

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
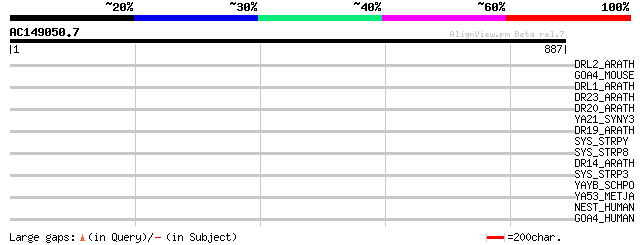
Query= AC149050.7 + phase: 0 /pseudo
(887 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
DRL2_ARATH (P60839) Probable disease resistance protein At1g12290 38 0.14
GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member... 37 0.23
DRL1_ARATH (P60838) Probable disease resistance protein At1g12280 37 0.30
DR23_ARATH (O82484) Putative disease resistance protein At4g10780 37 0.30
DR20_ARATH (Q9SH22) Putative disease resistance protein At1g63360 36 0.39
YA21_SYNY3 (P72929) Hypothetical protein sll1021 34 1.5
DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350 34 1.5
SYS_STRPY (Q99YE2) Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--... 33 4.4
SYS_STRP8 (Q8NZN7) Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--... 33 4.4
DR14_ARATH (Q9SI85) Probable disease resistance protein At1g6263... 33 4.4
SYS_STRP3 (Q8K635) Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--... 32 5.7
YAYB_SCHPO (Q10218) Hypothetical protein C4H3.11c in chromosome I 32 9.7
YA53_METJA (Q58453) Hypothetical protein MJ1053 32 9.7
NEST_HUMAN (P48681) Nestin 32 9.7
GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member... 32 9.7
>DRL2_ARATH (P60839) Probable disease resistance protein At1g12290
Length = 884
Score = 37.7 bits (86), Expect = 0.14
Identities = 19/57 (33%), Positives = 32/57 (55%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENE 85
Y+ K+ T+LEE +E LK ++ L +V+T G + ++ WL V TIE++
Sbjct: 28 YIQNIKENLTSLEEAMEDLKALRDDLLRKVQTAEEGGLQRLHQIKVWLKRVKTIESQ 84
>GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member 4
(tGolgin-1)
Length = 2238
Score = 37.0 bits (84), Expect = 0.23
Identities = 27/100 (27%), Positives = 48/100 (48%), Gaps = 5/100 (5%)
Query: 18 LVVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLY 77
+ +E + + LTQ K+ +L LE L++ K+A+ T E + + E+ + +
Sbjct: 533 IALEKSRSEYLKLTQEKEQQESLA--LEELELQKKAILTESENKLQ---ELGQEAEAYRT 587
Query: 78 DVTTIENELQKWLSDDNAQI*HLIIHWERKPLKALKILQA 117
+ +E L+K L + Q HL +H E + K K L A
Sbjct: 588 RILELETSLEKSLQESKTQSEHLAVHLEAEKNKHNKELTA 627
>DRL1_ARATH (P60838) Probable disease resistance protein At1g12280
Length = 894
Score = 36.6 bits (83), Expect = 0.30
Identities = 20/75 (26%), Positives = 43/75 (56%), Gaps = 1/75 (1%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETER-RKGYEIAPNVQKWLYDVTTIENELQ 87
Y+ + K +++++E LK + ++ RV+ E + E VQ WL +V+T+EN+
Sbjct: 28 YICELSKNVVAMKKDMEVLKKKRDDVKRRVDIEEFTRRRERLSQVQGWLTNVSTVENKFN 87
Query: 88 KWLSDDNAQI*HLII 102
+ L+ ++A++ L +
Sbjct: 88 ELLTTNDAELQRLCL 102
>DR23_ARATH (O82484) Putative disease resistance protein At4g10780
Length = 892
Score = 36.6 bits (83), Expect = 0.30
Identities = 24/75 (32%), Positives = 34/75 (45%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y+ + K LE+ +E L + + RV+ E KG E VQ WL V I N+
Sbjct: 28 YIHKLKDNIVALEKAIEDLTATRDDVLRRVQMEEGKGLERLQQVQVWLKRVEIIRNQFYD 87
Query: 89 WLSDDNAQI*HLIIH 103
LS N +I L +
Sbjct: 88 LLSARNIEIQRLCFY 102
>DR20_ARATH (Q9SH22) Putative disease resistance protein At1g63360
Length = 884
Score = 36.2 bits (82), Expect = 0.39
Identities = 21/74 (28%), Positives = 36/74 (48%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y +K LE+ ++ LK + L+ R++ E +G + Q WL V T+E+ +
Sbjct: 26 YTHNLEKNLAALEKTMKELKAKRDDLERRLKREEARGLQRLSEFQVWLDSVATVEDIIIT 85
Query: 89 WLSDDNAQI*HLII 102
L D N +I L +
Sbjct: 86 LLRDRNVEIQRLCL 99
>YA21_SYNY3 (P72929) Hypothetical protein sll1021
Length = 673
Score = 34.3 bits (77), Expect = 1.5
Identities = 37/122 (30%), Positives = 50/122 (40%), Gaps = 7/122 (5%)
Query: 6 ELGKETVTKLGELVVESTMKHFKYLTQHKKITTNLEEELERLKMIKQ-----ALQTRVET 60
EL + GEL E K + IT E EL +L K+ A Q R
Sbjct: 278 ELTTRVAIEQGELEAEKKSLAIKREQEDANITQQKEIELLKLAQRKELESQEAQQQREIQ 337
Query: 61 ERRKGYEIAPNVQKWLYDVTTIENELQKWLSDDNAQI*HLIIHWERKPLKALKILQA*RK 120
E + E K L + E +QK L+ N+QI I ER K LK+ QA +K
Sbjct: 338 EAKDKEEAKKERNKILQEQAVEEERIQKELAIQNSQIASAIALEERN--KELKVAQALQK 395
Query: 121 KK 122
++
Sbjct: 396 QE 397
>DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350
Length = 898
Score = 34.3 bits (77), Expect = 1.5
Identities = 20/74 (27%), Positives = 36/74 (48%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y +K LE +E LK + L +++ E +G + ++ WL V TIE+ +
Sbjct: 26 YTHNLEKNLVALETTMEELKAKRDDLLRKLKREEDRGLQTLGEIKVWLNRVETIESRVND 85
Query: 89 WLSDDNAQI*HLII 102
L+ NA++ L +
Sbjct: 86 LLNARNAELQRLCL 99
>SYS_STRPY (Q99YE2) Seryl-tRNA synthetase (EC 6.1.1.11)
(Serine--tRNA ligase) (SerRS)
Length = 425
Score = 32.7 bits (73), Expect = 4.4
Identities = 18/68 (26%), Positives = 34/68 (49%)
Query: 19 VVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYD 78
V E T+ H K L + ++ EEL+ + I A + + ++ + ++QK D
Sbjct: 23 VSEDTLTHLKELDEKRRALLVQSEELKAERNIASAAIAQAKRQKEDATQQIADMQKVSAD 82
Query: 79 VTTIENEL 86
+ TI+N+L
Sbjct: 83 IKTIDNQL 90
>SYS_STRP8 (Q8NZN7) Seryl-tRNA synthetase (EC 6.1.1.11)
(Serine--tRNA ligase) (SerRS)
Length = 425
Score = 32.7 bits (73), Expect = 4.4
Identities = 18/68 (26%), Positives = 34/68 (49%)
Query: 19 VVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYD 78
V E T+ H K L + ++ EEL+ + I A + + ++ + ++QK D
Sbjct: 23 VSEDTLTHLKELDEKRRTLLVQSEELKAERNIASAAIAQAKRQKEDATQQIADMQKVSAD 82
Query: 79 VTTIENEL 86
+ TI+N+L
Sbjct: 83 IKTIDNQL 90
>DR14_ARATH (Q9SI85) Probable disease resistance protein At1g62630
(pNd4)
Length = 893
Score = 32.7 bits (73), Expect = 4.4
Identities = 21/74 (28%), Positives = 34/74 (45%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y +K LE +E LK + L R++ E +G + Q WL V T+E+ +
Sbjct: 26 YTHNLEKNLVALETTMEELKAKRDDLLRRLKREEDRGLQRLSEFQVWLNRVATVEDIIIT 85
Query: 89 WLSDDNAQI*HLII 102
L D + +I L +
Sbjct: 86 LLRDRDVEIQRLCL 99
>SYS_STRP3 (Q8K635) Seryl-tRNA synthetase (EC 6.1.1.11)
(Serine--tRNA ligase) (SerRS)
Length = 425
Score = 32.3 bits (72), Expect = 5.7
Identities = 18/68 (26%), Positives = 34/68 (49%)
Query: 19 VVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYD 78
V E T+ H K L + ++ EEL+ + I A + + ++ + ++QK D
Sbjct: 23 VSEDTLTHLKELDEKRRTLLVQSEELKAGRNIASAAIAQAKRQKEDATQQIADMQKVSAD 82
Query: 79 VTTIENEL 86
+ TI+N+L
Sbjct: 83 IKTIDNQL 90
>YAYB_SCHPO (Q10218) Hypothetical protein C4H3.11c in chromosome I
Length = 783
Score = 31.6 bits (70), Expect = 9.7
Identities = 32/125 (25%), Positives = 57/125 (45%), Gaps = 14/125 (11%)
Query: 7 LGKETVTKLGELVVESTMKHFKY---------LTQHKKITTNLEEELERLKMI-KQALQT 56
+ KE +TKLGE V+ M +Y L++H K +L+E+ E ++ + K L
Sbjct: 342 MSKEMLTKLGESNVDDGMAASRYDTVKREKERLSEHLK---SLQEQYEHIQSVYKNVLLD 398
Query: 57 RVETERRKGYEIAPNVQKWLYDVTTIENELQKWLSDDNAQI*HLIIHWERKPLKALKILQ 116
R R G +I+ N + L + ++ +LQ +L + + I K L + +
Sbjct: 399 RESYIMRLGNKISEN-NELLNENRVLKEKLQTYLDKKESNVTSKIKSTAENSSKPLSMNE 457
Query: 117 A*RKK 121
A +K
Sbjct: 458 ADERK 462
>YA53_METJA (Q58453) Hypothetical protein MJ1053
Length = 163
Score = 31.6 bits (70), Expect = 9.7
Identities = 24/76 (31%), Positives = 41/76 (53%), Gaps = 7/76 (9%)
Query: 6 ELGKET--VTKLGELVVESTMKHFKYLTQHK---KITT--NLEEELERLKMIKQALQTRV 58
+L +ET + K GEL +E K + + + + KIT LEEE+E+LK + L ++
Sbjct: 88 KLVRETYEMMKRGELNIEEVEKFLEAVVRKEELEKITDIKKLEEEIEKLKKENEELAAKL 147
Query: 59 ETERRKGYEIAPNVQK 74
E + K E+ ++K
Sbjct: 148 EKVKEKLKEVLSELEK 163
>NEST_HUMAN (P48681) Nestin
Length = 1618
Score = 31.6 bits (70), Expect = 9.7
Identities = 25/99 (25%), Positives = 49/99 (49%), Gaps = 12/99 (12%)
Query: 2 DCLTELGKETVTKLGELVVES-----TMKHFKYLTQHKKITTNLEEELERLKMIKQALQT 56
D L KET+ LGE + ES H ++++ +LEE+LE LK +++ +
Sbjct: 535 DSQVPLEKETLKSLGEEIQESLKTLENQSHETLERENQECPRSLEEDLETLKSLEKENKR 594
Query: 57 RV------ETERRKG-YEIAPNVQKWLYDVTTIENELQK 88
+ ET R++G ++ P ++ + +++ E Q+
Sbjct: 595 AIKGCGGSETSRKRGCRQLKPTGKEDTQTLQSLQKENQE 633
>GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member 4
(Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (72.1
protein)
Length = 2230
Score = 31.6 bits (70), Expect = 9.7
Identities = 26/109 (23%), Positives = 53/109 (47%), Gaps = 7/109 (6%)
Query: 6 ELGKETVTKLGELVVESTMKHFKYLTQHKKITTNLEEE----LERLKMIKQALQTRVETE 61
EL K+ T+ E + + K +++ KI+ E++ LE L++ K+A+ T E +
Sbjct: 495 ELTKKLQTREREFQEQMKVALEKSQSEYLKISQEKEQQESLALEELELQKKAILTESENK 554
Query: 62 RRKGYEIAPNVQKWLYDVTTIENELQKWLSDDNAQI*HLIIHWERKPLK 110
R ++ + + + +E+ L+K L ++ Q L +H E + K
Sbjct: 555 LR---DLQQEAETYRTRILELESSLEKSLQENKNQSKDLAVHLEAEKNK 600
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.362 0.160 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,834,581
Number of Sequences: 164201
Number of extensions: 2965484
Number of successful extensions: 11528
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 11510
Number of HSP's gapped (non-prelim): 27
length of query: 887
length of database: 59,974,054
effective HSP length: 119
effective length of query: 768
effective length of database: 40,434,135
effective search space: 31053415680
effective search space used: 31053415680
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149050.7


