEEYI 214
>CALU_HUMAN (O43852) Calumenin precursor (Crocalbin) (IEF SSP 9302)
Length = 315
Score = 64.3 bits (155), Expect = 5e-10
Identities = 67/269 (24%), Positives = 119/269 (43%), Gaps = 30/269 (11%)
Query: 90 SEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQ 149
+E +T++ LT + RL + +D D KDGFV +EL+ W+ + +
Sbjct: 55 AEEAKTFDQLTPEESKE---RLGKIVSKIDGD-KDGFVTVDELKDWIKFAQKRWIYEDVE 110
Query: 150 VELESKDKNGDLALSFREYL----------PDLSEKDIEKKNMAHGEAGWLMEKFDVADY 199
+ + D N D +S+ EY PD + K+ M E +F +AD
Sbjct: 111 RQWKGHDLNEDGLVSWEEYKNATYGYVLDDPDPDDGFNYKQMMVRDE-----RRFKMADK 165
Query: 200 DHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYES 259
D + + E FLHPE+ + + +V + D DG I+ ++ ++Y
Sbjct: 166 DGDLIATKEEFTAFLHPEEYDYMKDI--VVQETMEDIDKNADGFIDLEEYIGDMYSH--- 220
Query: 260 YVDFETNGEGDIPTAKDKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMN 318
D T+ + T +++F E D N+D + EE ++ P + +A+ +L+
Sbjct: 221 --DGNTDEPEWVKTEREQFVEFRDKNRDGKMDKEETKD---WILPSDYDHAEAEARHLVY 275
Query: 319 EADDNEDRKLTLDEMLDHEFAFFNTVHAD 347
E+D N+D KLT +E++D F + D
Sbjct: 276 ESDQNKDGKLTKEEIVDKYDLFVGSQATD 304
Score = 31.2 bits (69), Expect = 4.3
Identities = 47/217 (21%), Positives = 83/217 (37%), Gaps = 48/217 (22%)
Query: 163 LSFREYLPDLS-------EKDIEKKNMAHGEAGWLMEKFDVA---DYDHNGLLNFTELRD 212
+ R++L LS K EKK+ H E + + A DYDH+ L E +
Sbjct: 1 MDLRQFLMCLSLCTAFALSKPTEKKDRVHHEPQLSDKVHNDAQSFDYDHDAFLGAEEAKT 60
Query: 213 F--LHPEDSQNK-------------------EMLKWM-----------VNDKFNMDDYEH 240
F L PE+S+ + E+ W+ V ++ D
Sbjct: 61 FDQLTPEESKERLGKIVSKIDGDKDGFVTVDELKDWIKFAQKRWIYEDVERQWKGHDLNE 120
Query: 241 DGKINFNQFEDNV--YVTYESYVDFETNGEGDIPTAKDKFAELDVNKDQFLSPEELFPII 298
DG +++ ++++ YV + D N + + + +F D + D + EE
Sbjct: 121 DGLVSWEEYKNATYGYVLDDPDPDDGFNYKQMMVRDERRFKMADKDGDLIATKEE---FT 177
Query: 299 PYVYPGELAYAK-YYTSYLMNEADDNEDRKLTLDEML 334
+++P E Y K M + D N D + L+E +
Sbjct: 178 AFLHPEEYDYMKDIVVQETMEDIDKNADGFIDLEEYI 214
>RCN1_HUMAN (Q15293) Reticulocalbin 1 precursor
Length = 331
Score = 60.5 bits (145), Expect = 7e-09
Identities = 74/288 (25%), Positives = 135/288 (46%), Gaps = 41/288 (14%)
Query: 91 EIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQV 150
E +T++ LT + RL + +D D DGFV EL++W+ +R +R +
Sbjct: 67 EDSKTFDQLTPDESKE---RLGKIVDRIDNDG-DGFVTTEELKTWI-KRVQKRYIFDNVA 121
Query: 151 EL-ESKDKNGDLALSFREY--------------LPDLSEKDIEKKNMAHGEAGWLMEKFD 195
++ + D++ D +S+ EY D S+ KK + E +F
Sbjct: 122 KVWKDYDRDKDDKISWEEYKQATYGYYLGNPAEFHDSSDHHTFKKMLPRDE-----RRFK 176
Query: 196 VADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYV 255
AD + + E FLHPE+ ++ M + +V + D DG ++ +++ +++
Sbjct: 177 AADLNGDLTATREEFTAFLHPEEFEH--MKEIVVLETLEDIDKNGDGFVDQDEYIADMF- 233
Query: 256 TYESYVDFETNG-EGD-IPTAKDKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYY 312
E NG E D + + +++F E D+NKD L +E I ++ P + +A+
Sbjct: 234 ------SHEENGPEPDWVLSEREQFNEFRDLNKDGKLDKDE---IRHWILPQDYDHAQAE 284
Query: 313 TSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGHVEIDDDDHDEL 360
+L+ E+D N+D KLT +E+L++ F + A + E +HDEL
Sbjct: 285 ARHLVYESDKNKDEKLTKEEILENWNMFVGS-QATNYGEDLTKNHDEL 331
>RCN1_FUGRU (O93434) Reticulocalbin 1 precursor
Length = 322
Score = 60.1 bits (144), Expect = 9e-09
Identities = 63/265 (23%), Positives = 128/265 (47%), Gaps = 35/265 (13%)
Query: 114 LFPLLDRDPKDG--FVGFNELESWVTQRALERLDYATQVELESK-DKNGDLALSFREY-- 168
L ++DR DG ++ +EL++W+ +R +R Y V++ + D N D +S+ EY
Sbjct: 75 LSKIVDRIDGDGNSYITTDELKAWI-KRVQKRYVYENVVKVWADYDLNKDNKISWEEYKQ 133
Query: 169 ------------LPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHP 216
+ +++ KK + E +F AD D + N E FLHP
Sbjct: 134 ATYGYYLSNPEEFDETTDQFSFKKMLPRDE-----RRFKRADLDGDSAANREEFTSFLHP 188
Query: 217 EDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKD 276
E+ ++ + + +V + D DG ++ +++ +++ + + E + T ++
Sbjct: 189 EEFEHMKDI--VVLETLEDIDKNSDGHVDEDEYIADMFAHEDRGPEPEW-----VKTERE 241
Query: 277 KFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLD 335
+F++ D+NKD + +E I ++ P + +A+ +L+ E+D ++D+ LT +E+LD
Sbjct: 242 QFSDFRDLNKDGKMDLDE---IRHWIMPQDYDHAQAEARHLVYESDKDKDQMLTKEEILD 298
Query: 336 HEFAFFNTVHADGHVEIDDDDHDEL 360
+ F + A + E +HDEL
Sbjct: 299 NWNMFVGS-QATNYGEDLTRNHDEL 322
>RCN1_MOUSE (Q05186) Reticulocalbin 1 precursor
Length = 325
Score = 57.0 bits (136), Expect = 7e-08
Identities = 65/255 (25%), Positives = 118/255 (45%), Gaps = 37/255 (14%)
Query: 124 DGFVGFNELESWVTQRALERLDYATQVEL-ESKDKNGDLALSFREY-------------- 168
DG V EL+ W+ +R +R Y ++ + D++ D +S+ EY
Sbjct: 90 DGLVTTEELKLWI-KRVQKRYIYDNVAKVWKDYDRDKDEKISWEEYKQATYGYYLGNPAE 148
Query: 169 LPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWM 228
D S+ KK + E +F +D D + E FLHPE+ ++ M + +
Sbjct: 149 FHDSSDHHTFKKMLPRDE-----RRFKASDLDGDLTATREEFTAFLHPEEFEH--MKEIV 201
Query: 229 VNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNG-EGD-IPTAKDKFAEL-DVNK 285
V + D DG ++ +++ +++ E NG E D + + +++F + D+NK
Sbjct: 202 VLETLEDIDKNGDGFVDQDEYIADMF-------SHEDNGPEPDWVLSEREQFNDFRDLNK 254
Query: 286 DQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVH 345
D L +E I ++ P + +A+ +L+ E+D N+D LT +E+LD+ F +
Sbjct: 255 DGKLDKDE---IRHWILPQDYDHAQAEARHLVYESDKNKDEMLTKEEILDNWNMFVGS-Q 310
Query: 346 ADGHVEIDDDDHDEL 360
A + E +HDEL
Sbjct: 311 ATNYGEDLTKNHDEL 325
>CALU_RAT (O35783) Calumenin precursor (Crocalbin) (CBP-50)
Length = 315
Score = 56.6 bits (135), Expect = 9e-08
Identities = 61/253 (24%), Positives = 110/253 (43%), Gaps = 35/253 (13%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYL 169
+L ++ +D D KDGFV EL+S + + + + + + D N D +S+ EY
Sbjct: 72 KLGMIVDKIDTD-KDGFVTEGELKSRIKHAQKKYIYDNVENQWQEFDMNQDGLISWDEYR 130
Query: 170 ----------PDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDS 219
PD + K M E +F +AD D + + E FLHPE+
Sbjct: 131 NVTYGTYLDDPDPDDGFNYKPIMVRDE-----RRFKMADQDGDLIATKEEFTAFLHPEEY 185
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIP----TAK 275
+ + ++ + D DG I+ ++ ++Y +G D P T +
Sbjct: 186 DYMKDI--VLQETMEDIDQNADGFIDLEEYIGDMY---------SHDGNADEPQWVKTER 234
Query: 276 DKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEML 334
++F E D N+D + EE ++ P + +A+ +L+ E+D ++D KLT +E++
Sbjct: 235 EQFVEFRDKNRDGKMDKEETKD---WILPSDYDHAEAEARHLVYESDQDKDGKLTKEEIV 291
Query: 335 DHEFAFFNTVHAD 347
D F + D
Sbjct: 292 DKYDLFVGSQATD 304
Score = 36.2 bits (82), Expect = 0.13
Identities = 36/143 (25%), Positives = 62/143 (43%), Gaps = 27/143 (18%)
Query: 163 LSFREYLPDLS-------EKDIEKKNMAHGEAGWLMEKFDVA---DYDHNGLLNFTELRD 212
+ R++L LS K EKK+ H E + + A DYDH+ L E +
Sbjct: 1 MDLRQFLMCLSLCTAFALSKPTEKKDRVHHEPQLSDKVHNDAQNFDYDHDAFLGAEEAKS 60
Query: 213 F--LHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGD 270
F L PE+S+ K M+ DK D + DG + + + + + Y+
Sbjct: 61 FGQLTPEESKEK---LGMIVDKI---DTDKDGFVTEGELKSRIKHAQKKYI--------- 105
Query: 271 IPTAKDKFAELDVNKDQFLSPEE 293
++++ E D+N+D +S +E
Sbjct: 106 YDNVENQWQEFDMNQDGLISWDE 128
>RCN3_MOUSE (Q8BH97) Reticulocalbin 3 precursor
Length = 328
Score = 54.3 bits (129), Expect = 5e-07
Identities = 62/254 (24%), Positives = 107/254 (41%), Gaps = 36/254 (14%)
Query: 124 DGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFRE--------YLP----- 170
DG+V EL +W+ + + + D + D + + E Y P
Sbjct: 94 DGWVSLAELRAWIAHTQQRHIRDSVSAAWHTYDTDRDGRVGWEELRNATYGHYEPGEEFH 153
Query: 171 DLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
D+ + + KK +A E +F VAD D + + EL FLHPE+ + M +V
Sbjct: 154 DVEDAETYKKMLARDE-----RRFRVADQDGDSMATREELTAFLHPEEFPH--MRDIVVA 206
Query: 231 DKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGD---IPTAKDKFAEL-DVNKD 286
+ D DG + ++ ++Y E GE + + T + +F E D+NKD
Sbjct: 207 ETLEDLDKNKDGYVQVEEYIADLY--------SEEPGEEEPAWVQTERQQFREFRDLNKD 258
Query: 287 QFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHA 346
L E + +V P ++L++E+D ++D +L+ E+L + F + A
Sbjct: 259 GRLDGSE---VGYWVLPPSQDQPLVEANHLLHESDTDKDGRLSKAEILSNWNMFVGS-QA 314
Query: 347 DGHVEIDDDDHDEL 360
+ E HDEL
Sbjct: 315 TNYGEDLTRHHDEL 328
>RCN3_HUMAN (Q96D15) Reticulocalbin 3 precursor (EF-hand calcium
binding protein RLP49) (UNQ239/PRO272)
Length = 328
Score = 52.8 bits (125), Expect = 1e-06
Identities = 62/257 (24%), Positives = 107/257 (41%), Gaps = 42/257 (16%)
Query: 124 DGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFRE--------YLP----- 170
DG+V EL +W+ + + ++ D + D + + E Y P
Sbjct: 94 DGWVSLAELRAWIAHTQQRHIRDSVSAAWDTYDTDRDGRVGWEELRNATYGHYAPGEEFH 153
Query: 171 DLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
D+ + + KK +A E +F VAD D + + EL FLHPE+ + M ++
Sbjct: 154 DVEDAETYKKMLARDE-----RRFRVADQDGDSMATREELTAFLHPEEFPH--MRDIVIA 206
Query: 231 DKFNMDDYEHDGKINFNQFEDNVYVTYESYV-DFETNGEGD-----IPTAKDKFAEL-DV 283
+ D DG YV E Y+ D + G+ + T + +F + D+
Sbjct: 207 ETLEDLDRNKDG-----------YVQVEEYIADLYSAEPGEEEPAWVQTERQQFRDFRDL 255
Query: 284 NKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNT 343
NKD L E + +V P ++L++E+D ++D +L+ E+L + F +
Sbjct: 256 NKDGHLDGSE---VGHWVLPPAQDQPLVEANHLLHESDTDKDGRLSKAEILGNWNMFVGS 312
Query: 344 VHADGHVEIDDDDHDEL 360
A + E HDEL
Sbjct: 313 -QATNYGEDLTRHHDEL 328
>CB45_MOUSE (Q61112) 45 kDa calcium-binding protein precursor
(Cab45) (Stromal cell-derived factor 4) (SDF-4)
Length = 361
Score = 50.4 bits (119), Expect = 7e-06
Identities = 65/254 (25%), Positives = 112/254 (43%), Gaps = 32/254 (12%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQ---VELESKDKNGDLALSFR 166
+L+++F +D + D + E++ W+ ++ E A + + + D +GD +S+
Sbjct: 101 KLMVIFSKVDVNT-DRRISAKEMQHWIMEKTAEHFQEAVKENKLHFRAVDPDGDGHVSWD 159
Query: 167 EYLPDL------SEKDIEKKNMAHGEA----------GWLMEKFDVADYDHNGLL-NFTE 209
EY +E++I + H E G L +++ AD LL E
Sbjct: 160 EYKVKFLASKGHNEREIAEAIKNHEELKVDEETQEVLGNLRDRWYQADNPPADLLLTEDE 219
Query: 210 LRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESY--VDFETNG 267
FLHPE S+ MLK+MV + F D + D +++ +F T E+ D + N
Sbjct: 220 FLSFLHPEHSRG--MLKFMVKEIFRDLDQDGDKQLSLPEFISLPVGTVENQQGQDIDDNW 277
Query: 268 EGDIPTAKDKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDR 326
D K +F EL D N D ++ EEL Y+ P A ++ AD+N++
Sbjct: 278 VKD---RKKEFEELIDSNHDGIVTMEEL---ENYMDPMNEYNALNEAKQMIAIADENQNH 331
Query: 327 KLTLDEMLDHEFAF 340
L +E+L + F
Sbjct: 332 HLEPEEILKYSEFF 345
>CB45_RAT (Q91ZS3) 45 kDa calcium-binding protein precursor (Cab45)
(Stromal cell-derived factor 4) (SDF-4)
Length = 361
Score = 47.0 bits (110), Expect = 7e-05
Identities = 64/254 (25%), Positives = 110/254 (43%), Gaps = 32/254 (12%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQ---VELESKDKNGDLALSFR 166
+L+++F +D + D + E++ W+ ++ E A + + + D +GD +S+
Sbjct: 101 KLMVIFSKVDVNT-DRRISAKEMQHWIMEKTAEHFQEAVKENKLHFRAVDPDGDGHVSWD 159
Query: 167 EYLPDL------SEKDIEKKNMAHGEA----------GWLMEKFDVADYDHNGLL-NFTE 209
EY +E++I H E G L +++ AD LL E
Sbjct: 160 EYKVKFLASKGHNEREIADAIKNHEELKVDEETQEVLGNLRDRWYQADNPPADLLLTEDE 219
Query: 210 LRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESY--VDFETNG 267
FLHPE S+ MLK+MV + D + D +++ +F T E+ D + N
Sbjct: 220 FLSFLHPEHSRG--MLKFMVKEIVRDLDQDGDKQLSLPEFISLPVGTVENQQGQDIDDNW 277
Query: 268 EGDIPTAKDKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDR 326
D K +F EL D N D ++ EEL Y+ P A ++ AD+N++
Sbjct: 278 VKD---RKKEFEELIDSNHDGIVTMEEL---ENYMDPMNEYNALNEAKQMIAIADENQNH 331
Query: 327 KLTLDEMLDHEFAF 340
L +E+L + F
Sbjct: 332 HLEPEEILKYSEFF 345
>CB45_HUMAN (Q9BRK5) 45 kDa calcium-binding protein precursor
(Cab45) (Stromal cell-derived factor 4) (SDF-4)
Length = 362
Score = 47.0 bits (110), Expect = 7e-05
Identities = 62/254 (24%), Positives = 110/254 (42%), Gaps = 32/254 (12%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYA---TQVELESKDKNGDLALSFR 166
+L+++F +D + D + E++ W+ ++ E A ++ + D +GD +S+
Sbjct: 102 KLMVIFSKVDVNT-DRKISAKEMQRWIMEKTAEHFQEAMEESKTHFRAVDPDGDGHVSWD 160
Query: 167 EY-LPDLSEKDIEKKNMAHG-----------EAGWLMEKFDVADYDHNG-----LLNFTE 209
EY + L+ K +K +A E ++E Y + LL E
Sbjct: 161 EYKVKFLASKGHSEKEVADAIRLNEELKVDEETQEVLENLKDRWYQADSPPADLLLTEEE 220
Query: 210 LRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESY--VDFETNG 267
FLHPE S+ ML++MV + D + D +++ +F T E+ D + N
Sbjct: 221 FLSFLHPEHSRG--MLRFMVKEIVRDLDQDGDKQLSVPEFISLPVGTVENQQGQDIDDNW 278
Query: 268 EGDIPTAKDKFAEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDR 326
D K +F EL D N D ++ EEL Y+ P A ++ AD+N++
Sbjct: 279 VKD---RKKEFEELIDSNHDGIVTAEEL---ESYMDPMNEYNALNEAKQMIAVADENQNH 332
Query: 327 KLTLDEMLDHEFAF 340
L +E+L + F
Sbjct: 333 HLEPEEVLKYSEFF 346
>CLM4_MOUSE (Q9JM83) Calmodulin 4 (Calcium-binding protein Dd112)
Length = 148
Score = 45.1 bits (105), Expect = 3e-04
Identities = 36/128 (28%), Positives = 59/128 (45%), Gaps = 11/128 (8%)
Query: 123 KDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEKDIEKKNM 182
KDG + EL + Q + + + D +GD +SF E+L IEK
Sbjct: 24 KDGHISVEELGDVMKQLGKNLPEKDLKALISKLDTDGDGKISFEEFL-----TAIEKYKK 78
Query: 183 AHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHP-EDSQNKEMLKWMVNDKFNMDDYEHD 241
H AG L F+V D + +G + EL++ L +S ++E L+ D + D + D
Sbjct: 79 GH-RAGELRAVFNVLDQNGDGYITVDELKESLSKLGESLSQEELE----DMIRVADVDQD 133
Query: 242 GKINFNQF 249
GK+ + +F
Sbjct: 134 GKVKYEEF 141
>CALM_PARTE (P07463) Calmodulin (CaM)
Length = 148
Score = 44.3 bits (103), Expect = 5e-04
Identities = 36/135 (26%), Positives = 62/135 (45%), Gaps = 9/135 (6%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 16 FALFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLSLMAR 74
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K E+ + L+E F V D D NGL++ ELR H + +++ V++
Sbjct: 75 KMKEQDSEEE-----LIEAFKVFDRDGNGLISAAELR---HVMTNLGEKLTDDEVDEMIR 126
Query: 235 MDDYEHDGKINFNQF 249
D + DG IN+ +F
Sbjct: 127 EADIDGDGHINYEEF 141
>CALM_TETPY (P02598) Calmodulin (CaM)
Length = 148
Score = 43.1 bits (100), Expect = 0.001
Identities = 37/139 (26%), Positives = 63/139 (44%), Gaps = 17/139 (12%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 16 FSLFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLSLMAR 74
Query: 175 K----DIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
K D E++ L+E F V D D NGL++ ELR H + +++ V+
Sbjct: 75 KMKDTDTEEE---------LIEAFKVFDRDGNGLISAAELR---HVMTNLGEKLTDEEVD 122
Query: 231 DKFNMDDYEHDGKINFNQF 249
+ D + DG IN+ +F
Sbjct: 123 EMIREADIDGDGHINYEEF 141
>CALM_STYLE (P27166) Calmodulin (CaM)
Length = 148
Score = 43.1 bits (100), Expect = 0.001
Identities = 37/139 (26%), Positives = 63/139 (44%), Gaps = 17/139 (12%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 16 FSLFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLSLMAR 74
Query: 175 K----DIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
K D E++ L+E F V D D NGL++ ELR H + +++ V+
Sbjct: 75 KMKDTDTEEE---------LVEAFKVFDRDGNGLISAAELR---HVMTNLGEKLTDEEVD 122
Query: 231 DKFNMDDYEHDGKINFNQF 249
+ D + DG IN+ +F
Sbjct: 123 EMIREADVDGDGHINYEEF 141
>CALM_CHLRE (P04352) Calmodulin (CaM)
Length = 162
Score = 42.7 bits (99), Expect = 0.001
Identities = 35/135 (25%), Positives = 62/135 (45%), Gaps = 9/135 (6%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 19 FALFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMISEVDADGNGTIDFPEFLMLMAR 77
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K K H + L E F V D D NG ++ ELR H + +++ + V++
Sbjct: 78 K---MKETDHEDE--LREAFKVFDKDGNGFISAAELR---HVMTNLGEKLSEEEVDEMIR 129
Query: 235 MDDYEHDGKINFNQF 249
D + DG++N+ +F
Sbjct: 130 EADVDGDGQVNYEEF 144
>CALM_PYUSP (P11121) Calmodulin (CaM)
Length = 148
Score = 42.0 bits (97), Expect = 0.002
Identities = 33/135 (24%), Positives = 60/135 (44%), Gaps = 9/135 (6%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +GD + F E+L ++
Sbjct: 16 FSLFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDFPEFLTMMAR 74
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + + E F V D D NG ++ ELR H + +++ V++
Sbjct: 75 KMKDTDSEEE-----IREAFRVFDKDGNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 126
Query: 235 MDDYEHDGKINFNQF 249
D + DG++N+ +F
Sbjct: 127 EADIDGDGQVNYEEF 141
>CALM_EUGGR (P11118) Calmodulin (CaM)
Length = 148
Score = 42.0 bits (97), Expect = 0.002
Identities = 36/139 (25%), Positives = 62/139 (43%), Gaps = 17/139 (12%)
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D++G + F E+L +S
Sbjct: 16 FSLFDKDG-DGTITTKELGTVMRSLGQNPTEAELQDMINEVDQDGSGTIDFPEFLTLMSR 74
Query: 175 K----DIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
K D E++ + E F V D D NG ++ ELR H + +++ V+
Sbjct: 75 KMHDTDTEEE---------IKEAFRVFDKDGNGFISAAELR---HVMTNLGEKLTDEEVD 122
Query: 231 DKFNMDDYEHDGKINFNQF 249
+ D + DG+IN+ +F
Sbjct: 123 EMIREADVDGDGQINYEEF 141
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,084,859
Number of Sequences: 164201
Number of extensions: 2226020
Number of successful extensions: 5363
Number of sequences better than 10.0: 211
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 199
Number of HSP's that attempted gapping in prelim test: 5149
Number of HSP's gapped (non-prelim): 330
length of query: 360
length of database: 59,974,054
effective HSP length: 111
effective length of query: 249
effective length of database: 41,747,743
effective search space: 10395188007
effective search space used: 10395188007
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 66 (30.0 bits)
Medicago: description of AC149127.7


