

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148995.8 + phase: 0
(325 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
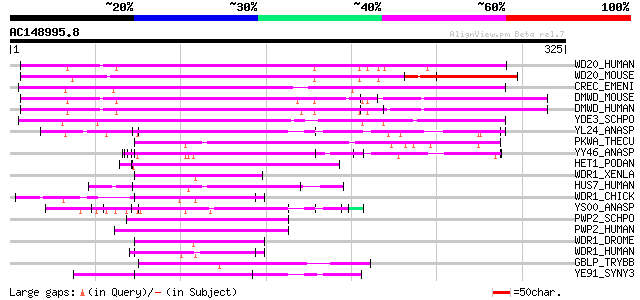
Score E
Sequences producing significant alignments: (bits) Value
WD20_HUMAN (Q8TBZ3) WD-repeat protein 20 (DMR protein) 208 2e-53
WD20_MOUSE (Q9D5R2) WD-repeat protein 20 191 2e-48
CREC_EMENI (Q9P4R5) Catabolite repression protein creC 183 4e-46
DMWD_MOUSE (Q08274) Dystrophia myotonica-containing WD repeat mo... 163 5e-40
DMWD_HUMAN (Q09019) Dystrophia myotonica-containing WD repeat mo... 163 6e-40
YDE3_SCHPO (Q10437) Hypothetical WD-repeat protein C12B10.03 in ... 158 2e-38
YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124 66 1e-10
PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkw... 62 2e-09
YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466 60 1e-08
HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1 55 2e-07
WDR1_XENLA (Q9W7F2) WD-repeat protein 1 (Actin interacting prote... 54 7e-07
HUS7_HUMAN (Q9NVX2) WD-repeat protein HUSSY-07 54 7e-07
WDR1_CHICK (O93277) WD-repeat protein 1 (Actin interacting prote... 53 1e-06
YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800 52 2e-06
PWP2_SCHPO (Q9C1X1) Periodic tryptophan protein 2 homolog 52 2e-06
PWP2_HUMAN (Q15269) Periodic tryptophan protein 2 homolog 52 2e-06
WDR1_DROME (Q9VU68) Putative actin interacting protein 1 (AIP1) 52 3e-06
WDR1_HUMAN (O75083) WD-repeat protein 1 (Actin interacting prote... 51 3e-06
GBLP_TRYBB (Q94775) Guanine nucleotide-binding protein beta subu... 51 3e-06
YE91_SYNY3 (P74598) Hypothetical WD-repeat protein sll1491 50 6e-06
>WD20_HUMAN (Q8TBZ3) WD-repeat protein 20 (DMR protein)
Length = 569
Score = 208 bits (529), Expect = 2e-53
Identities = 144/414 (34%), Positives = 188/414 (44%), Gaps = 130/414 (31%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WV G + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVHGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 509 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPG 562
>WD20_MOUSE (Q9D5R2) WD-repeat protein 20
Length = 567
Score = 191 bits (485), Expect = 2e-48
Identities = 116/290 (40%), Positives = 152/290 (52%), Gaps = 51/290 (17%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLYNKD-----GAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG D F+VAH+ GN+Y+ + G + +LK F+V + + SKS
Sbjct: 150 SRVTCVKWVPGSDSLFLVAHSSGNMYSYNVEHTYGTTAPHYQLLKQGESFAVHNCK-SKS 208
Query: 62 NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAW 121
+W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC W
Sbjct: 209 TRDLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSAELHGTMKSYFGGLLCLCW 268
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE---- 177
S DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 SPDGKYIVTGGEDDLVTVWSFLDCRVIARGRGHKSWVSVVAFDPYTTSVEESDPMEFSDS 328
Query: 178 ---------------------------------TITYRFGSVGQDTQLLLWELEMDEIV- 203
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 DKNFQDLLHFGRDRANSTQSRLSKQNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDILFP 388
Query: 204 -VPLRRGPPGGS--PTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVP 250
PL R + G P S GS + N+VP P P +P
Sbjct: 389 HQPLSRARAHANVMNATGLPAGSNGSAVTTPGNSVP----PPLPRSNSLP 434
Score = 56.2 bits (134), Expect = 1e-07
Identities = 29/67 (43%), Positives = 42/67 (62%), Gaps = 2/67 (2%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 502 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCIVTACQEGFICTWARPG 560
Query: 292 -VAESQP 297
V++ QP
Sbjct: 561 KVSKFQP 567
>CREC_EMENI (Q9P4R5) Catabolite repression protein creC
Length = 630
Score = 183 bits (465), Expect = 4e-46
Identities = 108/318 (33%), Positives = 163/318 (50%), Gaps = 41/318 (12%)
Query: 6 NSRCTCISWVPGGDGAFVVAHADGNL--YNKDGAGESSFPILKDQTLFSVAHARYS---- 59
+S T I W+PG + F+ AH +G L Y+K+ P L +Q+ SV +R+S
Sbjct: 288 SSPVTHIKWIPGSENFFIAAHENGQLVVYDKEKEDALFIPELPEQSAESVKPSRWSLQVL 347
Query: 60 --------KSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKS 111
K+NP+A W + I +FS D +LA V DG LR+ DY +E ++ +S
Sbjct: 348 KSVNSKNQKANPVAVWRLANQKITQFAFSPDHRHLAVVLEDGTLRLMDYLQEEVLDVFRS 407
Query: 112 YYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPN 171
YYGG C WS DGKYI+TGG+DDLV +WS+ +RK++A +GH+SWVS VAFD +
Sbjct: 408 YYGGFTCVCWSPDGKYIVTGGQDDLVTIWSLPERKIIARCQGHDSWVSAVAFDPW----- 462
Query: 172 SNDNGETITYRFGSVGQDTQLLLWELEM-----------------DEIVVPLRRGPPGGS 214
+ TYR GSVG D LLLW+ + ++ P + P
Sbjct: 463 ---RCDERTYRIGSVGDDCNLLLWDFSVGMLHRPKVHHQTSARHRTSLIAPSSQQPNRHR 519
Query: 215 PTFGSPTFSAGSQSSHWDN-AVPLGTLQ-PAPSMRDVPKISPLVAHRVHTEPLSSLIFTQ 272
+ SQ + D+ + P +Q P S + P+++ V +P+ L F +
Sbjct: 520 ADSSGNRMRSDSQRTAADSESAPDQPVQHPVESRARTALLPPIMSKAVGEDPICWLGFQE 579
Query: 273 ESVLTACREGHIKIWTRP 290
++++T+ EGHI+ W RP
Sbjct: 580 DTIMTSSLEGHIRTWDRP 597
>DMWD_MOUSE (Q08274) Dystrophia myotonica-containing WD repeat motif
protein (DMR-N9 protein)
Length = 650
Score = 163 bits (413), Expect = 5e-40
Identities = 99/254 (38%), Positives = 141/254 (54%), Gaps = 47/254 (18%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
++ T + W+P + F+ +HA G+LY + + + +LK F+V +A SK+
Sbjct: 196 TKVTYLKWLPESESLFLASHASGHLYLYNVSHPCTSTPPQYSLLKQGEGFAV-YAAKSKA 254
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+A+W + +G +N +FS DG +LA V +DG LRVF + L KSY+GGLLC
Sbjct: 255 PRNPLAKWAVGEGPLNEFAFSPDGRHLACVSQDGCLRVFHFDSMLLRGLMKSYFGGLLCV 314
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYW--------SSPN 171
WS DG+Y++TGGEDDLV VWS + +VVA G GH SWV+ VAFD Y +S +
Sbjct: 315 CWSPDGRYVVTGGEDDLVTVWSFTEGRVVARGHGHKSWVNAVAFDPYTTRAEEAASASAD 374
Query: 172 SNDNGE----------------------TITYRFGSVGQDTQLLLWELEMDEIVVP---L 206
+ +GE +ITYRFGS GQDTQ LW+L ++++ P L
Sbjct: 375 GDPSGEEEEPEVTSSDTGAPVSPLPKAGSITYRFGSAGQDTQFCLWDL-TEDVLSPHPSL 433
Query: 207 RR-----GPPGGSP 215
R G PG +P
Sbjct: 434 ARTRTLPGTPGATP 447
Score = 57.4 bits (137), Expect = 5e-08
Identities = 36/110 (32%), Positives = 54/110 (48%), Gaps = 3/110 (2%)
Query: 206 LRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPL 265
+ RG GG+ + + S + S D A LGT P + +VP + PLV ++ E L
Sbjct: 526 ISRGGSGGNSS--NDKLSGPAPRSRLDPAKVLGTAL-CPRIHEVPLLEPLVCKKIAQERL 582
Query: 266 SSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLS 315
+ L+F ++ ++TAC+EG I W RPG A + S PS S
Sbjct: 583 TVLLFLEDCIITACQEGLICTWARPGKAFTDEETEAQAGQASWPRSPSKS 632
>DMWD_HUMAN (Q09019) Dystrophia myotonica-containing WD repeat motif
protein (DMR-N9 protein) (Protein 59) (Fragment)
Length = 553
Score = 163 bits (412), Expect = 6e-40
Identities = 100/263 (38%), Positives = 138/263 (52%), Gaps = 51/263 (19%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
++ T + W+P + F+ +HA G+LY + + + +LK FSV +A SK+
Sbjct: 92 TKVTYLKWLPESESLFLASHASGHLYLYNVSHPCASAPPQYSLLKQGEGFSV-YAAKSKA 150
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+A+W + +G +N +FS DG +LA V +D LRVF + L KSY+GGLLC
Sbjct: 151 PRNPLAKWAVGEGPLNEFAFSPDGRHLACVSQDACLRVFHFDSMLLRGLMKSYFGGLLCV 210
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSS---------- 169
WS DG+Y++TGGEDDLV VWS + +VVA G GH SWV+ VAFDS +++
Sbjct: 211 CWSPDGRYVVTGGEDDLVTVWSFTEGRVVARGHGHKSWVNAVAFDSLYTTRAEEAATAAG 270
Query: 170 -------------PNSNDNGE-------------TITYRFGSVGQDTQLLLWELEMDEIV 203
P + G +ITYRFGS GQDTQ LW+L D +
Sbjct: 271 ADGERSGEEEEEEPEAAGTGSAGGAPLSPLPKAGSITYRFGSAGQDTQFCLWDLTEDVLY 330
Query: 204 --VPLRR-----GPPGGSPTFGS 219
PL R G PG +P S
Sbjct: 331 PHPPLARTRTLPGTPGTTPPAAS 353
Score = 57.0 bits (136), Expect = 6e-08
Identities = 38/110 (34%), Positives = 53/110 (47%), Gaps = 2/110 (1%)
Query: 206 LRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPL 265
+ RG GGS + G S S D A LGT P + +VP + PLV ++ E L
Sbjct: 428 ISRGGSGGSGSGGEKP-SGPVPRSRLDPAKVLGTAL-CPRIHEVPLLEPLVCKKIAQERL 485
Query: 266 SSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLS 315
+ L+F ++ ++TAC+EG I W RPG A + S PS S
Sbjct: 486 TVLLFLEDCIITACQEGLICTWPRPGKAFTDEETEAQTGEGSWPRSPSKS 535
>YDE3_SCHPO (Q10437) Hypothetical WD-repeat protein C12B10.03 in
chromosome I
Length = 543
Score = 158 bits (399), Expect = 2e-38
Identities = 104/313 (33%), Positives = 150/313 (47%), Gaps = 38/313 (12%)
Query: 6 NSRCTCISWVPGGDGAFVVAHADG--NLYNKDGAGESSFPILKDQTL------------- 50
+S T I WV G D F+V+ +G LY+K + ++ ++ L
Sbjct: 241 SSSVTAIKWVAGKDSQFLVSFRNGWLVLYDKYRHEQPLHIVVPEKNLKSLYLSSPGTFNI 300
Query: 51 -FSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGG 109
S+ H K NP+A + + IN FS D YLA V G L++FD+ KEH++
Sbjct: 301 LISINHRDDRKLNPVACYAFSKSPINGFCFSPDYQYLALVSERGTLKLFDFVKEHVLDVF 360
Query: 110 KSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSS 169
SY+ GL C WS DGK+I GG+DDLV ++S RK+VA +GH SWV+ V FD+ W
Sbjct: 361 HSYFAGLTCVTWSPDGKFIAIGGKDDLVSIYSFPLRKLVARCQGHKSWVTDVIFDA-WRC 419
Query: 170 PNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVP------LRRGPPGGSPTF------ 217
+ N YR SVG D +LLLW+ + I P + P
Sbjct: 420 DDDN-------YRIASVGLDRKLLLWDFSVSAIHRPKSAVYYVNHHSNNSKPAISDFDDV 472
Query: 218 GSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLT 277
G T + +S++ N T+ P S +P ISP+ + V PLSS+ F + ++T
Sbjct: 473 GDLTMGSEIDNSNYVNGDI--TIHPTLSRSLIPVISPITIYDVDDSPLSSVFFDPDCMIT 530
Query: 278 ACREGHIKIWTRP 290
G I+ W RP
Sbjct: 531 CATNGRIRTWQRP 543
>YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124
Length = 1683
Score = 65.9 bits (159), Expect = 1e-10
Identities = 52/223 (23%), Positives = 100/223 (44%), Gaps = 32/223 (14%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
++N++ FS DG LA+ D ++++D T L+ + G++ +S DG+ I G
Sbjct: 1157 TVNNVYFSPDGKNLASASSDHSIKLWDTTSGQLLMTLTGHSAGVITVRFSPDGQTIAAGS 1216
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQL 192
ED V++W +D K++ GH WV+ ++F + +G+T+ S D +
Sbjct: 1217 EDKTVKLWHRQDGKLLKTLNGHQDWVNSLSF---------SPDGKTL----ASASADKTI 1263
Query: 193 LLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQ---SSHWDNAVPLGTLQPAPSMRDV 249
LW + ++V L+ G + + FS+ + S+ DN + L
Sbjct: 1264 KLWRIADGKLVKTLK----GHNDSVWDVNFSSDGKAIASASRDNTIKLWNRHGI------ 1313
Query: 250 PKISPLVAHRVHTEPLSSLIFTQES--VLTACREGHIKIWTRP 290
L H+ + ++ F +S + +A + I++W RP
Sbjct: 1314 ----ELETFTGHSGGVYAVNFLPDSNIIASASLDNTIRLWQRP 1352
Score = 43.9 bits (102), Expect = 5e-04
Identities = 38/183 (20%), Positives = 78/183 (41%), Gaps = 24/183 (13%)
Query: 19 DGAFVV-AHADGNL---YNKDGAGESSFPILKDQTLFSVAH-------ARYSKSNPIARW 67
DG+ + A ADGN+ +++DG+ + P ++ ++ ++ A + + W
Sbjct: 1374 DGSIIATAGADGNIQLWHSQDGSLLKTLP--GNKAIYGISFTPQGDLIASANADKTVKIW 1431
Query: 68 HICQGS-----------INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGL 116
+ G +N ++FS DG LA+ RD +++++ + K + +
Sbjct: 1432 RVRDGKALKTLIGHDNEVNKVNFSPDGKTLASASRDNTVKLWNVSDGKFKKTLKGHTDEV 1491
Query: 117 LCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNG 176
++S DGK I + D +++W ++ HN V V F+ S S
Sbjct: 1492 FWVSFSPDGKIIASASADKTIRLWDSFSGNLIKSLPAHNDLVYSVNFNPDGSMLASTSAD 1551
Query: 177 ETI 179
+T+
Sbjct: 1552 KTV 1554
Score = 43.5 bits (101), Expect = 7e-04
Identities = 49/217 (22%), Positives = 90/217 (40%), Gaps = 32/217 (14%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDD 135
SIS S DG +A+ D ++++ L + + ++S DG+ I +GG D
Sbjct: 1077 SISISRDGQTIASGSLDKTIKLWS-RDGRLFRTLNGHEDAVYSVSFSPDGQTIASGGSDK 1135
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLW 195
+++W D ++ GH V+ V F SP+ + S D + LW
Sbjct: 1136 TIKLWQTSDGTLLKTITGHEQTVNNVYF-----SPDGKN--------LASASSDHSIKLW 1182
Query: 196 ELEMDEIVVPLRRGPPGGSPTFGSP---TFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKI 252
+ ++++ L G SP T +AGS+ D V L Q ++ +
Sbjct: 1183 DTTSGQLLMTLTGHSAGVITVRFSPDGQTIAAGSE----DKTVKLWHRQDGKLLKTL--- 1235
Query: 253 SPLVAHRVHTEPLSSLIFTQE--SVLTACREGHIKIW 287
H + ++SL F+ + ++ +A + IK+W
Sbjct: 1236 ------NGHQDWVNSLSFSPDGKTLASASADKTIKLW 1266
Score = 42.0 bits (97), Expect = 0.002
Identities = 33/126 (26%), Positives = 56/126 (44%), Gaps = 14/126 (11%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
Q +NS+SFS DG LA+ D ++++ LV K + + +S DGK I +
Sbjct: 1239 QDWVNSLSFSPDGKTLASASADKTIKLWRIADGKLVKTLKGHNDSVWDVNFSSDGKAIAS 1298
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
D+ +++W+ ++ + GH+ V V F P+SN S D
Sbjct: 1299 ASRDNTIKLWNRHGIELETF-TGHSGGVYAVNF-----LPDSN--------IIASASLDN 1344
Query: 191 QLLLWE 196
+ LW+
Sbjct: 1345 TIRLWQ 1350
Score = 41.2 bits (95), Expect = 0.004
Identities = 30/154 (19%), Positives = 63/154 (40%), Gaps = 24/154 (15%)
Query: 56 ARYSKSNPIARWHICQGSINS-----------ISFSADGAYLATVGRDGYLRVFDYTKEH 104
A S+ N + W++ G +SFS DG +A+ D +R++D +
Sbjct: 1462 ASASRDNTVKLWNVSDGKFKKTLKGHTDEVFWVSFSPDGKIIASASADKTIRLWDSFSGN 1521
Query: 105 LVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
L+ ++ + ++ DG + + D V++W D ++ GH++ V +F
Sbjct: 1522 LIKSLPAHNDLVYSVNFNPDGSMLASTSADKTVKLWRSHDGHLLHTFSGHSNVVYSSSF- 1580
Query: 165 SYWSSPNSNDNGETITYRFGSVGQDTQLLLWELE 198
SP+ S +D + +W+++
Sbjct: 1581 ----SPDGR--------YIASASEDKTVKIWQID 1602
Score = 41.2 bits (95), Expect = 0.004
Identities = 19/73 (26%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ S SFS DG Y+A+ D ++++ HL+ + G++ +S DGK +++G
Sbjct: 1575 VYSSSFSPDGRYIASASEDKTVKIWQIDG-HLLTTLPQHQAGVMSAIFSPDGKTLISGSL 1633
Query: 134 DDLVQVWSMEDRK 146
D ++W + ++
Sbjct: 1634 DTTTKIWRFDSQQ 1646
Score = 33.1 bits (74), Expect = 0.97
Identities = 23/94 (24%), Positives = 43/94 (45%), Gaps = 6/94 (6%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFD--YTKEHLVCGGKSYYGGLLCCAWSMDGKYIL 129
G + +++F D +A+ D +R++ V G S G+ ++ DG I
Sbjct: 1323 GGVYAVNFLPDSNIIASASLDNTIRLWQRPLISPLEVLAGNS---GVYAVSFLHDGSIIA 1379
Query: 130 TGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
T G D +Q+W +D ++ G N + G++F
Sbjct: 1380 TAGADGNIQLWHSQDGSLLKTLPG-NKAIYGISF 1412
Score = 32.3 bits (72), Expect = 1.7
Identities = 23/82 (28%), Positives = 39/82 (47%), Gaps = 14/82 (17%)
Query: 115 GLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSND 174
G++ + S DG+ I +G D +++WS D ++ GH V V+F +
Sbjct: 1074 GVISISISRDGQTIASGSLDKTIKLWS-RDGRLFRTLNGHEDAVYSVSF---------SP 1123
Query: 175 NGETITYRFGSVGQDTQLLLWE 196
+G+TI S G D + LW+
Sbjct: 1124 DGQTI----ASGGSDKTIKLWQ 1141
>PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkwA
(EC 2.7.1.37)
Length = 742
Score = 62.0 bits (149), Expect = 2e-09
Identities = 66/239 (27%), Positives = 103/239 (42%), Gaps = 30/239 (12%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYT--KEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
+ +++FS DGA LA+ D +R++D +E V G ++Y +L A+S DG + +G
Sbjct: 504 VRAVAFSPDGALLASGSDDATVRLWDVAAAEERAVFEGHTHY--VLDIAFSPDGSMVASG 561
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D ++W++ A +GH +V VAF S S TI + G++
Sbjct: 562 SRDGTARLWNVATGTEHAVLKGHTDYVYAVAFSPDGSMVASGSRDGTIRLWDVATGKERD 621
Query: 192 LLLWELEMDEIVVPLRRGPPGGSPTFGSPTF-----SAGSQSSH-------WDNAV---P 236
+L E VV L P G GS + A ++ H W AV P
Sbjct: 622 VLQAPAEN---VVSLAFSPDGSMLVHGSDSTVHLWDVASGEALHTFEGHTDWVRAVAFSP 678
Query: 237 LGTLQPAPS------MRDVPKISPLVAHRVHTEPLSSLIFTQE--SVLTACREGHIKIW 287
G L + S + DV HTEP+ S+ F E ++ +A +G I+IW
Sbjct: 679 DGALLASGSDDRTIRLWDVAAQEEHTTLEGHTEPVHSVAFHPEGTTLASASEDGTIRIW 737
Score = 33.9 bits (76), Expect = 0.57
Identities = 16/44 (36%), Positives = 21/44 (47%)
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
A+S G + G D L+ VW + + EGH WV VAF
Sbjct: 466 AFSPGGSLLAGGSGDKLIHVWDVASGDELHTLEGHTDWVRAVAF 509
>YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466
Length = 1526
Score = 59.7 bits (143), Expect = 1e-08
Identities = 52/229 (22%), Positives = 96/229 (41%), Gaps = 34/229 (14%)
Query: 68 HICQGS---INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMD 124
+I QG +NS+ F+ DG+ LA+ D +R+++ +C + + + ++ D
Sbjct: 1194 YILQGHTSWVNSVVFNPDGSTLASGSSDQTVRLWEINSSKCLCTFQGHTSWVNSVVFNPD 1253
Query: 125 GKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFG 184
G + +G D V++W + K + +GH +WV+ VAF N +G + G
Sbjct: 1254 GSMLASGSSDKTVRLWDISSSKCLHTFQGHTNWVNSVAF---------NPDGSMLASGSG 1304
Query: 185 SVGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGS---QSSHWDNAVPLGTLQ 241
D + LWE+ + + + G + S TFS S D V L ++
Sbjct: 1305 ----DQTVRLWEISSSKCLHTFQ----GHTSWVSSVTFSPDGTMLASGSDDQTVRLWSIS 1356
Query: 242 PAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGH--IKIWT 288
L HT + S+IF+ + + A G +++W+
Sbjct: 1357 SGEC---------LYTFLGHTNWVGSVIFSPDGAILASGSGDQTVRLWS 1396
Score = 58.2 bits (139), Expect = 3e-08
Identities = 29/106 (27%), Positives = 56/106 (52%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ S+ FS DGA LA+ G D +R++D + + + + Y + +S +G + G
Sbjct: 1077 VRSVVFSPDGAMLASGGDDQIVRLWDISSGNCLYTLQGYTSWVRFLVFSPNGVTLANGSS 1136
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
D +V++W + +K + +GH +WV+ VAF ++ S +T+
Sbjct: 1137 DQIVRLWDISSKKCLYTLQGHTNWVNAVAFSPDGATLASGSGDQTV 1182
Score = 53.9 bits (128), Expect = 5e-07
Identities = 38/132 (28%), Positives = 61/132 (45%), Gaps = 17/132 (12%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYT--KEHLVCGGKSYYGGLLCCAWSMDGKYIL 129
GS+ +++FS DG AT G +R ++ KE L C G + + + +S DGK +
Sbjct: 865 GSVLTVAFSPDGKLFATGDSGGIVRFWEAATGKELLTCKGHNSWVNSV--GFSQDGKMLA 922
Query: 130 TGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQD 189
+G +D V++W + + + +GH S V V F SPNS S D
Sbjct: 923 SGSDDQTVRLWDISSGQCLKTFKGHTSRVRSVVF-----SPNS--------LMLASGSSD 969
Query: 190 TQLLLWELEMDE 201
+ LW++ E
Sbjct: 970 QTVRLWDISSGE 981
Score = 53.1 bits (126), Expect = 9e-07
Identities = 45/216 (20%), Positives = 89/216 (40%), Gaps = 25/216 (11%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ + FS +G LA D +R++D + + + + + + A+S DG + +G
Sbjct: 1119 VRFLVFSPNGVTLANGSSDQIVRLWDISSKKCLYTLQGHTNWVNAVAFSPDGATLASGSG 1178
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLL 193
D V++W + K + +GH SWV+ V F N +G T+ S D +
Sbjct: 1179 DQTVRLWDISSSKCLYILQGHTSWVNSVVF---------NPDGSTL----ASGSSDQTVR 1225
Query: 194 LWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKIS 253
LWE+ + + + G + S F+ + + G+ + D+
Sbjct: 1226 LWEINSSKCLCTFQ----GHTSWVNSVVFNPDG------SMLASGSSDKTVRLWDISSSK 1275
Query: 254 PLVAHRVHTEPLSSLIFTQESVLTACREGH--IKIW 287
L + HT ++S+ F + + A G +++W
Sbjct: 1276 CLHTFQGHTNWVNSVAFNPDGSMLASGSGDQTVRLW 1311
Score = 52.4 bits (124), Expect = 2e-06
Identities = 37/144 (25%), Positives = 71/144 (48%), Gaps = 16/144 (11%)
Query: 67 WHICQGS---INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSM 123
++I QG + S+ FS+DGA LA+ D +R++D + + + + + + +S
Sbjct: 1025 FYIFQGHTSCVRSVVFSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVVFSP 1084
Query: 124 DGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRF 183
DG + +GG+D +V++W + + +G+ SWV + F SPN +T
Sbjct: 1085 DGAMLASGGDDQIVRLWDISSGNCLYTLQGYTSWVRFLVF-----SPNG------VTLAN 1133
Query: 184 GSVGQDTQLLLWELEMDEIVVPLR 207
GS D + LW++ + + L+
Sbjct: 1134 GS--SDQIVRLWDISSKKCLYTLQ 1155
Score = 50.8 bits (120), Expect = 4e-06
Identities = 31/118 (26%), Positives = 59/118 (49%), Gaps = 9/118 (7%)
Query: 68 HICQGS---INSISFSADGAYLATVGRDGYLRVFDYTKEHLV---CGGKSYYGGLLCCAW 121
H QG ++S++FS DG LA+ D +R++ + + G ++ G ++ +
Sbjct: 1320 HTFQGHTSWVSSVTFSPDGTMLASGSDDQTVRLWSISSGECLYTFLGHTNWVGSVI---F 1376
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
S DG + +G D V++WS+ K + +GHN+WV + F + S + +T+
Sbjct: 1377 SPDGAILASGSGDQTVRLWSISSGKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQTV 1434
Score = 48.5 bits (114), Expect = 2e-05
Identities = 28/113 (24%), Positives = 53/113 (46%), Gaps = 3/113 (2%)
Query: 70 CQGS---INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGK 126
C+G +NS+ FS DG LA+ D +R++D + + K + + +S +
Sbjct: 902 CKGHNSWVNSVGFSQDGKMLASGSDDQTVRLWDISSGQCLKTFKGHTSRVRSVVFSPNSL 961
Query: 127 YILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
+ +G D V++W + + + +GH WV VAF+ S + +T+
Sbjct: 962 MLASGSSDQTVRLWDISSGECLYIFQGHTGWVYSVAFNLDGSMLATGSGDQTV 1014
Score = 47.0 bits (110), Expect = 6e-05
Identities = 26/106 (24%), Positives = 51/106 (47%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ S+ FS + LA+ D +R++D + + + + G + A+++DG + TG
Sbjct: 951 VRSVVFSPNSLMLASGSSDQTVRLWDISSGECLYIFQGHTGWVYSVAFNLDGSMLATGSG 1010
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
D V++W + + +GH S V V F S + S + +T+
Sbjct: 1011 DQTVRLWDISSSQCFYIFQGHTSCVRSVVFSSDGAMLASGSDDQTV 1056
Score = 46.2 bits (108), Expect = 1e-04
Identities = 32/136 (23%), Positives = 59/136 (42%), Gaps = 13/136 (9%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
G + S++F+ DG+ LAT D +R++D + + + + +S DG + +G
Sbjct: 991 GWVYSVAFNLDGSMLATGSGDQTVRLWDISSSQCFYIFQGHTSCVRSVVFSSDGAMLASG 1050
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
+D V++W + + +GH S V V F SP+ S G D
Sbjct: 1051 SDDQTVRLWDISSGNCLYTLQGHTSCVRSVVF-----SPDG--------AMLASGGDDQI 1097
Query: 192 LLLWELEMDEIVVPLR 207
+ LW++ + L+
Sbjct: 1098 VRLWDISSGNCLYTLQ 1113
Score = 44.7 bits (104), Expect = 3e-04
Identities = 34/137 (24%), Positives = 64/137 (45%), Gaps = 19/137 (13%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKE---HLVCGGKSYYGGLLCCAWSMDGKYILT 130
+ S+ FS DGA LA+ D +R++ + + + G ++ G ++ +S DG + +
Sbjct: 1371 VGSVIFSPDGAILASGSGDQTVRLWSISSGKCLYTLQGHNNWVGSIV---FSPDGTLLAS 1427
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
G +D V++W++ + + GH + V VAF S S + ETI
Sbjct: 1428 GSDDQTVRLWNISSGECLYTLHGHINSVRSVAFSSDGLILASGSDDETIK---------- 1477
Query: 191 QLLLWELEMDEIVVPLR 207
LW+++ E + L+
Sbjct: 1478 ---LWDVKTGECIKTLK 1491
>HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1
Length = 1356
Score = 55.5 bits (132), Expect = 2e-07
Identities = 33/122 (27%), Positives = 55/122 (45%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
G ++S++FS DG +A+ DG ++++D + + G + A+S DG+ + +G
Sbjct: 1136 GWVHSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQTLEGHGGWVQSVAFSPDGQRVASG 1195
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D +++W EGH WV VAF S + TI + G TQ
Sbjct: 1196 SSDKTIKIWDTASGTCTQTLEGHGGWVQSVAFSPDGQRVASGSSDNTIKIWDTASGTCTQ 1255
Query: 192 LL 193
L
Sbjct: 1256 TL 1257
Score = 51.2 bits (121), Expect = 3e-06
Identities = 31/121 (25%), Positives = 52/121 (42%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S+ S++FS DG +A+ D ++++D + + G + A+S DG+ + +G
Sbjct: 969 SVLSVAFSPDGQRVASGSGDKTIKIWDTASGTCTQTLEGHGGSVWSVAFSPDGQRVASGS 1028
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQL 192
+D +++W EGH WV V F S + TI G TQ
Sbjct: 1029 DDKTIKIWDTASGTCTQTLEGHGGWVQSVVFSPDGQRVASGSDDHTIKIWDAVSGTCTQT 1088
Query: 193 L 193
L
Sbjct: 1089 L 1089
Score = 48.1 bits (113), Expect = 3e-05
Identities = 34/136 (25%), Positives = 57/136 (41%), Gaps = 7/136 (5%)
Query: 65 ARWHICQ-------GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLL 117
A W+ C S+ S++FSADG +A+ D ++++D + + G +
Sbjct: 828 AEWNACTQTLEGHGSSVLSVAFSADGQRVASGSDDKTIKIWDTASGTGTQTLEGHGGSVW 887
Query: 118 CCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE 177
A+S D + + +G +D +++W EGH V VAF S +
Sbjct: 888 SVAFSPDRERVASGSDDKTIKIWDAASGTCTQTLEGHGGRVQSVAFSPDGQRVASGSDDH 947
Query: 178 TITYRFGSVGQDTQLL 193
TI + G TQ L
Sbjct: 948 TIKIWDAASGTCTQTL 963
Score = 46.6 bits (109), Expect = 8e-05
Identities = 32/122 (26%), Positives = 54/122 (44%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
GS+ S++FS D +A+ D ++++D + + G + A+S DG+ + +G
Sbjct: 884 GSVWSVAFSPDRERVASGSDDKTIKIWDAASGTCTQTLEGHGGRVQSVAFSPDGQRVASG 943
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
+D +++W EGH S V VAF S +TI + G TQ
Sbjct: 944 SDDHTIKIWDAASGTCTQTLEGHGSSVLSVAFSPDGQRVASGSGDKTIKIWDTASGTCTQ 1003
Query: 192 LL 193
L
Sbjct: 1004 TL 1005
Score = 46.2 bits (108), Expect = 1e-04
Identities = 31/122 (25%), Positives = 50/122 (40%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
G + S+ FS DG +A+ D ++++D + + + A+S DG+ + +G
Sbjct: 1052 GWVQSVVFSPDGQRVASGSDDHTIKIWDAVSGTCTQTLEGHGDSVWSVAFSPDGQRVASG 1111
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D +++W EGH WV VAF S TI + G TQ
Sbjct: 1112 SIDGTIKIWDAASGTCTQTLEGHGGWVHSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQ 1171
Query: 192 LL 193
L
Sbjct: 1172 TL 1173
Score = 44.3 bits (103), Expect = 4e-04
Identities = 31/122 (25%), Positives = 52/122 (42%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
GS+ S++FS DG +A+ D ++++D + + G + +S DG+ + +G
Sbjct: 1010 GSVWSVAFSPDGQRVASGSDDKTIKIWDTASGTCTQTLEGHGGWVQSVVFSPDGQRVASG 1069
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
+D +++W EGH V VAF S TI + G TQ
Sbjct: 1070 SDDHTIKIWDAVSGTCTQTLEGHGDSVWSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQ 1129
Query: 192 LL 193
L
Sbjct: 1130 TL 1131
>WDR1_XENLA (Q9W7F2) WD-repeat protein 1 (Actin interacting protein
1) (XAIP1)
Length = 608
Score = 53.5 bits (127), Expect = 7e-07
Identities = 25/78 (32%), Positives = 41/78 (52%), Gaps = 3/78 (3%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVC---GGKSYYGGLLCCAWSMDGKYILT 130
+N + FS DG+ LA+ G DG + ++D VC G K++ GG+ +WS DG +L+
Sbjct: 192 VNCVRFSPDGSKLASAGADGQIFLYDGKTGEKVCSLGGSKAHDGGIYAVSWSPDGTQLLS 251
Query: 131 GGEDDLVQVWSMEDRKVV 148
D ++W + V
Sbjct: 252 ASGDKTTKIWDVAANSAV 269
Score = 35.0 bits (79), Expect = 0.26
Identities = 32/118 (27%), Positives = 49/118 (41%), Gaps = 25/118 (21%)
Query: 18 GDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWHICQGSINSI 77
G G V ADG ++ G S LKD+ K+ P +G++ +
Sbjct: 456 GSGTVAVGGADGKVHLYSIQGNS----LKDE----------GKTLP------AKGAVTDL 495
Query: 78 SFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYG---GLLCCAWSMDGKYILTGG 132
++S DGA+LA + + VF + SYYG L AWS D ++ + G
Sbjct: 496 AYSHDGAFLAVTDANKVVTVFSVADGY--SEKNSYYGHHAKALSVAWSPDNEHFASSG 551
Score = 31.2 bits (69), Expect = 3.7
Identities = 20/57 (35%), Positives = 31/57 (54%), Gaps = 2/57 (3%)
Query: 79 FSADGAYLATVGRDGYLRVFDYT-KEHLV-CGGKSYYGGLLCCAWSMDGKYILTGGE 133
++ G Y+A+ G LR++D T KEHL+ + + G + AW+ D K I GE
Sbjct: 66 YAPSGFYIASGDTSGKLRIWDTTQKEHLLKYEYQPFAGKIKDIAWTEDSKRIAVVGE 122
>HUS7_HUMAN (Q9NVX2) WD-repeat protein HUSSY-07
Length = 485
Score = 53.5 bits (127), Expect = 7e-07
Identities = 29/101 (28%), Positives = 56/101 (54%), Gaps = 5/101 (4%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKE--HLVCGGKSYYGGLLCCAWSMDGKYILT 130
++ S++FS G YLA+ D +R +D + E H C G ++ +L +WS DGK + +
Sbjct: 116 AVISVAFSPTGKYLASGSGDTTVRFWDLSTETPHFTCKGHRHW--VLSISWSPDGKKLAS 173
Query: 131 GGEDDLVQVWSMEDRKVVAWG-EGHNSWVSGVAFDSYWSSP 170
G ++ + +W K V GH+ W++G++++ ++P
Sbjct: 174 GCKNGQILLWDPSTGKQVGRTLAGHSKWITGLSWEPLHANP 214
Score = 48.5 bits (114), Expect = 2e-05
Identities = 38/149 (25%), Positives = 63/149 (41%), Gaps = 15/149 (10%)
Query: 47 DQTLFSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLV 106
D TLF + A K P+ R Q IN + FS D +A+ D ++++D +
Sbjct: 350 DFTLFLWSPAEDKK--PLTRMTGHQALINQVLFSPDSRIVASASFDKSIKLWDGRTGKYL 407
Query: 107 CGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSY 166
+ + + AWS D + +++G D ++VW ++ +K+ GH V V
Sbjct: 408 ASLRGHVAAVYQIAWSADSRLLVSGSSDSTLKVWDVKAQKLAMDLPGHADEVYAVD---- 463
Query: 167 WSSPNSNDNGETITYRFGSVGQDTQLLLW 195
WS R S G+D L +W
Sbjct: 464 WSPDGQ---------RVASGGKDKCLRIW 483
Score = 40.0 bits (92), Expect = 0.008
Identities = 22/84 (26%), Positives = 42/84 (49%), Gaps = 7/84 (8%)
Query: 85 YLATVGRDGYLRVFDYTK---EHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWS 141
Y+A+ +DG +R++D T E ++ G + + C W DG + + +D ++VW
Sbjct: 218 YVASSSKDGSVRIWDTTAGRCERILTG---HTQSVTCLRWGGDG-LLYSASQDRTIKVWR 273
Query: 142 MEDRKVVAWGEGHNSWVSGVAFDS 165
D + +GH WV+ +A +
Sbjct: 274 AHDGVLCRTLQGHGHWVNTMALST 297
>WDR1_CHICK (O93277) WD-repeat protein 1 (Actin interacting protein
1) (AIP1)
Length = 609
Score = 52.8 bits (125), Expect = 1e-06
Identities = 25/79 (31%), Positives = 40/79 (49%), Gaps = 3/79 (3%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVC---GGKSYYGGLLCCAWSMDGKYILT 130
+N + FS DG AT DG + ++D VC GGK++ GG+ +WS D +L+
Sbjct: 194 VNCVRFSPDGNRFATASADGQIFIYDGKTGEKVCALGGGKAHDGGIYAISWSPDSSQLLS 253
Query: 131 GGEDDLVQVWSMEDRKVVA 149
D ++W + VV+
Sbjct: 254 ASGDKTAKIWDVGANSVVS 272
Score = 51.6 bits (122), Expect = 3e-06
Identities = 43/146 (29%), Positives = 67/146 (45%), Gaps = 30/146 (20%)
Query: 4 DLNSRCTCISWVPGGDGAFVVAHADGN--LYNKDGAGESSFPILKDQTLFSVAHARYSKS 61
DL ++ PGG G+ V DGN LY+ G S D+TL +
Sbjct: 445 DLGYEPEAVAVHPGG-GSVAVGGTDGNVRLYSIQGTSLKS----DDKTLEA--------- 490
Query: 62 NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVF---DYTKEHLVCGGKSYYGGLLC 118
+G + +++S DGA+LA + + VF D EH V G ++ ++C
Sbjct: 491 ---------KGPVTDLAYSHDGAFLAVCDANKVVTVFSVPDGYVEHNVFYG--HHAKVVC 539
Query: 119 CAWSMDGKYILTGGEDDLVQVWSMED 144
AWS D ++ +GG D +V VW++ D
Sbjct: 540 IAWSPDNEHFASGGMDMMVYVWTVSD 565
Score = 33.1 bits (74), Expect = 0.97
Identities = 30/104 (28%), Positives = 48/104 (45%), Gaps = 7/104 (6%)
Query: 79 FSADGAYLATVGRDGYLRVFDYT-KEHLV-CGGKSYYGGLLCCAWSMDGKYILTGGE--D 134
++ G Y+A+ G LR++D T KEHL+ + + G + AW+ D K I GE +
Sbjct: 68 YAPSGFYIASGDVSGKLRIWDTTQKEHLLKYEYQPFAGKIKDLAWTEDSKRIAVVGEGRE 127
Query: 135 DLVQVWSMEDRKVVAWGEGHNSWVSGVAFDS---YWSSPNSNDN 175
V+ + V GHN ++ V Y + S+DN
Sbjct: 128 KFGAVFLWDSGSSVGEITGHNKVINSVDIKQTRPYRLATGSDDN 171
>YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800
Length = 1258
Score = 52.4 bits (124), Expect = 2e-06
Identities = 39/129 (30%), Positives = 65/129 (50%), Gaps = 17/129 (13%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYT--KEHLVCGGKSYYGGLLCCAWSMDGKYIL 129
G+I S +FS +G LAT D ++RV++ K L+C G S + + +S DG+ +
Sbjct: 643 GNILSAAFSPEGQLLATCDTDCHVRVWEVKSGKLLLICRGHSNWVRFV--VFSPDGEILA 700
Query: 130 TGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQD 189
+ G D+ V++WS+ D + GH V VAF + +GET+ S D
Sbjct: 701 SCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAF---------HPDGETL----ASASGD 747
Query: 190 TQLLLWELE 198
+ LW+++
Sbjct: 748 KTIKLWDIQ 756
Score = 51.2 bits (121), Expect = 3e-06
Identities = 44/167 (26%), Positives = 78/167 (46%), Gaps = 24/167 (14%)
Query: 49 TLFSVAHARYSK-------SNPIARW----HICQGSIN-------SISFSADGAYLATVG 90
+++S+A++ SK I W HIC +++ S++FS DG LA V
Sbjct: 854 SVYSIAYSPDSKILVSGSGDRTIKLWDCQTHICIKTLHGHTNEVCSVAFSPDGQTLACVS 913
Query: 91 RDGYLRVFDYTKEHLVCGGKSYYGGL---LCCAWSMDGKYILTGGEDDLVQVWSMEDRKV 147
D +R+++ + K++YG L A+S D + + +G D V++W + K
Sbjct: 914 LDQSVRLWNCRTGQCL---KAWYGNTDWALPVAFSPDRQILASGSNDKTVKLWDWQTGKY 970
Query: 148 VAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLL 194
++ EGH ++ G+AF + S ++ S GQ Q+LL
Sbjct: 971 ISSLEGHTDFIYGIAFSPDSQTLASASTDSSVRLWNISTGQCFQILL 1017
Score = 46.2 bits (108), Expect = 1e-04
Identities = 38/163 (23%), Positives = 65/163 (39%), Gaps = 24/163 (14%)
Query: 56 ARYSKSNPIARWHICQGS-----------INSISFSADGAYLATVGRDGYLRVFDYTKEH 104
A S + W C G + S FS +G +AT D ++++D+ +
Sbjct: 1078 ASASADQSVRLWDCCTGRCVGILRGHSNRVYSAIFSPNGEIIATCSTDQTVKIWDWQQGK 1137
Query: 105 LVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
+ + + A+S DGK + + D V++W + K GH VS VAF
Sbjct: 1138 CLKTLTGHTNWVFDIAFSPDGKILASASHDQTVRIWDVNTGKCHHICIGHTHLVSSVAF- 1196
Query: 165 SYWSSPNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPLR 207
+ +GE + S QD + +W ++ E + LR
Sbjct: 1197 --------SPDGEVV----ASGSQDQTVRIWNVKTGECLQILR 1227
Score = 45.1 bits (105), Expect = 2e-04
Identities = 25/104 (24%), Positives = 46/104 (44%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDD 135
S++F DG LA+ D ++++D + + + C A+S DG + + D
Sbjct: 731 SVAFHPDGETLASASGDKTIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADH 790
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
+++W + K + + H WV VAF + + S TI
Sbjct: 791 TIKLWDVSQGKCLRTLKSHTGWVRSVAFSADGQTLASGSGDRTI 834
Score = 43.9 bits (102), Expect = 5e-04
Identities = 42/172 (24%), Positives = 73/172 (42%), Gaps = 30/172 (17%)
Query: 22 FVVAHADGNLYNKDGAGES-------SFPILKDQT-----LFSVAH-------ARYSKSN 62
FVV DG + GA E+ +K T +FSVA A S
Sbjct: 689 FVVFSPDGEILASCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAFHPDGETLASASGDK 748
Query: 63 PIARWHICQGS-----------INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKS 111
I W I G+ + ++FS DG LA+ D ++++D ++ + KS
Sbjct: 749 TIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLRTLKS 808
Query: 112 YYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
+ G + A+S DG+ + +G D +++W+ + + GH + V +A+
Sbjct: 809 HTGWVRSVAFSADGQTLASGSGDRTIKIWNYHTGECLKTYIGHTNSVYSIAY 860
Score = 34.7 bits (78), Expect = 0.33
Identities = 31/162 (19%), Positives = 63/162 (38%), Gaps = 24/162 (14%)
Query: 56 ARYSKSNPIARWHICQGS-----------INSISFSADGAYLATVGRDGYLRVFDYTKEH 104
A S + + W+I G + ++ F G +AT D +++++ +
Sbjct: 994 ASASTDSSVRLWNISTGQCFQILLEHTDWVYAVVFHPQGKIIATGSADCTVKLWNISTGQ 1053
Query: 105 LVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
+ + +L AWS DG+ + + D V++W + V GH++ V F
Sbjct: 1054 CLKTLSEHSDKILGMAWSPDGQLLASASADQSVRLWDCCTGRCVGILRGHSNRVYSAIF- 1112
Query: 165 SYWSSPNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPL 206
+ NGE I + D + +W+ + + + L
Sbjct: 1113 --------SPNGEII----ATCSTDQTVKIWDWQQGKCLKTL 1142
>PWP2_SCHPO (Q9C1X1) Periodic tryptophan protein 2 homolog
Length = 854
Score = 52.0 bits (123), Expect = 2e-06
Identities = 29/96 (30%), Positives = 52/96 (53%), Gaps = 1/96 (1%)
Query: 69 ICQGSINSISFSADGAYLAT-VGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKY 127
I Q +I++++ ++ G ++A + G L V+++ E V +S+Y L +S DG+
Sbjct: 294 ITQSNIDTLTVNSTGDWIAIGSSKLGQLLVWEWQSESYVLKQQSHYDALSTLQYSSDGQR 353
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
I+TG +D ++VW M + H S VSG+ F
Sbjct: 354 IITGADDGKIKVWDMNSGFCIVTFTQHTSAVSGLCF 389
Score = 42.7 bits (99), Expect = 0.001
Identities = 25/105 (23%), Positives = 50/105 (46%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
+G ++S+SF++ G+ LA+ D +R++D + +L A+ DGK +
Sbjct: 467 EGPVSSLSFNSSGSLLASGSWDKTVRIWDIFSRSGIVEPLPIPSDVLSLAFHPDGKEVCV 526
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDN 175
D + W++++ K + +G G FD ++ NS+ N
Sbjct: 527 ASLDGQLTFWNVQEGKQTSLIDGRKDLSGGRRFDDARTAENSSLN 571
>PWP2_HUMAN (Q15269) Periodic tryptophan protein 2 homolog
Length = 919
Score = 52.0 bits (123), Expect = 2e-06
Identities = 32/103 (31%), Positives = 54/103 (52%), Gaps = 1/103 (0%)
Query: 62 NPIARWHICQGSINSISFSADGAYLAT-VGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCA 120
N I I SI S++ ++ G ++A G L V+++ E V + ++ ++ A
Sbjct: 321 NLIHSLSISDQSIASVAINSSGDWIAFGCSGLGQLLVWEWQSESYVLKQQGHFNSMVALA 380
Query: 121 WSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
+S DG+YI+TGG+D V+VW+ H+S V+GV F
Sbjct: 381 YSPDGQYIVTGGDDGKVKVWNTLSGFCFVTFTEHSSGVTGVTF 423
Score = 38.9 bits (89), Expect = 0.018
Identities = 43/187 (22%), Positives = 72/187 (37%), Gaps = 41/187 (21%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYG----GLLCCAWSMDGKYIL 129
+ ++F+A G + T DG +R FD H +++ C A G+ +
Sbjct: 418 VTGVTFTATGYVVVTSSMDGTVRAFDL---HRYRNFRTFTSPRPTQFSCVAVDASGEIVS 474
Query: 130 TGGEDDL-VQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQ 188
G +D + VWSM+ +++ GH +SG+ F+ S S
Sbjct: 475 AGAQDSFEIFVWSMQTGRLLDVLSGHEGPISGLCFNPMKSV-------------LASASW 521
Query: 189 DTQLLLWE------------LEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWD--NA 234
D + LW+ L D + V R P G+ + SQ + WD NA
Sbjct: 522 DKTVRLWDMFDSWRTKETLALTSDALAVTFR---PDGAEL---AVATLNSQITFWDPENA 575
Query: 235 VPLGTLQ 241
V G+++
Sbjct: 576 VQTGSIE 582
Score = 33.1 bits (74), Expect = 0.97
Identities = 40/151 (26%), Positives = 61/151 (39%), Gaps = 23/151 (15%)
Query: 71 QGSINSISFSADG-AYLATVGRDGYLRVFDYTKE--HLVCGGKSYYG---GLLCCAWSMD 124
+GS++S+SFS DG ++ T G + K + K+Y+G C W+ D
Sbjct: 96 KGSVHSVSFSPDGRKFVVTKGNIAQMYHAPGKKREFNAFVLDKTYFGPYDETTCIDWTDD 155
Query: 125 GKYILTGGEDDLVQVWSME--DRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYR 182
+ + G +D V+ E D + GH + F+S NS D
Sbjct: 156 SRCFVVGSKDMSTWVFGAERWDNLIYYALGGHKDAIVACFFES-----NSLD-------- 202
Query: 183 FGSVGQDTQLLLWELEMDEIVVPLRRGPPGG 213
S+ QD L +W + D LR PP G
Sbjct: 203 LYSLSQDGVLCMW--QCDTPPEGLRLKPPAG 231
>WDR1_DROME (Q9VU68) Putative actin interacting protein 1 (AIP1)
Length = 608
Score = 51.6 bits (122), Expect = 3e-06
Identities = 24/78 (30%), Positives = 44/78 (55%), Gaps = 2/78 (2%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLV--CGGKSYYGGLLCCAWSMDGKYILTG 131
+ ++ +S DG + A+ G DG + ++D T LV G ++ GG+ AW D +LT
Sbjct: 198 VQAVRYSPDGKFFASAGFDGKVFLYDGTSSELVGEFGSPAHKGGVYALAWKPDSTQLLTC 257
Query: 132 GEDDLVQVWSMEDRKVVA 149
D ++W++E R++V+
Sbjct: 258 SGDKTCRLWTVESRELVS 275
>WDR1_HUMAN (O75083) WD-repeat protein 1 (Actin interacting protein
1) (AIP1) (NORI-1)
Length = 606
Score = 51.2 bits (121), Expect = 3e-06
Identities = 24/79 (30%), Positives = 40/79 (50%), Gaps = 3/79 (3%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVC---GGKSYYGGLLCCAWSMDGKYILT 130
+N + FS DG AT DG + ++D VC G K++ GG+ +WS D ++L+
Sbjct: 192 VNCVRFSPDGNRFATASADGQIYIYDGKTGEKVCALGGSKAHDGGIYAISWSPDSTHLLS 251
Query: 131 GGEDDLVQVWSMEDRKVVA 149
D ++W + VV+
Sbjct: 252 ASGDKTSKIWDVSVNSVVS 270
Score = 48.9 bits (115), Expect = 2e-05
Identities = 23/78 (29%), Positives = 44/78 (55%), Gaps = 7/78 (8%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFD----YTKEHLVCGGKSYYGGLLCCAWSMDGK 126
+G + +++S DGA+LA + VF Y++ ++ G ++ ++C AWS D +
Sbjct: 489 KGPVTDVAYSHDGAFLAVCDASKVVTVFSVADGYSENNVFYG---HHAKIVCLAWSPDNE 545
Query: 127 YILTGGEDDLVQVWSMED 144
+ +GG D +V VW++ D
Sbjct: 546 HFASGGMDMMVYVWTLSD 563
Score = 33.1 bits (74), Expect = 0.97
Identities = 30/104 (28%), Positives = 48/104 (45%), Gaps = 7/104 (6%)
Query: 79 FSADGAYLATVGRDGYLRVFDYT-KEHLV-CGGKSYYGGLLCCAWSMDGKYILTGGE--D 134
++ G Y+A+ G LR++D T KEHL+ + + G + AW+ D K I GE +
Sbjct: 66 YAPSGFYIASGDVSGKLRIWDTTQKEHLLKYEYQPFAGKIKDIAWTEDSKRIAVVGEGRE 125
Query: 135 DLVQVWSMEDRKVVAWGEGHNSWVSGVAFDS---YWSSPNSNDN 175
V+ + V GHN ++ V Y + S+DN
Sbjct: 126 KFGAVFLWDSGSSVGEITGHNKVINSVDIKQSRPYRLATGSDDN 169
Score = 31.2 bits (69), Expect = 3.7
Identities = 30/117 (25%), Positives = 47/117 (39%), Gaps = 23/117 (19%)
Query: 11 CISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWHIC 70
C+ + P G+ F A ADG +Y DG +T V SK++
Sbjct: 194 CVRFSPDGN-RFATASADGQIYIYDG-----------KTGEKVCALGGSKAH-------- 233
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCG---GKSYYGGLLCCAWSMD 124
G I +IS+S D +L + D +++D + +V G + L C W D
Sbjct: 234 DGGIYAISWSPDSTHLLSASGDKTSKIWDVSVNSVVSTFPMGSTVLDQQLGCLWQKD 290
>GBLP_TRYBB (Q94775) Guanine nucleotide-binding protein beta
subunit-like protein (Activated protein kinase C
receptor homolog) (Track)
Length = 318
Score = 51.2 bits (121), Expect = 3e-06
Identities = 37/139 (26%), Positives = 64/139 (45%), Gaps = 16/139 (11%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAW---SMDGKYILTGG 132
S++FS D + + GRD LRV++ E + + + + C S+D I++GG
Sbjct: 114 SVAFSPDNRQIVSGGRDNALRVWNVKGECMHTLSRGAHTDWVSCVRFSPSLDAPVIVSGG 173
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQL 192
D+LV+VW + ++V +GH ++V+ V S S+D +D
Sbjct: 174 WDNLVKVWDLATGRLVTDLKGHTNYVTSVTVSPDGSLCASSD-------------KDGVA 220
Query: 193 LLWELEMDEIVVPLRRGPP 211
LW+L E + + G P
Sbjct: 221 RLWDLTKGEALSEMAAGAP 239
Score = 42.7 bits (99), Expect = 0.001
Identities = 30/135 (22%), Positives = 59/135 (43%), Gaps = 12/135 (8%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
++ ++ S +G + + D LR+++ + +L A+S D + I++GG
Sbjct: 70 VSDVALSNNGNFAVSASWDHSLRLWNLQNGQCQYKFLGHTKDVLSVAFSPDNRQIVSGGR 129
Query: 134 DDLVQVWSMEDRKVVAWGEG-HNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQL 192
D+ ++VW+++ + G H WVS V F +P S G D +
Sbjct: 130 DNALRVWNVKGECMHTLSRGAHTDWVSCVRFSPSLDAP-----------VIVSGGWDNLV 178
Query: 193 LLWELEMDEIVVPLR 207
+W+L +V L+
Sbjct: 179 KVWDLATGRLVTDLK 193
Score = 37.4 bits (85), Expect = 0.051
Identities = 20/78 (25%), Positives = 37/78 (46%), Gaps = 8/78 (10%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVC-------GGKSYYGGLLCCAWSMDGK 126
IN I FS + ++ G +R+FD + ++ G K + AWS DG
Sbjct: 240 INQICFSPNRYWMCAATEKG-IRIFDLENKDIIVELAPEHQGSKKIVPECVSIAWSADGS 298
Query: 127 YILTGGEDDLVQVWSMED 144
+ +G D++++VW + +
Sbjct: 299 TLYSGYTDNVIRVWGVSE 316
>YE91_SYNY3 (P74598) Hypothetical WD-repeat protein sll1491
Length = 348
Score = 50.4 bits (119), Expect = 6e-06
Identities = 44/169 (26%), Positives = 69/169 (40%), Gaps = 13/169 (7%)
Query: 38 GESSFPILKDQTLFSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRV 97
GE++ ++VA S S + QG I +++ + DG YLA D + +
Sbjct: 28 GENAHANASTPNPYTVAQTTASPSVAVENLSGFQGIITALNITPDGKYLAVATADNQITL 87
Query: 98 FDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSW 157
D + +V +S A S DG+++ D+ V V + D V GH
Sbjct: 88 IDLANQEVVYSQRSPVNNFADLAISADGQWLAIAA-DNNVDVRRVRDGMRVETLVGHTDK 146
Query: 158 VSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPL 206
VSGVAF + +GETI G D + +WE ++ L
Sbjct: 147 VSGVAF---------SPDGETIV---SVSGGDRTIRIWERASGNLIQTL 183
Score = 47.0 bits (110), Expect = 6e-05
Identities = 21/69 (30%), Positives = 39/69 (56%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
IN+I+ S + ++AT ++G + +FD V + + G +L A+S DG + +G E
Sbjct: 275 INTIAVSPNNRWVATANKEGTIMIFDLANGKQVTTLRGHQGWVLSLAFSPDGNTLYSGAE 334
Query: 134 DDLVQVWSM 142
D V++W +
Sbjct: 335 DKTVKIWDL 343
Score = 31.6 bits (70), Expect = 2.8
Identities = 29/128 (22%), Positives = 56/128 (43%), Gaps = 18/128 (14%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYT--KEHLVCGGKSYYGGLLCCAWSMDGKYIL 129
G IN ++ + DG L R+ +++ ++ KE G S + A S + +++
Sbjct: 232 GFINGLAVTPDGRKLVGAVRN-FVKAWNLADAKELFSVRGPSLEINTI--AVSPNNRWVA 288
Query: 130 TGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQD 189
T ++ + ++ + + K V GH WV +AF SP+ N S +D
Sbjct: 289 TANKEGTIMIFDLANGKQVTTLRGHQGWVLSLAF-----SPDGN--------TLYSGAED 335
Query: 190 TQLLLWEL 197
+ +W+L
Sbjct: 336 KTVKIWDL 343
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.133 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,242,074
Number of Sequences: 164201
Number of extensions: 2152895
Number of successful extensions: 6417
Number of sequences better than 10.0: 353
Number of HSP's better than 10.0 without gapping: 195
Number of HSP's successfully gapped in prelim test: 161
Number of HSP's that attempted gapping in prelim test: 4836
Number of HSP's gapped (non-prelim): 1350
length of query: 325
length of database: 59,974,054
effective HSP length: 110
effective length of query: 215
effective length of database: 41,911,944
effective search space: 9011067960
effective search space used: 9011067960
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC148995.8


