

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.4 - phase: 0
(1009 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
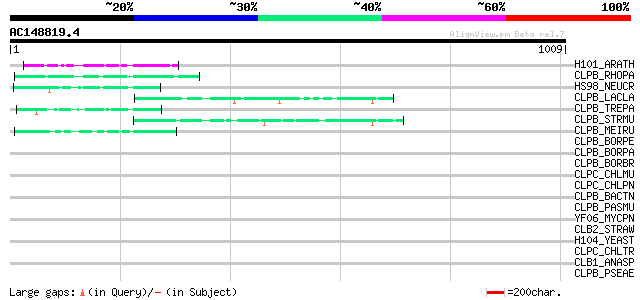
Score E
Sequences producing significant alignments: (bits) Value
H101_ARATH (P42730) Heat shock protein 101 78 1e-13
CLPB_RHOPA (Q6N1H2) Chaperone clpB 53 4e-06
HS98_NEUCR (P31540) Heat shock protein HSP98 52 6e-06
CLPB_LACLA (Q9CFF3) Chaperone clpB 52 6e-06
CLPB_TREPA (O83110) Chaperone clpB 51 2e-05
CLPB_STRMU (Q8DTC7) Chaperone clpB 46 6e-04
CLPB_MEIRU (Q7X2S8) Chaperone clpB 45 8e-04
CLPB_BORPE (Q7VYV6) Chaperone clpB 45 0.001
CLPB_BORPA (Q7W9E6) Chaperone clpB 45 0.001
CLPB_BORBR (Q7WHB6) Chaperone clpB 45 0.001
CLPC_CHLMU (Q9PKA8) Probable ATP-dependent Clp protease ATP-bind... 44 0.002
CLPC_CHLPN (Q9Z8A6) Probable ATP-dependent Clp protease ATP-bind... 44 0.003
CLPB_BACTN (Q89YY3) Chaperone clpB 44 0.003
CLPB_PASMU (Q9CKC0) Chaperone clpB 42 0.008
YF06_MYCPN (P75280) Hypothetical lipoprotein MPN506 precursor (P... 41 0.014
CLB2_STRAW (Q826F2) Chaperone clpB 2 41 0.014
H104_YEAST (P31539) Heat shock protein 104 41 0.018
CLPC_CHLTR (O84288) Probable ATP-dependent Clp protease ATP-bind... 41 0.018
CLB1_ANASP (Q8YUL9) Chaperone clpB 1 40 0.032
CLPB_PSEAE (Q9HVN5) Chaperone clpB 40 0.041
>H101_ARATH (P42730) Heat shock protein 101
Length = 911
Score = 78.2 bits (191), Expect = 1e-13
Identities = 75/284 (26%), Positives = 128/284 (44%), Gaps = 40/284 (14%)
Query: 26 LARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQCRALELCFNVALNRLPT 85
LA GHAQ TPLH+A L+S P+ A + +N ++ E N AL +LP+
Sbjct: 20 LAVNAGHAQFTPLHLAGALISDPTGIFPQAISSAGGEN----AAQSAERVINQALKKLPS 75
Query: 86 TTTSPLLQPQHVPSLSNALIAALKRAQAHQR-RGCIEQNQQQQQQPLLSVKVELDQLILS 144
+ P +P+ S++LI ++RAQA Q+ RG + +DQLI+
Sbjct: 76 QSP----PPDDIPA-SSSLIKVIRRAQAAQKSRG--------------DTHLAVDQLIMG 116
Query: 145 ILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFLSSYGYGSVL 204
+L+D + ++ E G ++ VK+ +E L S S ++ L +YG ++
Sbjct: 117 LLEDSQIRDLLNEVGVATARVKSEVEK---LRGKEGKKVESASGDTNFQALKTYG-RDLV 172
Query: 205 FSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKR 264
+ K + V+ E+I V +L R+ K N V++G+ +V + +R
Sbjct: 173 EQAGKLDPVI---------GRDEEIRRVVRILSRRTKNNPVLIGEPGVGKTAVVEGLAQR 223
Query: 265 FERGEVPDEMKTTHFVKFH-GLSSVSLKYMKKEEVEMNVIRVLK 307
+G+VP+ + + G KY + E E + VLK
Sbjct: 224 IVKGDVPNSLTDVRLISLDMGALVAGAKY--RGEFEERLKSVLK 265
Score = 33.5 bits (75), Expect = 3.0
Identities = 26/88 (29%), Positives = 46/88 (51%), Gaps = 12/88 (13%)
Query: 617 QKRMENLTLQRDHIYK-VLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQGN 670
Q E L D ++K V+ +N + V++++EA++ S++ + ++LFL G
Sbjct: 554 QNEKERLIGLADRLHKRVVGQN-----QAVNAVSEAILRSRAGLGRPQQPTGSFLFL-GP 607
Query: 671 DSVGKKRLALAIAESVFGSVEMFSHVDM 698
VGK LA A+AE +F + +DM
Sbjct: 608 TGVGKTELAKALAEQLFDDENLLVRIDM 635
>CLPB_RHOPA (Q6N1H2) Chaperone clpB
Length = 879
Score = 53.1 bits (126), Expect = 4e-06
Identities = 72/338 (21%), Positives = 130/338 (38%), Gaps = 43/338 (12%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T ++ + LA R GH Q +PLH+ LL S L + N+
Sbjct: 4 EKYTERVRGFIQSAQSLAMREGHQQFSPLHILKVLLD-DSEGLAGGLIDRAGGNS----- 57
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
RA+ AL ++P + S Q P+ + A AA K A+
Sbjct: 58 RAILKATEEALGKMPKVSGSGAGQVYLAPATARAFDAAEKAAEKAGDS------------ 105
Query: 130 PLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPL 189
V VE L LS+ D +++ + G + +N + IN+ ++ S
Sbjct: 106 ---FVTVERLLLALSLDKDSEAGQLLTKGGVTP-------QNLNAAINALRKGRTADSAT 155
Query: 190 SHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGD 249
+ N + + Y L + + ++ P + E+I VL R+ K N V++G+
Sbjct: 156 AENAYDALKKYARDLTQAARDGKL--DPVI----GRDEEIRRTIQVLSRRTKNNPVLIGE 209
Query: 250 TVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMKKEEVEMNVIRVLKR 308
+V + R G+VP+ +K + G KY + E LK
Sbjct: 210 PGVGKTAIVEGLALRILNGDVPESLKDKKLLALDMGALIAGAKYRGEFEER------LKA 263
Query: 309 KVSDYVALGVGAIFYVGDLKWIVDDN--DGSLNEKEVV 344
+++ A G I ++ ++ +V DG+++ ++
Sbjct: 264 VLNEVTAAEGGIILFIDEMHTLVGAGKADGAMDASNLL 301
>HS98_NEUCR (P31540) Heat shock protein HSP98
Length = 927
Score = 52.4 bits (124), Expect = 6e-06
Identities = 61/278 (21%), Positives = 113/278 (39%), Gaps = 39/278 (14%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSL---PSSSLRNACLKSQQQN 63
T + T A L+ ++ LA + H+Q+ P+H+A LL PS +NA +
Sbjct: 2 TSKMEFTDRAKKALEDAMALAEQYAHSQLLPVHLAVALLDPLPDPSKDQQNAPAGATSSL 61
Query: 64 NHPLQCRA------LELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRR 117
+ RA + AL RLP+ P HV S++ + L++A
Sbjct: 62 FRQVIERAHGDPQLFDRALKKALVRLPSQDPPP----DHV-SMAPSFHTVLRKA------ 110
Query: 118 GCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLIN 177
N+ Q+ Q + +D LI ++ ++PS+ ++EA P + + ++ I
Sbjct: 111 -----NELQKTQK--DTYIAVDHLITALAEEPSIMNALKEANIPKPKL---VTDAIQAIR 160
Query: 178 SSSVFHSSPSPLSHNHF-LSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVL 236
+ S + H L+ + + + K V +E+I V +L
Sbjct: 161 GTKRVDSRNADTEEEHENLAKFTIDMTAMAREGKIDPVI--------GREEEIRRVIRIL 212
Query: 237 LRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEM 274
R+ K N V++G+ +V + +R +VPD +
Sbjct: 213 SRRTKNNPVLIGEPGVGKTTVVEGLAQRIVNADVPDNL 250
>CLPB_LACLA (Q9CFF3) Chaperone clpB
Length = 867
Score = 52.4 bits (124), Expect = 6e-06
Identities = 93/497 (18%), Positives = 191/497 (37%), Gaps = 71/497 (14%)
Query: 227 EDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKT-THFVKFHGL 285
E+I V VL RK K N V++G+ +V + +R R +VP+ +K T F G
Sbjct: 187 EEIRDVIRVLSRKTKNNPVLIGEPGVGKTAIVEGLAQRIVRKDVPENLKDKTIFSLDMGA 246
Query: 286 SSVSLKYMKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVD--DNDGSLNEKEV 343
KY + E + + +K + I ++ +L IV +GS++ +
Sbjct: 247 LIAGAKYRGEFEERLKAVLNEVKKSDGQI------ILFIDELHTIVGAGKTEGSMDAGNL 300
Query: 344 VDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLG 403
+ ++ G++ L+ + Y + E ++ V P+
Sbjct: 301 LKPMLAR------------GELHLIGATTLDEYRKYMETDKALERRFQKVLVTEPTVEDT 348
Query: 404 LSL-----------HSSSVHDSKMSISQNPSPMLESKFFSNKEEHEK-LNCCEECVSNYE 451
+S+ H ++HD+ + + L +++ +++ +K ++ +E +
Sbjct: 349 ISILRGLKERFEIHHGVTIHDNALVAAAT----LSNRYITDRFLPDKAIDLVDEASATIR 404
Query: 452 KEAQLFKPDQKNLLPSWLQSHSTEARQKDELTQLNKKW-----NRLCQCLHQNKQPQNHW 506
E + +Q EA K E +KK + + +N Q + W
Sbjct: 405 VEMNSLPTELDQANRRLMQLEIEEAALKKERDDASKKRLEIIRGEIAELREENNQLKAQW 464
Query: 507 SNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQS 566
I + + ++ L + + + +A A IP +
Sbjct: 465 EAEKKEVGNISEKRNELEHARHE----LEEAQNEGNLEKAA---ALRYGKIPEIEK---- 513
Query: 567 CTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQ 626
E +EKA+ + ++ E E + T +G T + K +E +
Sbjct: 514 ---ELKAIEEKAKSDDLSLV--------QESVTEEQITEVVGRMT-GIPITKLVEGEREK 561
Query: 627 RDHIYKVLQENIPWHCETVSSIAEALVDSKS-----SKECATWLFLQGNDSVGKKRLALA 681
H+ + L + + E V ++++A++ +++ ++ ++LFL G VGK LA A
Sbjct: 562 LLHLPETLHQRVVGQDEAVEAVSDAIIRARAGIQDPNRPLGSFLFL-GPTGVGKTELAKA 620
Query: 682 IAESVFGSVEMFSHVDM 698
+AE++F S E +DM
Sbjct: 621 LAENLFDSEEHMVRIDM 637
>CLPB_TREPA (O83110) Chaperone clpB
Length = 878
Score = 50.8 bits (120), Expect = 2e-05
Identities = 64/268 (23%), Positives = 107/268 (39%), Gaps = 46/268 (17%)
Query: 13 TAEAASILKHSLGLARRRGHAQITPLHVAATLLS----LPSSSLRNACLKSQQQNNHPLQ 68
T +A+ L ++ LA H Q+ H+ LLS + S + K + LQ
Sbjct: 7 TVKASEALNDAISLAEAENHGQVEEEHLLHALLSQKDGIISPLIEKIGAKPDFLYDELLQ 66
Query: 69 CRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQ 128
C L R P T P Q + P+LS A A + A +NQ +
Sbjct: 67 C----------LRRKPRVT-GPAAQTRCAPTLSKACARAERLAL---------KNQDEY- 105
Query: 129 QPLLSVKVELDQLILSILD-DPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPS 187
V + L+L+I + D + +R++ G +S ++ L++ + S V +S
Sbjct: 106 -------VSCEHLLLAISETDSNTARLLHSQGITSKTISAALKD---IRGSKRV--TSQD 153
Query: 188 PLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIV 247
P S L Y + ++K V E+I V VL R+ K N V++
Sbjct: 154 PESTFQCLEKYCRDLTTLAREEKIDPVI--------GRDEEIRRVMQVLSRRTKNNPVLI 205
Query: 248 GDTVSLTEGLVSEIMKRFERGEVPDEMK 275
G+ +V + +R G+VP+ +K
Sbjct: 206 GEPGVGKTAIVEGLARRIVSGDVPESLK 233
>CLPB_STRMU (Q8DTC7) Chaperone clpB
Length = 860
Score = 45.8 bits (107), Expect = 6e-04
Identities = 101/510 (19%), Positives = 195/510 (37%), Gaps = 68/510 (13%)
Query: 226 KEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-G 284
++++N + +L RK K N V+VG + ++ ++R + +VP ++ G
Sbjct: 186 QDEMNDIIRILSRKNKNNAVLVGHSGVGKTAIIEGFVQRLVKNQVPKNLQGKVIYSLDIG 245
Query: 285 LSSVSLKYMKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDN--DGSLNEKE 342
KY + E + ++D + I ++ ++ IV +GS++
Sbjct: 246 ALLAGTKYRGEFEERFKAL------LNDIIDSNGQIILFIDEIHSIVSAGRTEGSVDAAS 299
Query: 343 VVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGL 402
++ ++ GKI ++ + ++ Y E ++ Q + V L
Sbjct: 300 ILKPLLAR------------GKIRIIGSTTHAEYRESIEYDRALERRF--QRILVHEPDL 345
Query: 403 GLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFKPDQ- 461
D M I + P+ E+ F + E + L Y + F PD+
Sbjct: 346 ----------DYTMEILKGLKPIYEN-FHGVRLEEDGLEAAASLSKRYISDR--FLPDKA 392
Query: 462 -----KNLLPSWLQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKI 516
+ L SH+ + + QL + RL + L + NNH +
Sbjct: 393 LDLIDEACAAKRLASHAYPVQMTNLSQQLISEKIRLLR-LTDSDTDDKFKLNNHLEQLEE 451
Query: 517 YPYNSSYPYWPNQGSSI--LPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFN 574
+ W + + + + S ++ + A +N + + R
Sbjct: 452 QK-SQLLANWQRERELLDQVQELRSHLTALEEQANLALENNQVADYVR----------LE 500
Query: 575 DEKAQK-NQVTATLE---LDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHI 630
DE+ +K +Q LE LDS K + T + A+ + Q MEN + H+
Sbjct: 501 DEEIRKCHQKMTALEKERLDSQKIINVTVTREDIAAVVERLTGIKVQGVMENERDRLLHL 560
Query: 631 YKVLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQGNDSVGKKRLALAIAES 685
++L + I + V +++A++ S++ + ++LFL G VGK LA +AE
Sbjct: 561 EELLHQKIVGQDQAVQKVSQAIIRSRAGIQNPKRPIGSFLFL-GPTGVGKTALAKRLAEV 619
Query: 686 VFGSVEMFSHVDMMKRENSETPFSEKVVGP 715
+FGS +DM E E ++VGP
Sbjct: 620 LFGSELEMVRLDM--SEYMEKHAVSRLVGP 647
>CLPB_MEIRU (Q7X2S8) Chaperone clpB
Length = 854
Score = 45.4 bits (106), Expect = 8e-04
Identities = 68/295 (23%), Positives = 116/295 (39%), Gaps = 45/295 (15%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T +A + S LAR H++I H+AA +L + K+ Q + Q
Sbjct: 4 ERYTEQARQAIAQSQVLARESAHSKIDLPHLAAVMLRDAAGLPAKIVQKAGQNPQNIYQA 63
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
EL RLP + + Q LS+ L +AL RA+ +
Sbjct: 64 AQSEL------GRLPKVSGTEGGQ-----YLSSRLASALGRAE-------------KLAD 99
Query: 130 PLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPL 189
L V LD L+L++ + G + +V+ L+ I +S +
Sbjct: 100 ELKDRFVALDTLLLALAETGY-------GGLQASAVRQALQE----IRGGRTVNSEHAEG 148
Query: 190 SHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGD 249
++N L YG L +++ E+ P + E+I V +LLR+ K N V++G+
Sbjct: 149 TYNA-LEQYG----LDLTRQAEEGKLDPVI----GRDEEIRRVIQILLRRTKNNPVLIGE 199
Query: 250 TVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMKKEEVEMNVI 303
+V + +R +G+VP+ +K V G KY + E + +
Sbjct: 200 PGVGKTAVVEGLAQRIVKGDVPEGLKGKRIVSLQMGSLLAGAKYRGEFEERLKAV 254
>CLPB_BORPE (Q7VYV6) Chaperone clpB
Length = 865
Score = 45.1 bits (105), Expect = 0.001
Identities = 67/304 (22%), Positives = 113/304 (37%), Gaps = 40/304 (13%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
+ LT + L + LA R H I P+HV A LL P S + ++ N
Sbjct: 4 DKLTTKFQQALADAQSLAARNDHPYIEPVHVLAALLGDPDSGAASLLARAGVAVNR---- 59
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
++ + AL LP +Q L + L+ K A RRG
Sbjct: 60 --VQPAIDSALKGLPQVQGDDNVQVGR--ELQSVLVRTDKEA---ARRG----------- 101
Query: 130 PLLSVKVELDQLILSILDDP-SVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
+ + +L++ DD R++REAG K LE + + S
Sbjct: 102 ---DTYIASELFLLALADDKGDAGRILREAGLQ----KKALEAAIDAVRGGENV-SGAEG 153
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
S+ LS Y L +++ Q P + ++I +L R+ K N V++G
Sbjct: 154 ESNREALSKY----TLDLTERARQGKLDPVI----GRDDEIRRTIQILQRRTKNNPVLIG 205
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKR 308
+ +V + +R EVP+ ++ + L+++ + E E + VLK
Sbjct: 206 EPGVGKTAIVEGLAQRIVNDEVPETLRGKRVLSL-DLAALLAGAKFRGEFEERLKAVLKE 264
Query: 309 KVSD 312
D
Sbjct: 265 LAQD 268
>CLPB_BORPA (Q7W9E6) Chaperone clpB
Length = 865
Score = 45.1 bits (105), Expect = 0.001
Identities = 67/304 (22%), Positives = 113/304 (37%), Gaps = 40/304 (13%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
+ LT + L + LA R H I P+HV A LL P S + ++ N
Sbjct: 4 DKLTTKFQQALADAQSLAARNDHPYIEPVHVLAALLGDPDSGAASLLARAGVAVNR---- 59
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
++ + AL LP +Q L + L+ K A RRG
Sbjct: 60 --VQPAIDSALKGLPQVQGEDNVQVGR--ELQSVLVRTDKEA---ARRG----------- 101
Query: 130 PLLSVKVELDQLILSILDDP-SVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
+ + +L++ DD R++REAG K LE + + S
Sbjct: 102 ---DTYIASELFLLALADDKGDAGRILREAGLQ----KKALEAAIDAVRGGENV-SGAEG 153
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
S+ LS Y L +++ Q P + ++I +L R+ K N V++G
Sbjct: 154 ESNREALSKY----TLDLTERARQGKLDPVI----GRDDEIRRTIQILQRRTKNNPVLIG 205
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKR 308
+ +V + +R EVP+ ++ + L+++ + E E + VLK
Sbjct: 206 EPGVGKTAIVEGLAQRIVNDEVPETLRGKRVLSL-DLAALLAGAKFRGEFEERLKAVLKE 264
Query: 309 KVSD 312
D
Sbjct: 265 LAQD 268
>CLPB_BORBR (Q7WHB6) Chaperone clpB
Length = 865
Score = 45.1 bits (105), Expect = 0.001
Identities = 67/304 (22%), Positives = 113/304 (37%), Gaps = 40/304 (13%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
+ LT + L + LA R H I P+HV A LL P S + ++ N
Sbjct: 4 DKLTTKFQQALADAQSLAARNDHPYIEPVHVLAALLGDPDSGAASLLARAGVAVNR---- 59
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
++ + AL LP +Q L + L+ K A RRG
Sbjct: 60 --VQPAIDSALKGLPQVQGEDNVQVGR--ELQSVLVRTDKEA---ARRG----------- 101
Query: 130 PLLSVKVELDQLILSILDDP-SVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
+ + +L++ DD R++REAG K LE + + S
Sbjct: 102 ---DTYIASELFLLALADDKGDAGRILREAGLQ----KKALEAAIDAVRGGENV-SGAEG 153
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
S+ LS Y L +++ Q P + ++I +L R+ K N V++G
Sbjct: 154 ESNREALSKY----TLDLTERARQGKLDPVI----GRDDEIRRTIQILQRRTKNNPVLIG 205
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKR 308
+ +V + +R EVP+ ++ + L+++ + E E + VLK
Sbjct: 206 EPGVGKTAIVEGLAQRIVNDEVPETLRGKRVLSL-DLAALLAGAKFRGEFEERLKAVLKE 264
Query: 309 KVSD 312
D
Sbjct: 265 LAQD 268
>CLPC_CHLMU (Q9PKA8) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 870
Score = 44.3 bits (103), Expect = 0.002
Identities = 68/322 (21%), Positives = 126/322 (39%), Gaps = 49/322 (15%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T A ++K + A+R H + H+ LL L N
Sbjct: 19 EKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVL------------- 65
Query: 70 RALELCFNVALNRLPTTTT-SPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQ 128
R L + F+ A N + P +Q P+L+ + + + A++ +E N +
Sbjct: 66 RTLGVDFDTAKNEVERLIGYGPEIQVYGDPALTGRVKKSFE--SANEEASILEHNYVGTE 123
Query: 129 QPLLSVKVELDQLILSILD----DPSVSR--VMREAGF--------SSPSVKNNLENSST 174
LL + + D + L +L+ DP R +++E SS + +N +SS+
Sbjct: 124 HLLLGILNQADGVALQVLENLHIDPKEIRKEILKELETFNLQLPPSSSITPRNTNSSSSS 183
Query: 175 LINSSSVFHSS-----PSPLSHNHFLSSYGYG-SVLFSSQKKEQVVYHPFLKSSESNKED 228
SSS+ S P LS L +YGY + +F + + V+ +
Sbjct: 184 SSKSSSLGGHSLGGDKPEKLSA---LKAYGYDLTEMFKEARLDPVI---------GRSAE 231
Query: 229 INLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSS 287
+ + +L R++K N V++G+ +V + ++ GEVP+ ++ + L
Sbjct: 232 VERLILILCRRRKNNPVLIGEAGVGKTAIVEGLAQKIVSGEVPEALRKKRLITLDLALMI 291
Query: 288 VSLKYMKKEEVEMNVIRVLKRK 309
KY + E + + RK
Sbjct: 292 AGTKYRGQFEERIKAVMDEVRK 313
>CLPC_CHLPN (Q9Z8A6) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 845
Score = 43.5 bits (101), Expect = 0.003
Identities = 68/315 (21%), Positives = 121/315 (37%), Gaps = 38/315 (12%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T A ++K + A+R H + H+ LL L N
Sbjct: 3 EKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVL------------- 49
Query: 70 RALELCFNVALNRLPTTTT-SPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQ 128
R L + F+ A + P +Q P+L+ + + + A++ +E N +
Sbjct: 50 RNLGIDFDTARQEVERLIGYGPEIQVYGDPALTGRVKKSFE--SANEEASLLEHNYVGTE 107
Query: 129 QPLLSVKVELDQLILSILDDPSVS--RVMREAGFSSPSVKNNLENSSTLINSSSVFH--S 184
LL + + D + L +L++ + V +E + L SS+ +SSS + S
Sbjct: 108 HLLLGILHQSDSVALQVLENLHIDPREVRKEILKELETFNLQLPPSSSSSSSSSRSNPSS 167
Query: 185 SPSPLSHN---------HFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDV 235
S SPL H+ L +YGY + K V +SSE + + +
Sbjct: 168 SKSPLGHSLGSDKNEKLSALKAYGYDLTEMVRESKLDPVIG---RSSE-----VERLILI 219
Query: 236 LLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMK 294
L R++K N V++G+ +V + ++ EVPD ++ + L KY
Sbjct: 220 LCRRRKNNPVLIGEAGVGKTAIVEGLAQKIILNEVPDALRKKRLITLDLALMIAGTKYRG 279
Query: 295 KEEVEMNVIRVLKRK 309
+ E + + RK
Sbjct: 280 QFEERIKAVMDEVRK 294
>CLPB_BACTN (Q89YY3) Chaperone clpB
Length = 862
Score = 43.5 bits (101), Expect = 0.003
Identities = 41/182 (22%), Positives = 80/182 (43%), Gaps = 16/182 (8%)
Query: 136 VELDQLILSILDDPS-VSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHF 194
V L+ ++L++L S VS ++++AG + ++N IN S S +++
Sbjct: 104 VSLEHILLALLTVKSTVSTILKDAGMTEKELRN-------AINELRKGEKVTSQSSEDNY 156
Query: 195 LSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLT 254
S Y L + + ++ P + E+I V +L R+ K N +++G+ +
Sbjct: 157 QSLEKYAINLNEAARSGKL--DPVI----GRDEEIRRVLQILSRRTKNNPILIGEPGTGK 210
Query: 255 EGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMKK-EEVEMNVIRVLKRKVSD 312
+V + R RG+VP+ +K G KY + EE +V+ +K+ D
Sbjct: 211 TAIVEGLAHRILRGDVPENLKNKQVYSLDMGALVAGAKYKGEFEERLKSVVNEVKKSEGD 270
Query: 313 YV 314
+
Sbjct: 271 II 272
Score = 39.7 bits (91), Expect = 0.041
Identities = 49/218 (22%), Positives = 90/218 (40%), Gaps = 24/218 (11%)
Query: 561 RRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTK-------EVKTTLALGNSTFS 613
R G E + +A ++ T E L+GM+G K + + + +
Sbjct: 486 REGDYGKVAEIRYGKLQALDKEIEDTQE--KLRGMQGDKAMIKEEVDAEDIADVVSRWTG 543
Query: 614 VSDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQ 668
+ K M++ + H+ L + + E + ++A+A+ S++ + +++FL
Sbjct: 544 IPVSKMMQSEKDKLLHLEDELHQRVIGQDEAIEAVADAVRRSRAGLQDPKRPIGSFIFL- 602
Query: 669 GNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVEN 728
G VGK LA A+AE +F M + +DM E E ++VG +V E
Sbjct: 603 GTTGVGKTELAKALAEFLFDDESMMTRIDM--SEYQEKHSVSRLVGAPPG---YVGYDEG 657
Query: 729 ADFGDTLIRK----MLADEFEIAKFGTLGQKIFILSNG 762
+ + RK +L DE E A + +L +G
Sbjct: 658 GQLTEAIRRKPYSVVLFDEIEKAHPDVFNILLQVLDDG 695
>CLPB_PASMU (Q9CKC0) Chaperone clpB
Length = 855
Score = 42.0 bits (97), Expect = 0.008
Identities = 66/296 (22%), Positives = 113/296 (37%), Gaps = 40/296 (13%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T + L+ + LA + + I P+H+ LL+ S A + + N PL
Sbjct: 4 EKFTTKFQQALQEAQSLAIGKDNQFIEPVHLLTALLNQQGGS--TAPILTASGANLPLLR 61
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
L N L++LP + S Q SL N L K AQ Q+Q
Sbjct: 62 NEL----NAELSKLPQVSGSGG-DVQVSRSLVNLLNLCDKLAQ-------------QRQD 103
Query: 130 PLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPL 189
+S ++ +L+ L+D ++ V+++ G K NL+ + + + P+
Sbjct: 104 KFISSEL----FLLAALEDKTLGDVLKKCGVK----KENLQQAIEKVRGGQNVND-PNAE 154
Query: 190 SHNHFLSSYGYG-SVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
L Y + S K + V+ E+I VL R+ K N V++G
Sbjct: 155 ESRQALEKYTIDLTARAESGKLDPVI---------GRDEEIRRAIQVLQRRTKNNPVLIG 205
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMKKEEVEMNVI 303
+ +V + +R GEVP+ +K + G KY + E + +
Sbjct: 206 EPGVGKTAIVEGLAQRIVNGEVPEGLKNKRVLSLDMGALIAGAKYRGEFEERLKAV 261
>YF06_MYCPN (P75280) Hypothetical lipoprotein MPN506 precursor
(P02_orf793)
Length = 793
Score = 41.2 bits (95), Expect = 0.014
Identities = 45/198 (22%), Positives = 84/198 (41%), Gaps = 17/198 (8%)
Query: 782 ISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEEN-EGNKKREFSFSR 840
I KK L+ +SSS+ + D + +IE++ + N K +
Sbjct: 170 ILNNAKKKEGTLNQKMTSSSEGKNSSGTLTVATDTETSSLWKKIEDSAKANGKSDEKGKG 229
Query: 841 QSSFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQ 900
+ N + ++ ++ E D+ +++ S D +++ GE+ + +D + FEF
Sbjct: 230 KKKDNKSATFSLVQLKQTQEKTDDSQDTKNSDDQVKKSWGEY----QEVDGVLKNFEFKA 285
Query: 901 SPAKNKEMTQMFMSRVKESFEEVLGNVKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQ 960
S +N F +RV +SF+++ N D I+ I +G S N++F
Sbjct: 286 SIFENWHDLLDFSTRVAKSFKKIHENSNKKGND--IQGI-LGVDSTPNSLF--------- 333
Query: 961 TSLERVNGGDKNGIVYTL 978
TS+ GGD N Y +
Sbjct: 334 TSVFAAGGGDYNNFFYKI 351
>CLB2_STRAW (Q826F2) Chaperone clpB 2
Length = 879
Score = 41.2 bits (95), Expect = 0.014
Identities = 71/324 (21%), Positives = 125/324 (37%), Gaps = 44/324 (13%)
Query: 12 LTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQCRA 71
LT ++ L+ + A R GH ++ H+ LL + QQ P + RA
Sbjct: 6 LTQKSQEALQEAQTAAGRMGHTEVDGEHLLLALLDQEDGLIPRLL---QQAGTEPKELRA 62
Query: 72 LELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQPL 131
L+ P T P P V ++ L L A+ +R L
Sbjct: 63 ---AVREELSHRPKAT-GPGAAPGQV-FVTQRLARLLDAAEREAKR-------------L 104
Query: 132 LSVKVELDQLILSILDDPSVSR---VMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
V ++ L+L++ ++ S + ++++AG + S + L T + + S+
Sbjct: 105 KDEYVSVEHLLLALAEESSSTAAGLLLKQAGITRDSFLSAL----TQVRGNQRVTSANPE 160
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
+++ L YG VL + + V +I V +L RK K N V++G
Sbjct: 161 VAYEA-LEKYGRDLVLEARSGRLDPVI--------GRDAEIRRVTQILSRKTKNNPVLIG 211
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMK-TTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLK 307
D +V + +R RG+VP+ ++ T F G KY + E LK
Sbjct: 212 DPGVGKTAIVEGLAQRIVRGDVPEGLRDKTVFALDMGSLVAGAKYRGEFEER------LK 265
Query: 308 RKVSDYVALGVGAIFYVGDLKWIV 331
+S+ A + +V +L +V
Sbjct: 266 AVLSEVKAAEGRILLFVDELHTVV 289
Score = 34.7 bits (78), Expect = 1.3
Identities = 35/140 (25%), Positives = 62/140 (44%), Gaps = 15/140 (10%)
Query: 632 KVLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQGNDSVGKKRLALAIAESV 686
++L+E + E V + +A++ ++S + +++FL G VGK LA +A ++
Sbjct: 572 EILRERVIGQDEAVKLVTDAIIRARSGIRDPRRPIGSFIFL-GPTGVGKTELAKTLARTL 630
Query: 687 FGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRK----MLAD 742
F S E +DM + + T S + P +V E + + RK +L D
Sbjct: 631 FDSEENMVRLDMSEYQERHT-VSRLMGAP----PGYVGYEEGGQLTEAVRRKPYSVVLFD 685
Query: 743 EFEIAKFGTLGQKIFILSNG 762
E E A + IL +G
Sbjct: 686 EIEKAHTDVFNTLLQILDDG 705
>H104_YEAST (P31539) Heat shock protein 104
Length = 908
Score = 40.8 bits (94), Expect = 0.018
Identities = 58/270 (21%), Positives = 105/270 (38%), Gaps = 45/270 (16%)
Query: 9 QETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSS----LRNACLKSQQQNN 64
Q T A +IL + LA H Q+ P+H+ A + P L+N K + +
Sbjct: 4 QTQFTERALTILTLAQKLASDHQHPQLQPIHILAAFIETPEDGSVPYLQNLIEKGRYDYD 63
Query: 65 HPLQCRALELCFNVALNRLPTTTTSPL-LQPQHVPSLSNALIAALKRAQAHQRRGCIEQN 123
+ N L R+P +P + P + +L L A K
Sbjct: 64 ------LFKKVVNRNLVRIPQQQPAPAEITPSY--ALGKVLQDAAK-------------I 102
Query: 124 QQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNN-LE-NSSTLINSSSV 181
Q+QQ+ ++ D ++ ++ +D S+ ++ +EA ++K LE +T I+S
Sbjct: 103 QKQQKDSFIA----QDHILFALFNDSSIQQIFKEAQVDIEAIKQQALELRGNTRIDSRGA 158
Query: 182 FHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKK 241
++P +LS Y + Q K V +E+I VL R+ K
Sbjct: 159 DTNTPL-----EYLSKYAIDMTEQARQGKLDPVI--------GREEEIRSTIRVLARRIK 205
Query: 242 KNTVIVGDTVSLTEGLVSEIMKRFERGEVP 271
N ++G+ ++ + +R +VP
Sbjct: 206 SNPCLIGEPGIGKTAIIEGVAQRIIDDDVP 235
>CLPC_CHLTR (O84288) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 854
Score = 40.8 bits (94), Expect = 0.018
Identities = 66/319 (20%), Positives = 123/319 (37%), Gaps = 44/319 (13%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T A ++K + A+R H + H+ LL L N
Sbjct: 3 EKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVL------------- 49
Query: 70 RALELCFNVALNRLPTTTT-SPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQ 128
R L + F+ A + + P +Q P+L+ + + + A++ +E N +
Sbjct: 50 RTLGVDFDTAKHEVERLIGYGPEIQVCGDPALTGRVKKSFE--SANEEAALLEHNYVGTE 107
Query: 129 QPLLSVKVELDQLILSILDDPSVS--RVMREAGFSSPSVKNNLENSSTLI----NSSSVF 182
LL + + D + L +L++ V + +E + L SS++ NSSS
Sbjct: 108 HLLLGILNQSDGVALQVLENLHVDPKEIRKEILKELETFNLQLPPSSSITPRNTNSSSSS 167
Query: 183 HSSPSPLSHNHF----------LSSYGYG-SVLFSSQKKEQVVYHPFLKSSESNKEDINL 231
SS SPL + L +YGY + +F + + V+ ++
Sbjct: 168 KSS-SPLGGHTLGGDKPEKLSALKAYGYDLTEMFKESRLDPVI---------GRSAEVER 217
Query: 232 VFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSL 290
+ +L R++K N V+VG+ +V + ++ GEVP+ ++ + L
Sbjct: 218 LILILCRRRKNNPVLVGEAGVGKTAIVEGLAQKIVSGEVPEALRKKRLITLDLALMIAGT 277
Query: 291 KYMKKEEVEMNVIRVLKRK 309
KY + E + + RK
Sbjct: 278 KYRGQFEERIKAVMDEVRK 296
>CLB1_ANASP (Q8YUL9) Chaperone clpB 1
Length = 880
Score = 40.0 bits (92), Expect = 0.032
Identities = 40/178 (22%), Positives = 74/178 (41%), Gaps = 25/178 (14%)
Query: 136 VELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFL 195
+ ++ ++L+ +DD V R + + GF+ S K +E S + S + +P S L
Sbjct: 108 ISVEHMLLAFVDDERVGRKVLK-GFNVDSAK--IEASIKAVRGSQKV-TDQNPESRYEAL 163
Query: 196 SSYGYG-SVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLT 254
+G + S K + V+ ++I V VL R+ K N V++G+
Sbjct: 164 QKFGRDLTEQAKSGKLDPVI---------GRDDEIRRVIQVLSRRSKNNPVLIGEPGVGK 214
Query: 255 EGLVSEIMKRFERGEVPDEMKTTHFV-----------KFHGLSSVSLKYMKKEEVEMN 301
+ + +R G+VP+ +K + K+ G LK + +E E N
Sbjct: 215 TAIAEALAQRIVNGDVPESLKNRQLISLDIGSLIAGAKYRGEFEDRLKAVLREVTESN 272
>CLPB_PSEAE (Q9HVN5) Chaperone clpB
Length = 854
Score = 39.7 bits (91), Expect = 0.041
Identities = 56/267 (20%), Positives = 110/267 (40%), Gaps = 38/267 (14%)
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
+ LT++ L + LA H I P+H+ + LL S++ ++
Sbjct: 4 DRLTSKLQLALSDAQSLAVGHDHPAIEPVHLLSALLEQQGGSIKPLLMQVG------FDI 57
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQ 129
AL N L+ LP + P +LS L L +A ++ QQ+
Sbjct: 58 AALRSGLNKELDALPK-----IQSPTGDVNLSQDLARLLNQA---------DRLAQQKGD 103
Query: 130 PLLSVKVELDQLILSILDDPS-VSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
+S ++ ++L+ +D+ + + +++ G S +++N + N L +V + P+
Sbjct: 104 QFISSEL----VLLAAMDENTRLGKLLLGQGVSRKALENAVAN---LRGGEAV--NDPNV 154
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
L Y + +++ E+ P + ++I VL R+ K N V++G
Sbjct: 155 EESRQALDKY----TVDMTKRAEEGKLDPVI----GRDDEIRRTIQVLQRRTKNNPVLIG 206
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMK 275
+ +V + +R GEVPD +K
Sbjct: 207 EPGVGKTAIVEGLAQRIINGEVPDGLK 233
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 115,469,720
Number of Sequences: 164201
Number of extensions: 5020492
Number of successful extensions: 14445
Number of sequences better than 10.0: 176
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 115
Number of HSP's that attempted gapping in prelim test: 14154
Number of HSP's gapped (non-prelim): 373
length of query: 1009
length of database: 59,974,054
effective HSP length: 120
effective length of query: 889
effective length of database: 40,269,934
effective search space: 35799971326
effective search space used: 35799971326
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC148819.4


