

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149039.3 + phase: 0 /pseudo
(2185 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
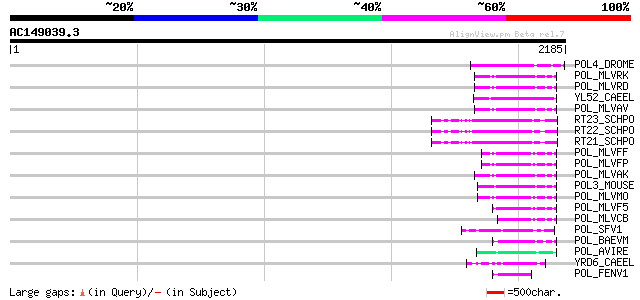
Score E
Sequences producing significant alignments: (bits) Value
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 122 1e-26
POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC 3.4.2... 103 4e-21
POL_MLVRD (P11227) Pol polyprotein [Contains: Protease (EC 3.4.2... 103 4e-21
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 103 5e-21
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 102 9e-21
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 101 2e-20
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 101 2e-20
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 101 2e-20
POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.2... 101 2e-20
POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC 3.4.2... 100 3e-20
POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse transcript... 100 4e-20
POL3_MOUSE (P11367) Retrovirus-related Pol polyprotein (Endonucl... 100 4e-20
POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.2... 100 8e-20
POL_MLVF5 (P26810) Pol polyprotein [Contains: Protease (EC 3.4.2... 100 8e-20
POL_MLVCB (P08361) Pol polyprotein [Contains: Reverse transcript... 96 1e-18
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 91 3e-17
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 89 2e-16
POL_AVIRE (P03360) Pol polyprotein [Contains: Reverse transcript... 88 3e-16
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 87 5e-16
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 86 1e-15
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-); Reverse
transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 122 bits (306), Expect = 1e-26
Identities = 100/381 (26%), Positives = 170/381 (44%), Gaps = 29/381 (7%)
Query: 1814 RNYDMVLLRCV----DEHEAEQLMHDVHDGTFRTHATGHTMSRKLLRAGYYWMAMEHDCY 1869
+N + LL V +E E E ++ +HD + TG T + ++ YYW M
Sbjct: 874 KNLKVALLNPVTQINNEKEKEAILSTLHDDPIQGGHTGITKTLAKVKRHYYWKNMSKYIK 933
Query: 1870 QHARKCHKCQIYADKIHVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDY 1929
++ RKC KCQ H + F +D IG + PK+ NG+ + + I
Sbjct: 934 EYVRKCQKCQKAKTTKHTKTPMTITETPEHAFDRVVVDTIGPL-PKSENGNEYAVTLICD 992
Query: 1930 FTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEH 1989
TK++ A N + + VAK I + I +YG ITD GT N+++ LC+ KI++
Sbjct: 993 LTKYLVAIPIANKSAKTVAKAIFESFILKYGPMKTFITDMGTEYKNSIITDLCKYLKIKN 1052
Query: 1990 HNSSPYRPQMNGAVEATNKNIKRIVQKMVTTYK-DWHEMLPYALHGYRTTVRSSTGATPF 2048
S+ + Q G VE +++ + ++ ++T K DW L Y ++ + TT P+
Sbjct: 1053 ITSTAHHHQTVGVVERSHRTLNEYIRSYISTDKTDWDVWLQYFVYCFNTTQSMVHNYCPY 1112
Query: 2049 SLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARG----QS 2104
LV+G + LP KL E + D + + A AR ++
Sbjct: 1113 ELVFGRTSNLPKHFN----------KLHSIEPIYNIDDYAKESKYRLEVAYARARKLLEA 1162
Query: 2105 YQARMKTAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVVKK-AFSGGALI 2163
++ + K +D KV E +VG+ VL R + K Y GPY ++ + +
Sbjct: 1163 HKEKNKENYDLKVKDIELEVGDKVLLRN-----EVGHKLDFKYTGPYKIESIGDNNNITL 1217
Query: 2164 LTHMDGVELPNPVNADIVKKY 2184
LT+ + ++ V+ D +KK+
Sbjct: 1218 LTNKNKKQI---VHKDRLKKF 1235
Score = 37.7 bits (86), Expect = 0.35
Identities = 24/93 (25%), Positives = 44/93 (46%), Gaps = 1/93 (1%)
Query: 1470 IYYLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVT 1529
+ Y S+ FT E+ + E+ A+ WA R Y+ + + P+ Y+F +
Sbjct: 642 VAYASRAFTKGESNKSTTEQELAAIHWAIIHFRPYIYGKHFTVKTDHRPLTYLFSMVNPS 701
Query: 1530 GKIARWQVLLSEYDIVFKTQKAIKGSILADHLA 1562
K+ R ++ L EY+ + K K + +AD L+
Sbjct: 702 SKLTRIRLELEEYNFTVEYLKG-KDNHVADALS 733
>POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)] (Fragment)
Length = 581
Score = 103 bits (258), Expect = 4e-21
Identities = 90/330 (27%), Positives = 150/330 (45%), Gaps = 28/330 (8%)
Query: 1830 EQLMHDVHDGTFRTHATGHTMSRKLLRAG---YYWMAMEHDCYQHARKCHKC-QIYADKI 1885
+Q + ++ D R G+ + LL G YY + + A C C Q+ A K
Sbjct: 222 DQFVFELLDSLHRLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADSCTVCAQVNASKA 281
Query: 1886 HVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQ 1945
+ + P + W ID ++P G+ ++LV +D F+ WVEA + T +
Sbjct: 282 KIGAGVR--VRGHRPGTHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKHETAK 337
Query: 1946 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2005
+V K + I R+G+P + TDNG + V Q++ + I+ YRPQ +G VE
Sbjct: 338 IVTKKLLEEIFPRFGMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVER 397
Query: 2006 TNKNIKRIVQK--MVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
N+ IK + K + T +DW +LP AL+ R T G TP+ ++YG L +
Sbjct: 398 MNRTIKETLTKLTLATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAPPPL-VNFH 455
Query: 2064 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFK 2123
P + +K + + Q+ L ++ + +A +YQ ++ D+ V P F+
Sbjct: 456 DPEM-----SKFTNSPSLQAHLQALQAVQREVWKPLA--AAYQDQL----DQPVIPHPFR 504
Query: 2124 VGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
VG+ V RR + P ++GPY V
Sbjct: 505 VGDTVWVRRHQTK-----NLEPRWKGPYTV 529
>POL_MLVRD (P11227) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 103 bits (258), Expect = 4e-21
Identities = 90/330 (27%), Positives = 150/330 (45%), Gaps = 28/330 (8%)
Query: 1830 EQLMHDVHDGTFRTHATGHTMSRKLLRAG---YYWMAMEHDCYQHARKCHKC-QIYADKI 1885
+Q + ++ D R G+ + LL G YY + + A C C Q+ A K
Sbjct: 837 DQFVFELLDSLHRLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADSCTVCAQVNASKA 896
Query: 1886 HVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQ 1945
+ + P + W ID ++P G+ ++LV +D F+ WVEA + T +
Sbjct: 897 KIGAGVR--VRGHRPGTHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKHETAK 952
Query: 1946 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2005
+V K + I R+G+P + TDNG + V Q++ + I+ YRPQ +G VE
Sbjct: 953 IVTKKLLEEIFPRFGMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVER 1012
Query: 2006 TNKNIKRIVQK--MVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
N+ IK + K + T +DW +LP AL+ R T G TP+ ++YG L +
Sbjct: 1013 MNRTIKETLTKLTLATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAPPPL-VNFH 1070
Query: 2064 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFK 2123
P + +K + + Q+ L ++ + +A +YQ ++ D+ V P F+
Sbjct: 1071 DPEM-----SKFTNSPSLQAHLQALQAVQREVWKPLA--AAYQDQL----DQPVIPHPFR 1119
Query: 2124 VGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
VG+ V RR + P ++GPY V
Sbjct: 1120 VGDTVWVRRHQTK-----NLEPRWKGPYTV 1144
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 103 bits (257), Expect = 5e-21
Identities = 87/335 (25%), Positives = 145/335 (42%), Gaps = 14/335 (4%)
Query: 1824 VDEHEAEQLMHDVHDGTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYAD 1883
V E L+ ++H+G H M R + R +YW M R C KC D
Sbjct: 1460 VPEKIRTPLLKELHEGMLAGHFGIKKMWRMVHRK-FYWPQMRVCVENCVRTCAKCLCAND 1518
Query: 1884 KIHVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVT 1943
+ +L +P + D++ + G+ +IL ID FTK+ A +
Sbjct: 1519 HSKLTS-SLTPYRMTFPLEIVACDLMD--VGLSVQGNRYILTIIDLFTKYGTAVPIPDKK 1575
Query: 1944 KQVVAK-FIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGA 2002
+ V K F++ I +P K++TD G N + KIEH + Y + NGA
Sbjct: 1576 AETVLKAFVERWAIGEGRIPLKLLTDQGKEFVNGLFAQFTHMLKIEHITTKGYNSRANGA 1635
Query: 2003 VEATNKNIKRIVQKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2062
VE NK I I++K +W + + YA++ Y V +TG TP L++G + + PLE+
Sbjct: 1636 VERFNKTIMHIMKKKTAVPMEWDDQVVYAVYAYNNCVHENTGETPMFLMHGRDVMGPLEM 1695
Query: 2063 EIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVH---- 2118
I A + E + ++ + L + + + A+ +SY++ + K H
Sbjct: 1696 SGEDAVGINYADMDEYKHLLTQ-ELLKVQKIAKEHAMREQESYKSLFDQKYASKKHRFPQ 1754
Query: 2119 PREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
P + E+ ++ +Q P KW+ GPY V
Sbjct: 1755 PGSRVLLEIPSEKLGAQCPKLVNKWS----GPYRV 1785
Score = 34.7 bits (78), Expect = 3.0
Identities = 20/93 (21%), Positives = 45/93 (47%), Gaps = 1/93 (1%)
Query: 1470 IYYLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVT 1529
I + SK + ETRY + + A+ +A +R + + + + P+ + + + +
Sbjct: 1271 IAFASKALSPAETRYHITDLEALAMMFALRRFKTIIYGTAITVFTDHKPLISLLKGSPLA 1330
Query: 1530 GKIARWQVLLSEYDIVFKTQKAIKGSILADHLA 1562
++ RW + + E+D+ A K + +AD L+
Sbjct: 1331 DRLWRWSIEILEFDVKI-VYLAGKANAVADALS 1362
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 102 bits (255), Expect = 9e-21
Identities = 88/330 (26%), Positives = 151/330 (45%), Gaps = 28/330 (8%)
Query: 1830 EQLMHDVHDGTFRTHATGHTMSRKLLRAG---YYWMAMEHDCYQHARKCHKC-QIYADKI 1885
+Q + ++ D R G+ + LL G YY + + A C C Q+ A K
Sbjct: 837 DQFVFELLDSLHRLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADSCTVCAQVNASKA 896
Query: 1886 HVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQ 1945
+ + P S W ID ++P G+ ++LV +D F+ WVEA T +
Sbjct: 897 KIGAGVR--VRGHRPGSHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKRETAR 952
Query: 1946 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2005
VV+K + I R+G+P + +DNG + V Q++ + I+ YRPQ +G VE
Sbjct: 953 VVSKKLLEEIFPRFGMPQVLGSDNGPAFTSQVSQSVADLLGIDWKLHCAYRPQSSGQVER 1012
Query: 2006 TNKNIKRIVQKMVTT--YKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
N+ IK + K+ +DW +LP AL+ R T G TP+ ++YG L +
Sbjct: 1013 MNRTIKETLTKLTLAAGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAPPPL-VNFH 1070
Query: 2064 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFK 2123
P + ++L+ + Q+ L ++ + +A ++Y+ ++ D+ V P F+
Sbjct: 1071 DPDM-----SELTNSPSLQAHLQALQTVQREIWKPLA--EAYRDQL----DQPVIPHPFR 1119
Query: 2124 VGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
+G+ V RR + P ++GPY V
Sbjct: 1120 IGDSVWVRRHQTK-----NLEPRWKGPYTV 1144
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 101 bits (252), Expect = 2e-20
Identities = 115/505 (22%), Positives = 211/505 (41%), Gaps = 53/505 (10%)
Query: 1659 IDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQ 1718
++ I+ I D +I +I E E + +L ++ + + E+++ P N
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDF-----NFEINYRPGSANH 824
Query: 1719 MADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYYDIKQ 1778
+ADAL+ + PI K + V I D + V +Y D K
Sbjct: 825 IADALSRIVD-------ETEPIPKDSEDNSINFVNQISITDDFKNQVVTEYTN---DTKL 874
Query: 1779 FLLSREYPPGASKQDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHD 1838
L + +DK+ + L DG ++ + D +LL D ++ H+
Sbjct: 875 LNL-------LNNEDKRVEENIQ---LKDGLLINSK--DQILLPN-DTQLTRTIIKKYHE 921
Query: 1839 GTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVIS-S 1897
H ++ +LR + W + ++ + CH CQI + H P L I S
Sbjct: 922 EGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPS 980
Query: 1898 PWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYT-NVTKQVVAKFIKNNII 1956
P+ +D I + S+G++ + V +D F+K T ++T + A+ +I
Sbjct: 981 ERPWESLSMDFITALPE--SSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVI 1038
Query: 1957 CRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK 2016
+G P +II DN + + ++ S PYRPQ +G E TN+ ++++++
Sbjct: 1039 AYFGNPKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRC 1098
Query: 2017 MVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKL 2075
+ +T+ + W + + Y + S+T TPF +V+ L +E+PS +
Sbjct: 1099 VCSTHPNTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPALS-PLELPSFSDKTDENS 1157
Query: 2076 SEA-EWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHP-REFKVGELVL-KRR 2132
E + Q+ + LN + +MK FD K+ EF+ G+LV+ KR
Sbjct: 1158 QETIQVFQTVKEHLN--------------TNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT 1203
Query: 2133 ISQQPDPRGKWTPNYEGP-YVVKKA 2156
+ K P++ GP YV++K+
Sbjct: 1204 KTGFLHKSNKLAPSFAGPFYVLQKS 1228
Score = 38.9 bits (89), Expect = 0.16
Identities = 30/124 (24%), Positives = 57/124 (45%), Gaps = 20/124 (16%)
Query: 1472 YLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVTGK 1531
Y S K + + Y++ +K A+ + K RHYL S ++P K + + + G+
Sbjct: 737 YYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLE-------STIEPFKILTDHRNLIGR 789
Query: 1532 I-----------ARWQVLLSEYDIVFKTQKAIKGSILADHLAYQPLDDYQPIEFDFPDEE 1580
I ARWQ+ L +++ + + +AD L+ + +D+ +PI D D
Sbjct: 790 ITNESEPENKRLARWQLFLQDFNFEINYRPG-SANHIADALS-RIVDETEPIPKDSEDNS 847
Query: 1581 IMYL 1584
I ++
Sbjct: 848 INFV 851
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 101 bits (252), Expect = 2e-20
Identities = 115/505 (22%), Positives = 211/505 (41%), Gaps = 53/505 (10%)
Query: 1659 IDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQ 1718
++ I+ I D +I +I E E + +L ++ + + E+++ P N
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDF-----NFEINYRPGSANH 824
Query: 1719 MADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYYDIKQ 1778
+ADAL+ + PI K + V I D + V +Y D K
Sbjct: 825 IADALSRIVD-------ETEPIPKDSEDNSINFVNQISITDDFKNQVVTEYTN---DTKL 874
Query: 1779 FLLSREYPPGASKQDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHD 1838
L + +DK+ + L DG ++ + D +LL D ++ H+
Sbjct: 875 LNL-------LNNEDKRVEENIQ---LKDGLLINSK--DQILLPN-DTQLTRTIIKKYHE 921
Query: 1839 GTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVIS-S 1897
H ++ +LR + W + ++ + CH CQI + H P L I S
Sbjct: 922 EGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPS 980
Query: 1898 PWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYT-NVTKQVVAKFIKNNII 1956
P+ +D I + S+G++ + V +D F+K T ++T + A+ +I
Sbjct: 981 ERPWESLSMDFITALPE--SSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVI 1038
Query: 1957 CRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK 2016
+G P +II DN + + ++ S PYRPQ +G E TN+ ++++++
Sbjct: 1039 AYFGNPKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRC 1098
Query: 2017 MVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKL 2075
+ +T+ + W + + Y + S+T TPF +V+ L +E+PS +
Sbjct: 1099 VCSTHPNTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPALS-PLELPSFSDKTDENS 1157
Query: 2076 SEA-EWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHP-REFKVGELVL-KRR 2132
E + Q+ + LN + +MK FD K+ EF+ G+LV+ KR
Sbjct: 1158 QETIQVFQTVKEHLN--------------TNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT 1203
Query: 2133 ISQQPDPRGKWTPNYEGP-YVVKKA 2156
+ K P++ GP YV++K+
Sbjct: 1204 KTGFLHKSNKLAPSFAGPFYVLQKS 1228
Score = 38.9 bits (89), Expect = 0.16
Identities = 30/124 (24%), Positives = 57/124 (45%), Gaps = 20/124 (16%)
Query: 1472 YLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVTGK 1531
Y S K + + Y++ +K A+ + K RHYL S ++P K + + + G+
Sbjct: 737 YYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLE-------STIEPFKILTDHRNLIGR 789
Query: 1532 I-----------ARWQVLLSEYDIVFKTQKAIKGSILADHLAYQPLDDYQPIEFDFPDEE 1580
I ARWQ+ L +++ + + +AD L+ + +D+ +PI D D
Sbjct: 790 ITNESEPENKRLARWQLFLQDFNFEINYRPG-SANHIADALS-RIVDETEPIPKDSEDNS 847
Query: 1581 IMYL 1584
I ++
Sbjct: 848 INFV 851
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 101 bits (252), Expect = 2e-20
Identities = 115/505 (22%), Positives = 211/505 (41%), Gaps = 53/505 (10%)
Query: 1659 IDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQ 1718
++ I+ I D +I +I E E + +L ++ + + E+++ P N
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDF-----NFEINYRPGSANH 824
Query: 1719 MADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYYDIKQ 1778
+ADAL+ + PI K + V I D + V +Y D K
Sbjct: 825 IADALSRIVD-------ETEPIPKDSEDNSINFVNQISITDDFKNQVVTEYTN---DTKL 874
Query: 1779 FLLSREYPPGASKQDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHD 1838
L + +DK+ + L DG ++ + D +LL D ++ H+
Sbjct: 875 LNL-------LNNEDKRVEENIQ---LKDGLLINSK--DQILLPN-DTQLTRTIIKKYHE 921
Query: 1839 GTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVIS-S 1897
H ++ +LR + W + ++ + CH CQI + H P L I S
Sbjct: 922 EGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPS 980
Query: 1898 PWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYT-NVTKQVVAKFIKNNII 1956
P+ +D I + S+G++ + V +D F+K T ++T + A+ +I
Sbjct: 981 ERPWESLSMDFITALPE--SSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVI 1038
Query: 1957 CRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK 2016
+G P +II DN + + ++ S PYRPQ +G E TN+ ++++++
Sbjct: 1039 AYFGNPKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRC 1098
Query: 2017 MVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKL 2075
+ +T+ + W + + Y + S+T TPF +V+ L +E+PS +
Sbjct: 1099 VCSTHPNTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPALS-PLELPSFSDKTDENS 1157
Query: 2076 SEA-EWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHP-REFKVGELVL-KRR 2132
E + Q+ + LN + +MK FD K+ EF+ G+LV+ KR
Sbjct: 1158 QETIQVFQTVKEHLN--------------TNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT 1203
Query: 2133 ISQQPDPRGKWTPNYEGP-YVVKKA 2156
+ K P++ GP YV++K+
Sbjct: 1204 KTGFLHKSNKLAPSFAGPFYVLQKS 1228
Score = 38.9 bits (89), Expect = 0.16
Identities = 30/124 (24%), Positives = 57/124 (45%), Gaps = 20/124 (16%)
Query: 1472 YLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVTGK 1531
Y S K + + Y++ +K A+ + K RHYL S ++P K + + + G+
Sbjct: 737 YYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLE-------STIEPFKILTDHRNLIGR 789
Query: 1532 I-----------ARWQVLLSEYDIVFKTQKAIKGSILADHLAYQPLDDYQPIEFDFPDEE 1580
I ARWQ+ L +++ + + +AD L+ + +D+ +PI D D
Sbjct: 790 ITNESEPENKRLARWQLFLQDFNFEINYRPG-SANHIADALS-RIVDETEPIPKDSEDNS 847
Query: 1581 IMYL 1584
I ++
Sbjct: 848 INFV 851
>POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 101 bits (252), Expect = 2e-20
Identities = 87/298 (29%), Positives = 135/298 (45%), Gaps = 25/298 (8%)
Query: 1859 YYWMAMEHDCYQHARKCHKC-QIYADKIHVPPHALNVISSPWPFSMWGIDMIGRIEPKAS 1917
YY + + C C Q+ A K V + P + W ID ++P
Sbjct: 874 YYMLNRDRTLKDITETCQACAQVNASKSAVKQGTR--VRGHRPGTHWEIDFT-EVKP-GL 929
Query: 1918 NGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNV 1977
G+ ++LV ID F+ WVEA T +VV K + I R+G+P + TDNG + V
Sbjct: 930 YGYKYLLVFIDTFSGWVEAFPTKKETAKVVTKKLLEEIFPRFGMPQVLGTDNGPAFVSKV 989
Query: 1978 VQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--MVTTYKDWHEMLPYALHGY 2035
Q + + ++ YRPQ +G VE N+ IK + K + T +DW +LP AL+
Sbjct: 990 SQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLTLATGSRDWVLLLPLALYRA 1049
Query: 2036 RTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKR 2095
R T G TP+ ++YG P V P + AK++ Q+ L L++ +
Sbjct: 1050 RNT-PGPHGLTPYEILYGAP---PPLVNFPDPDM---AKVTHNPSLQAHLQALYLVQHEV 1102
Query: 2096 MDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
+A +YQ ++ D+ V P F+VG+ V RR + P ++GPY V
Sbjct: 1103 WRPLA--AAYQEQL----DRPVVPHPFRVGDTVWVRRHQTK-----NLEPRWKGPYTV 1149
>POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 100 bits (250), Expect = 3e-20
Identities = 86/298 (28%), Positives = 135/298 (44%), Gaps = 25/298 (8%)
Query: 1859 YYWMAMEHDCYQHARKCHKC-QIYADKIHVPPHALNVISSPWPFSMWGIDMIGRIEPKAS 1917
YY + + C C Q+ A K V + P + W ID ++P
Sbjct: 874 YYMLNRDRTLKDITETCKACAQVNASKSAVKQGTR--VRGHRPGTHWEIDFT-EVKP-GL 929
Query: 1918 NGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNV 1977
G+ ++LV +D F+ WVEA T +VV K + I R+G+P + TDNG + V
Sbjct: 930 YGYKYLLVFVDTFSGWVEAFPTKKETAKVVTKKLLEEIFPRFGMPQVLGTDNGPAFVSKV 989
Query: 1978 VQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--MVTTYKDWHEMLPYALHGY 2035
Q + + ++ YRPQ +G VE N+ IK + K + T +DW +LP AL+
Sbjct: 990 SQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLTLATGSRDWVLLLPLALYRA 1049
Query: 2036 RTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKR 2095
R T G TP+ ++YG P V P + AK++ Q+ L L++ +
Sbjct: 1050 RNT-PGPHGLTPYEILYGAP---PPLVNFPDPDM---AKVTHNPSLQAHLQALYLVQHEV 1102
Query: 2096 MDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
+A +YQ ++ D+ V P F+VG+ V RR + P ++GPY V
Sbjct: 1103 WRPLA--AAYQEQL----DRPVVPHPFRVGDTVWVRRHQTK-----NLEPRWKGPYTV 1149
>POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 843
Score = 100 bits (249), Expect = 4e-20
Identities = 88/330 (26%), Positives = 153/330 (45%), Gaps = 29/330 (8%)
Query: 1830 EQLMHDVHDGTFRTHATGHTMSRKLLRAG---YYWMAMEHDCYQHARKCHKC-QIYADKI 1885
+Q + ++ D R G+ + LL G YY + + A C C Q+ A K
Sbjct: 485 DQFVFELLDSLHRLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADSCTVCAQVNASKA 544
Query: 1886 HVPPHALNVISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQ 1945
+ + P S W ID ++P G+ ++LV +D F+ WVEA T +
Sbjct: 545 KIGAGVR--VRGHRPGSHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKRETAR 600
Query: 1946 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2005
VV+K + I R+G+P + +DNG + V Q++ + I+ + + YRPQ +G VE
Sbjct: 601 VVSKKLLEEIFPRFGMPQVLGSDNGPAFTSQVSQSVADLLGIDKLHCA-YRPQSSGQVER 659
Query: 2006 TNKNIKRIVQKMVTT--YKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
N+ IK + K+ +DW +LP AL+ R T G TP+ ++YG L +
Sbjct: 660 MNRTIKETLTKLTLAAGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAPPPL-VNFH 717
Query: 2064 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFK 2123
P + ++L+ + Q+ L ++ + +A ++Y+ ++ D+ V P F+
Sbjct: 718 DPDM-----SELTNSPSLQAHLQALQTVQREIWKPLA--EAYRDQL----DQPVIPHPFR 766
Query: 2124 VGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
+G+ V RR + P ++GPY V
Sbjct: 767 IGDSVWVRRHQTK-----NLEPRWKGPYTV 791
>POL3_MOUSE (P11367) Retrovirus-related Pol polyprotein (Endonuclease)
(Fragment)
Length = 390
Score = 100 bits (249), Expect = 4e-20
Identities = 88/316 (27%), Positives = 146/316 (45%), Gaps = 27/316 (8%)
Query: 1843 THATGHTMSRKLLR--AGYYWMAMEHDCYQHARKCHKC-QIYADKIHVPPHALNVISSPW 1899
TH + M L R + YY + + ++ A C C Q+ A K + A +
Sbjct: 60 THLSYQKMRALLDRKESPYYMLNKDKILHEVAESCQACVQVNASKTKI--RAGTRVRGHR 117
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
+ W ID ++P G+ ++LV +D F+ WVEA + T ++V K + I R+
Sbjct: 118 LGTHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKHETAKIVTKKLLEEIFPRF 175
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--M 2017
G+P + TDNG + V Q++ + I+ YRPQ +G VE N+ IK + K +
Sbjct: 176 GMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 235
Query: 2018 VTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSE 2077
T +DW +LP AL+ R T G TP+ ++YG L + P + +K +
Sbjct: 236 ATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAPPPL-VNFHDPEM-----SKFTN 288
Query: 2078 AEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQP 2137
+ Q+ L ++ + +A +YQ ++ D+ V P F+VG+ V RR +
Sbjct: 289 SPSLQAHLQALQAVQREVWKPLA--AAYQDQL----DQPVIPHPFRVGDTVWVRRHQTK- 341
Query: 2138 DPRGKWTPNYEGPYVV 2153
P ++GPY V
Sbjct: 342 ----NLEPRWKGPYTV 353
>POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1199
Score = 99.8 bits (247), Expect = 8e-20
Identities = 90/316 (28%), Positives = 142/316 (44%), Gaps = 27/316 (8%)
Query: 1843 THATGHTMSRKLLRAG--YYWMAMEHDCYQHARKCHKC-QIYADKIHVPPHALNVISSPW 1899
TH + M L R+ YY + + C C Q+ A K V +
Sbjct: 851 THLSFSKMKALLERSHSPYYMLNRDRTLKNITETCKACAQVNASKSAVKQGTR--VRGHR 908
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
P + W ID I+P G+ ++LV ID F+ W+EA T +VV K + I R+
Sbjct: 909 PGTHWEIDFT-EIKP-GLYGYKYLLVFIDTFSGWIEAFPTKKETAKVVTKKLLEEIFPRF 966
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--M 2017
G+P + TDNG + V Q + + I+ YRPQ +G VE N+ IK + K +
Sbjct: 967 GMPQVLGTDNGPAFVSKVSQTVADLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 1026
Query: 2018 VTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSE 2077
T +DW +LP AL+ R T G TP+ ++YG P V P + +++
Sbjct: 1027 ATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAP---PPLVNFPDPDM---TRVTN 1079
Query: 2078 AEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQP 2137
+ Q+ L L++ + +A +YQ ++ D+ V P ++VG+ V RR +
Sbjct: 1080 SPSLQAHLQALYLVQHEVWRPLA--AAYQEQL----DRPVVPHPYRVGDTVWVRRHQTK- 1132
Query: 2138 DPRGKWTPNYEGPYVV 2153
P ++GPY V
Sbjct: 1133 ----NLEPRWKGPYTV 1144
>POL_MLVF5 (P26810) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 99.8 bits (247), Expect = 8e-20
Identities = 78/256 (30%), Positives = 123/256 (47%), Gaps = 22/256 (8%)
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
P + W ID ++P G+ ++LV +D F+ WVEA T +VV K + I R+
Sbjct: 914 PGTHWEIDFT-EVKP-GLYGYKYLLVFVDTFSGWVEAFPTKKETAKVVTKKLLEEIFPRF 971
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--M 2017
G+P + TDNG + V Q + + ++ YRPQ +G VE N+ IK + K +
Sbjct: 972 GMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 1031
Query: 2018 VTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSE 2077
T +DW +LP AL+ R T G TP+ ++YG P V P + AK++
Sbjct: 1032 ATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYGAP---PPLVNFPDPDM---AKVTH 1084
Query: 2078 AEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQP 2137
Q+ L L++ + +A +YQ ++ D+ V P F+VG+ V RR +
Sbjct: 1085 NPSLQAHLQALYLVQHEVWRPLA--AAYQEQL----DRPVVPHPFRVGDTVWVRRHQTK- 1137
Query: 2138 DPRGKWTPNYEGPYVV 2153
P ++GPY V
Sbjct: 1138 ----NLEPRWKGPYTV 1149
>POL_MLVCB (P08361) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 282
Score = 95.5 bits (236), Expect = 1e-18
Identities = 70/237 (29%), Positives = 115/237 (47%), Gaps = 20/237 (8%)
Query: 1919 GHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVV 1978
G+ ++LV +D F+ W+EA T +VV K + I R+G+P + TDNG + V
Sbjct: 12 GYKYLLVFVDTFSGWIEAFPTKKETAKVVTKKLLEEIFPRFGMPQVLGTDNGPAFVSKVS 71
Query: 1979 QALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK--MVTTYKDWHEMLPYALHGYR 2036
Q + + I+ YRPQ +G VE N+ IK + K + T +DW +LP AL+ R
Sbjct: 72 QTVADLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTKLTLATGSRDWVLLLPLALYRAR 131
Query: 2037 TTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRM 2096
T G TP+ ++YG P V P + +++ + Q+ L L++ +
Sbjct: 132 NT-PGPHGLTPYEILYGAP---PPLVNFPDPDM---TRVTNSPSLQAHLQALYLVQHEVW 184
Query: 2097 DAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVV 2153
+A +YQ ++ D+ V P ++VG+ V RR + P ++GPY V
Sbjct: 185 RPLA--AAYQEQL----DRPVVPHPYRVGDTVWVRRHQTK-----NLEPRWKGPYTV 230
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 91.3 bits (225), Expect = 3e-17
Identities = 86/365 (23%), Positives = 150/365 (40%), Gaps = 43/365 (11%)
Query: 1780 LLSREYPPGASKQDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDG 1839
LL YPPG KQ K TL + I+ + N ++ D + H++
Sbjct: 777 LLQGHYPPGYPKQYKYTLEE-------NKLIVERPNGIRIVPPKADREKIISTAHNIAH- 828
Query: 1840 TFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVISSPW 1899
TG + + + Y+W + D + R+C +C + P L +
Sbjct: 829 ------TGRDATFLKVSSKYWWPNLRKDVVKSIRQCKQCLVTNATNLTSPPILRPVKPLK 882
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
PF + ID IG + P SNG+ +LV +D T +V Y A N++
Sbjct: 883 PFDKFYIDYIGPLPP--SNGYLHVLVVVDSMTGFVWL--YPTKAPSTSATVKALNMLTSI 938
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQK-MV 2018
+P + +D G ++ +E I+ S+PY PQ +G VE N +IKR++ K ++
Sbjct: 939 AIPKVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLI 998
Query: 2019 TTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEA 2078
W+++LP + S+ TP L++G+++ P +
Sbjct: 999 GRPAKWYDLLPVVQLALNNSYSPSSKYTPHQLLFGVDSNTPF--------------ANSD 1044
Query: 2079 EWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPD 2138
SR ++L+L++E R +Q A + P VG+LV + R+++
Sbjct: 1045 TLDLSREEELSLLQE------IRSSLHQPTSPPASSRSWSP---SVGQLV-QERVARPAS 1094
Query: 2139 PRGKW 2143
R +W
Sbjct: 1095 LRPRW 1099
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 88.6 bits (218), Expect = 2e-16
Identities = 77/256 (30%), Positives = 114/256 (44%), Gaps = 25/256 (9%)
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
P W ID ++P + G+ ++LV +D F+ WVEA T +VAK I I R+
Sbjct: 904 PGVYWEIDFT-EVKPHYA-GYKYLLVFVDTFSGWVEAFPTRQETAHIVAKKILEEIFPRF 961
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMV- 2018
G+P I +DNG + V Q L I YRPQ +G VE N+ IK + K+
Sbjct: 962 GLPKVIGSDNGPAFVSQVSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 1021
Query: 2019 -TTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSE 2077
T KDW +L AL R T + G TP+ ++YG PL + S + +
Sbjct: 1022 ETGLKDWRRLLSLALLRARNT-PNRFGLTPYEILYG--GPPPLSTLLNSF-----SPSNS 1073
Query: 2078 AEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQP 2137
Q+R L ++ + +A + + HP F+VG+ V RR Q
Sbjct: 1074 KTDLQARLKGLQAVQAQIWAPLAE------LYRPGHSQTSHP--FQVGDSVYVRRHRSQ- 1124
Query: 2138 DPRGKWTPNYEGPYVV 2153
P ++GPY+V
Sbjct: 1125 ----GLEPRWKGPYIV 1136
>POL_AVIRE (P03360) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 473
Score = 87.8 bits (216), Expect = 3e-16
Identities = 82/319 (25%), Positives = 126/319 (38%), Gaps = 20/319 (6%)
Query: 1836 VHDGTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNV- 1894
V + T R G + +L+R Y + +C C + LN
Sbjct: 123 VLEQTHRATHLGESKLTELVRKHYPICGIYRAARDITTRCVACAQVNPRAAPVEKGLNSR 182
Query: 1895 ISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNN 1954
I P W +D I K G+ ++LV +D F+ WVEA T QVV K + +
Sbjct: 183 IRGAAPGEHWEVDFTEMITAKG--GYKYLLVLVDTFSGWVEAYPAKRETSQVVIKHLILD 240
Query: 1955 IICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIV 2014
II R+G+P +I +DNG V Q LCE + YRPQ +G VE N+ +K+ +
Sbjct: 241 IIPRFGLPVQIGSDNGPAFVAKVTQQLCEALNVSWKLHCAYRPQSSGQVERMNRTLKKAI 300
Query: 2015 QKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAK 2074
K+ + + P + T G +PF ++YG++ + V L I
Sbjct: 301 AKLEDRDRRGLGLPPPSGFAPGTVYPGREGLSPFEILYGLKPPVVPRVGCDKLASITN-- 358
Query: 2075 LSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVLKRRIS 2134
Q+ L ++ R A A + +A P V +K+
Sbjct: 359 -------QTLLKSLQALQATRSLARAAARPTAPERSSARPYPTVPN--LVTSFFVKKHDF 409
Query: 2135 QQPDPRGKWTPNYEGPYVV 2153
QQ PR ++GPY V
Sbjct: 410 QQLGPR------WDGPYTV 422
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 87.0 bits (214), Expect = 5e-16
Identities = 79/320 (24%), Positives = 137/320 (42%), Gaps = 44/320 (13%)
Query: 1797 LRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFRTHATGHTMSRKLLR 1856
L+ + LLD ++ ++ ++L+ +H+ H G + ++ R
Sbjct: 763 LKLIHGCLLLDDRVIVPKSLQKIVLK---------QLHEGHPGIVQM--------KQKAR 805
Query: 1857 AGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVISSPWPF--SMWG---IDMIGR 1911
+ +W ++ D R C+ CQ + V P +PWP + W ID G
Sbjct: 806 SFVFWRGLDSDIENMVRHCNNCQENSKMPRVVP------LNPWPVPEAPWKRIHIDFAGP 859
Query: 1912 IEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGT 1971
+ NG ++LV +D TK+ E T V + I +G P II+DNGT
Sbjct: 860 L-----NGC-YLLVVVDAKTKYAEV-KLTRSISAVTTIDLLEEIFSIHGYPETIISDNGT 912
Query: 1972 NLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMVTTYKDWHEMLPYA 2031
L +++ +C+ IEH S+ Y P+ NGA E +KR + K+ ++L
Sbjct: 913 QLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAERFVDTLKRGIAKIKGEGSVNQQILNKF 972
Query: 2032 LHGYRTTVRSS-TGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNL 2090
L YR T S+ G+TP +G + + + +P+ RV+ KL++ Q N+
Sbjct: 973 LISYRNTPHSALNGSTPAECHFGRKIRTTMSLLMPTDRVLKVPKLTQY--------QQNM 1024
Query: 2091 IEEKRMDAVARGQSYQARMK 2110
+ AR +++Q K
Sbjct: 1025 KHHYELRNGARAKAFQVNQK 1044
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 85.5 bits (210), Expect = 1e-15
Identities = 56/156 (35%), Positives = 78/156 (49%), Gaps = 5/156 (3%)
Query: 1900 PFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRY 1959
P W ID ++P + G+ ++LV +D F+ WVEA T +VAK I I R+
Sbjct: 761 PGVYWEIDFT-EVKPHYA-GYKYLLVFVDTFSGWVEAYPTRQETAHMVAKKILEEIFPRF 818
Query: 1960 GVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMV- 2018
G+P I +DNG + V Q L I YRPQ +G VE N+ IK + K+
Sbjct: 819 GLPKVIGSDNGPAFVSQVSQGLARTLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 878
Query: 2019 -TTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYG 2053
T KDW +L AL R T + G TP+ ++YG
Sbjct: 879 ETGLKDWRRLLSLALLRARNT-PNRFGLTPYEILYG 913
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.340 0.148 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 232,328,298
Number of Sequences: 164201
Number of extensions: 9537673
Number of successful extensions: 36543
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 105
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 36235
Number of HSP's gapped (non-prelim): 269
length of query: 2185
length of database: 59,974,054
effective HSP length: 126
effective length of query: 2059
effective length of database: 39,284,728
effective search space: 80887254952
effective search space used: 80887254952
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 74 (33.1 bits)
Medicago: description of AC149039.3


