

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149080.7 - phase: 0
(165 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
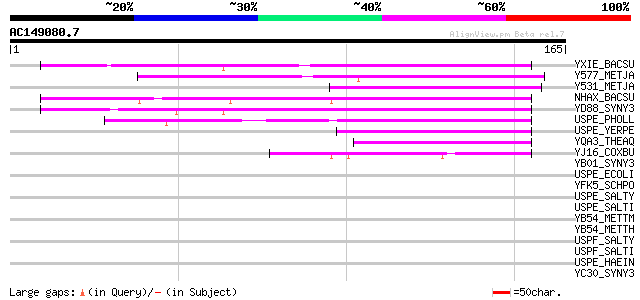
YXIE_BACSU (P42297) Hypothetical protein yxiE precursor 56 3e-08
Y577_METJA (Q57997) Protein MJ0577 55 6e-08
Y531_METJA (Q57951) Hypothetical protein MJ0531 54 2e-07
NHAX_BACSU (O07552) Stress response protein nhaX 49 5e-06
YD88_SYNY3 (P74148) Hypothetical protein sll1388 48 9e-06
USPE_PHOLL (P60005) Universal stress protein E 45 7e-05
USPE_YERPE (Q8ZE81) Universal stress protein E 44 2e-04
YQA3_THEAQ (P74897) Hypothetical 14.6 kDa protein in QAH/OAS sul... 44 2e-04
YJ16_COXBU (P45680) Hypothetical protein CBU1916 43 3e-04
YB01_SYNY3 (P72745) Hypothetical protein slr1101 41 0.001
USPE_ECOLI (P03807) Universal stress protein E 41 0.001
YFK5_SCHPO (P87132) Hypothetical protein C167.05 in chromosome I 40 0.002
USPE_SALTY (Q8ZP84) Universal stress protein E 40 0.002
USPE_SALTI (Q8Z788) Universal stress protein E 40 0.002
YB54_METTM (Q50777) Hypothetical 16.1 kDa protein in MTR region ... 38 0.009
YB54_METTH (O27222) Hypothetical protein MTH1154 38 0.012
USPF_SALTY (P67091) Universal stress protein F 38 0.012
USPF_SALTI (P67092) Universal stress protein F 38 0.012
USPE_HAEIN (P44195) Universal stress protein E homolog 37 0.020
YC30_SYNY3 (P73475) Hypothetical protein slr1230 36 0.035
>YXIE_BACSU (P42297) Hypothetical protein yxiE precursor
Length = 148
Score = 56.2 bits (134), Expect = 3e-08
Identities = 45/149 (30%), Positives = 72/149 (48%), Gaps = 7/149 (4%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYL---FSSD 66
+++VAID + S AL + +L + + IL + V S+ G Y+ F +
Sbjct: 4 KMLVAIDGSDMSAKALDAAV-HLAKEQQAELSILHVGREAVVTTSSLTGIVYVPEHFIDE 62
Query: 67 ITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSH 126
I ++K ++ + EKA ET NG+P I ++ GV ++V+GS
Sbjct: 63 IRNEVKKEGLKILENAKEKA---AEKGVQAETIYANGEPAHEILNHAKEKGVSLIVVGSR 119
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
G +K LGSVS+ +Q CPVLIV+
Sbjct: 120 GISGLKEMMLGSVSHKVSQLSTCPVLIVR 148
>Y577_METJA (Q57997) Protein MJ0577
Length = 162
Score = 55.5 bits (132), Expect = 6e-08
Identities = 36/132 (27%), Positives = 69/132 (52%), Gaps = 14/132 (10%)
Query: 39 DHLILLYVKPPRVVYSAFDGTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVET 98
+ +ILL+V R + + L + + ++E++ ++ + + E+AK N ++N++
Sbjct: 34 EEVILLHVIDEREIKKRDIFSLLLGVAGLNKSVEEFENELKNKLTEEAK---NKMENIKK 90
Query: 99 RIEN-----------GDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNV 147
+E+ G P + I + + GVDI++MGSHG +K LGSV+ + +
Sbjct: 91 ELEDVGFKVKDIIVVGIPHEEIVKIAEDEGVDIIIMGSHGKTNLKEILLGSVTENVIKKS 150
Query: 148 KCPVLIVKKPKS 159
PVL+VK+ S
Sbjct: 151 NKPVLVVKRKNS 162
>Y531_METJA (Q57951) Hypothetical protein MJ0531
Length = 170
Score = 53.5 bits (127), Expect = 2e-07
Identities = 25/63 (39%), Positives = 38/63 (59%)
Query: 96 VETRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
+ T + G P + I + +K D++VMG+ G ++R LGSV+ +N CPVL+VK
Sbjct: 106 IHTEMLEGVPANEIVEFAEKKKADLIVMGTTGKTGLERILLGSVAERVIKNAHCPVLVVK 165
Query: 156 KPK 158
KPK
Sbjct: 166 KPK 168
>NHAX_BACSU (O07552) Stress response protein nhaX
Length = 166
Score = 48.9 bits (115), Expect = 5e-06
Identities = 45/163 (27%), Positives = 74/163 (44%), Gaps = 19/163 (11%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNS----------KDHLILLYVKPPRVVYSAFDGT 59
RI+VA D E S AL + N+ KD+ + + PPR A +
Sbjct: 6 RIIVAFDGSENSKKALLTAIDLAKTVNAAITVAHSHDMKDNQTV--IDPPRPAAEASYIS 63
Query: 60 GYLFS------SDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQA 112
G + S SD+T+ + + V+ +A+++ N+ Q + + I GDP + I +
Sbjct: 64 GGMTSVPDPLISDVTSPEPMIYEDRTEEVIAEARMMLNEQQADGDIDILEGDPAESIIEH 123
Query: 113 VQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
++ D++V GS +K+ GSVS + PVLIVK
Sbjct: 124 ANRISADMIVTGSRDQNRLKKLIFGSVSEKLSAKSDIPVLIVK 166
>YD88_SYNY3 (P74148) Hypothetical protein sll1388
Length = 154
Score = 48.1 bits (113), Expect = 9e-06
Identities = 39/151 (25%), Positives = 69/151 (44%), Gaps = 7/151 (4%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKP----PRVVYSAFDGTGYL-FS 64
+I+VA+D E + L + + Q L++ Y P +Y +F G + FS
Sbjct: 5 KILVALDRSELAKEVLQQAIA--LGQKESSQLMVFYCIPVDSQDLSIYPSFYGEAAIGFS 62
Query: 65 SDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMG 124
I +E+ + + + + V D E ++ G+P I + D++V+G
Sbjct: 63 QIIKEHLEEQQTEAREWLQSIVQQVQEDGVACEWDVKVGEPGRWIRDMAKNWDADLVVLG 122
Query: 125 SHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
G + FLGSVS++ +V+C VLIV+
Sbjct: 123 RRGLKGLAEVFLGSVSSYVIHHVQCSVLIVQ 153
>USPE_PHOLL (P60005) Universal stress protein E
Length = 314
Score = 45.1 bits (105), Expect = 7e-05
Identities = 35/128 (27%), Positives = 56/128 (43%), Gaps = 10/128 (7%)
Query: 29 LKNLVFQNSKDHLILLY-VKPPRVVYSAFDGTGYLFSSDITATMEKYSQQVADCVLEKAK 87
L N V +N + HL+ Y V P + D +++ + Q + E +
Sbjct: 183 LANQVMKNPEIHLVSAYPVAPLNIAIELPDFNPSVYNHALRG-------QHLIAMKELRQ 235
Query: 88 IVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNV 147
C D + T I G P VI Q +M I+V+G G + AFLG+ + H ++
Sbjct: 236 TFCIDEKY--THIHEGLPESVIPQMCDEMNAGIIVLGILGRTGLSAAFLGNTAEHVIDHL 293
Query: 148 KCPVLIVK 155
KC +L +K
Sbjct: 294 KCDILTIK 301
>USPE_YERPE (Q8ZE81) Universal stress protein E
Length = 318
Score = 43.9 bits (102), Expect = 2e-04
Identities = 18/58 (31%), Positives = 33/58 (56%)
Query: 98 TRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
T +E G P +VI + + ++V+G+ G + AF+G+ + H N+KC +L +K
Sbjct: 243 THVEKGLPEEVIPDLAEHLNAGVVVLGTLGRTGLSAAFIGNTTEHVIDNLKCDLLAIK 300
Score = 29.6 bits (65), Expect = 3.2
Identities = 33/149 (22%), Positives = 64/149 (42%), Gaps = 9/149 (6%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYS-AFDGTGYLFSSDI 67
+ I+VAID ++ AL + LV +N +K +Y ++D T L +
Sbjct: 5 QNILVAIDPNQDDQPALRRAVY-LVQRNGGT------IKAFLAIYDLSYDMTTLLSPDER 57
Query: 68 TATMEKYSQQVADCVLEKAKIVCNDVQNVETRIE-NGDPRDVICQAVQKMGVDILVMGSH 126
TA + Q + + E+ + + ++ ++ + P + I Q V K D+L+ +H
Sbjct: 58 TAMRKGVISQRSAWISEQCRFYIDSGIPIDIKVVWHNRPYEAIIQEVLKAKHDLLLKMAH 117
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
+ ++ H + CPV +VK
Sbjct: 118 QHDRLESIIFTPTDWHLLRKCPCPVWMVK 146
>YQA3_THEAQ (P74897) Hypothetical 14.6 kDa protein in QAH/OAS
sulfhydrylase 3'region
Length = 137
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/53 (39%), Positives = 28/53 (52%)
Query: 103 GDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
G P + I QA D++VMG+ G G + FLGS S CPVL+V+
Sbjct: 85 GRPAEAILQAAIGEKADLIVMGTRGLGAVGSLFLGSQSQKVVAEAPCPVLLVR 137
>YJ16_COXBU (P45680) Hypothetical protein CBU1916
Length = 146
Score = 43.1 bits (100), Expect = 3e-04
Identities = 28/85 (32%), Positives = 48/85 (55%), Gaps = 9/85 (10%)
Query: 78 VADCVLEKAKIVCNDVQ---NVETR---IENGDPRDVICQAVQKMGVDILVMGSHG-YGV 130
V D +LE+AK N++ N+ + ++ G + +I + + GVD++++GSHG +G+
Sbjct: 60 VEDMLLEEAKKRMNEIASQLNISSDHQIVKVGPAKFLILEQAKNWGVDLIIVGSHGRHGI 119
Query: 131 IKRAFLGSVSNHCAQNVKCPVLIVK 155
+ LGS SN KC VL V+
Sbjct: 120 --QLLLGSTSNAVLHGAKCDVLAVR 142
>YB01_SYNY3 (P72745) Hypothetical protein slr1101
Length = 108
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/55 (34%), Positives = 28/55 (50%)
Query: 103 GDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
G P +ICQ ++ DI+V+G G + LGSV N+ + C V +V P
Sbjct: 53 GSPGKIICQRAKQDNSDIIVVGHRGRWGLSEILLGSVGNYVFHHAHCCVFVVPTP 107
>USPE_ECOLI (P03807) Universal stress protein E
Length = 315
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/64 (29%), Positives = 35/64 (54%)
Query: 98 TRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
T +E G P +VI + + I+V+G+ G I AFLG+ + +++C +L++K
Sbjct: 242 THVEKGLPEEVIPDLAEHLQAGIVVLGTVGRTGISAAFLGNTAEQVIDHLRCDLLVIKPD 301
Query: 158 KSTT 161
+ T
Sbjct: 302 QYQT 305
>YFK5_SCHPO (P87132) Hypothetical protein C167.05 in chromosome I
Length = 601
Score = 40.4 bits (93), Expect = 0.002
Identities = 37/152 (24%), Positives = 67/152 (43%), Gaps = 24/152 (15%)
Query: 13 VAIDEGEESIYALTWCLKNLVFQNSKDHLILLYV-----KPPRVVYSAFDGTGYLFSSDI 67
+ +D ES++A W + L+ + D LI++ V R V + S+
Sbjct: 435 LTLDLSSESLHAAEWAVGILL--RNGDTLIIVDVIECDDPSARAVKDRME-------SEQ 485
Query: 68 TATMEKYSQQVADCVLEKAKIVCNDVQNVETRIE---NGDPRDVICQAVQKMGVDILVMG 124
T+EK ++ + K++ V VE IE + + +I + + + ++VMG
Sbjct: 486 LETLEKITKYIL-------KLLSKTVLEVEVNIEVIHHEKAKHLIIEMIDYIEPSLVVMG 538
Query: 125 SHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
S G +K LGS SN+ PV++ +K
Sbjct: 539 SRGRSHLKGVLLGSFSNYLVNKSSVPVMVARK 570
>USPE_SALTY (Q8ZP84) Universal stress protein E
Length = 314
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/64 (28%), Positives = 35/64 (54%)
Query: 98 TRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
T +E G P +VI + + I+V+G+ G + AFLG+ + +++C +L++K
Sbjct: 242 THVEKGLPEEVIPDLAEHLQAGIVVLGTVGRTGLSAAFLGNTAEQVIDHLRCDLLVIKPD 301
Query: 158 KSTT 161
+ T
Sbjct: 302 EYQT 305
>USPE_SALTI (Q8Z788) Universal stress protein E
Length = 314
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/64 (28%), Positives = 35/64 (54%)
Query: 98 TRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
T +E G P +VI + + I+V+G+ G + AFLG+ + +++C +L++K
Sbjct: 242 THVEKGLPEEVIPDLAEHLQAGIVVLGTVGRTGLSAAFLGNTAEQVIDHLRCDLLVIKPD 301
Query: 158 KSTT 161
+ T
Sbjct: 302 EYQT 305
>YB54_METTM (Q50777) Hypothetical 16.1 kDa protein in MTR region
(ORF143)
Length = 143
Score = 38.1 bits (87), Expect = 0.009
Identities = 24/97 (24%), Positives = 46/97 (46%), Gaps = 12/97 (12%)
Query: 70 TMEKYSQQVADCVLEKAKIVCNDVQNVETRIEN-----------GDPRDVICQAVQKMGV 118
T +K + + + E+ K + D++ T EN G+P D I + ++ V
Sbjct: 46 TPKKVKEMMVKELTERGKEILRDMEKGLTGPENPNVKFRGVMLEGNPADEIVKLAEEEDV 105
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
D+++MG+ G ++ + LGSVS C + +V+
Sbjct: 106 DVIIMGT-GKSLVDKHLLGSVSEKVVHYAPCTIHLVR 141
>YB54_METTH (O27222) Hypothetical protein MTH1154
Length = 146
Score = 37.7 bits (86), Expect = 0.012
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 1/56 (1%)
Query: 100 IENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
+ GDP D I + ++ VD++VMG+ G ++ + LGSVS C + +V+
Sbjct: 90 MREGDPADEIVKVAEEEDVDVIVMGT-GKSLVDKHLLGSVSEKVVHYAPCTIHLVR 144
>USPF_SALTY (P67091) Universal stress protein F
Length = 144
Score = 37.7 bits (86), Expect = 0.012
Identities = 32/149 (21%), Positives = 67/149 (44%), Gaps = 9/149 (6%)
Query: 9 RRIMVAID--EGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSD 66
R I+V ID + E + ++ ++K H + + P + Y A G Y
Sbjct: 3 RTILVPIDISDSELTQRVISHVEAEAKIDDAKVHFLTVI---PSLPYYASLGLAYSAELP 59
Query: 67 ITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSH 126
++ ++ + +++K + + VQ + G P+D I + +K+ D++++ SH
Sbjct: 60 AMDDLKAEAKSQLEAIIKKFNLPADRVQ---AHVAEGSPKDKILEMAKKLPADMVIIASH 116
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
I LGS + ++ +C VL+V+
Sbjct: 117 RPD-ITTYLLGSNAAAVVRHAECSVLVVR 144
>USPF_SALTI (P67092) Universal stress protein F
Length = 144
Score = 37.7 bits (86), Expect = 0.012
Identities = 32/149 (21%), Positives = 67/149 (44%), Gaps = 9/149 (6%)
Query: 9 RRIMVAID--EGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSD 66
R I+V ID + E + ++ ++K H + + P + Y A G Y
Sbjct: 3 RTILVPIDISDSELTQRVISHVEAEAKIDDAKVHFLTVI---PSLPYYASLGLAYSAELP 59
Query: 67 ITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSH 126
++ ++ + +++K + + VQ + G P+D I + +K+ D++++ SH
Sbjct: 60 AMDDLKAEAKSQLEAIIKKFNLPADRVQ---AHVAEGSPKDKILEMAKKLPADMVIIASH 116
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
I LGS + ++ +C VL+V+
Sbjct: 117 RPD-ITTYLLGSNAAAVVRHAECSVLVVR 144
>USPE_HAEIN (P44195) Universal stress protein E homolog
Length = 309
Score = 37.0 bits (84), Expect = 0.020
Identities = 15/61 (24%), Positives = 33/61 (53%)
Query: 98 TRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
T + G P +VI + +++ +++++G+ G + A LG+ + H + C +L +K
Sbjct: 246 THVREGFPEEVIPEVAKEIEAELVILGTVGRTGLSAALLGNTAEHVISKLSCNLLGIKPS 305
Query: 158 K 158
K
Sbjct: 306 K 306
>YC30_SYNY3 (P73475) Hypothetical protein slr1230
Length = 287
Score = 36.2 bits (82), Expect = 0.035
Identities = 21/74 (28%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Query: 79 ADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGS 138
A+ VLEKA +E + G + I + + +D+L+MG+HG+ I+ +GS
Sbjct: 217 AEKVLEKAGF------KLEVELLVGHAEEAIVRYQEDNAIDLLLMGAHGHSRIRHLVIGS 270
Query: 139 VSNHCAQNVKCPVL 152
+ + PVL
Sbjct: 271 TTAQVLRKTSIPVL 284
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.137 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,698,611
Number of Sequences: 164201
Number of extensions: 769302
Number of successful extensions: 1776
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1734
Number of HSP's gapped (non-prelim): 63
length of query: 165
length of database: 59,974,054
effective HSP length: 102
effective length of query: 63
effective length of database: 43,225,552
effective search space: 2723209776
effective search space used: 2723209776
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149080.7


