

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
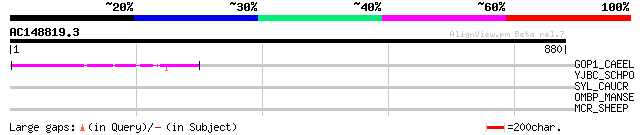
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
GOP1_CAEEL (P46578) Hypothetical protein gop-1 151 8e-36
YJBC_SCHPO (O74949) Hypothetical protein C553.12c in chromosome III 35 0.87
SYL_CAUCR (Q9A217) Leucyl-tRNA synthetase (EC 6.1.1.4) (Leucine-... 33 3.3
OMBP_MANSE (P31420) Ommochrome-binding protein precursor (OBP) (... 32 9.6
MCR_SHEEP (Q9BDJ7) Mineralocorticoid receptor (MR) (Fragment) 32 9.6
>GOP1_CAEEL (P46578) Hypothetical protein gop-1
Length = 892
Score = 151 bits (381), Expect = 8e-36
Identities = 96/308 (31%), Positives = 179/308 (57%), Gaps = 26/308 (8%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQ 62
++EALR+IAE++ +GDQ+D + F+FF+E+Q++ F++I++ T + +QLLQT++I+ +
Sbjct: 44 LVEALRAIAEILIWGDQNDASVFDFFLERQMLLYFLKIMEQGNT-PLNVQLLQTLNILFE 102
Query: 63 NLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRN 122
N++ E ++Y++ SN H+N +I++ FD +N+E+++YYISFL+ + KLN TI N
Sbjct: 103 NIRHETSLYFLLSNNHVNSIISHKFDLQNDEIMAYYISFLKTLSFKLNPATIHFFF---N 159
Query: 123 DEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHT-DYFSNLV 181
+ FPL VE ++ ESM+R AVR + LN+ V DDS+ I + HT +Y S L+
Sbjct: 160 ETTEEFPLLVEVLKLYNWNESMVRIAVRNILLNIVRVQDDSM--IIFAIKHTKEYLSELI 217
Query: 182 SFFRKQCMDLN---RLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERL 238
++++ R L N + VD++ D ++Y +++ DVE
Sbjct: 218 DSLVGLSLEMDTFVRSAENVLAN-----RERLRGKVDDLIDLIHYIGELL-----DVE-A 266
Query: 239 ITDSILMLL----IFPVLLPSLRTVADN-DMQSGVVTSLYLLCCILRIVKIKDLANTIVA 293
+ +S+ +L+ + P+LL S+ DN + +++L+ L IV+ + T ++
Sbjct: 267 VAESLSILVTTRYLSPLLLSSISPRRDNHSLLLTPISALFFFSEFLLIVRHHETIYTFLS 326
Query: 294 ALYYPLES 301
+ + ++
Sbjct: 327 SFLFDTQN 334
>YJBC_SCHPO (O74949) Hypothetical protein C553.12c in chromosome III
Length = 521
Score = 35.0 bits (79), Expect = 0.87
Identities = 41/176 (23%), Positives = 72/176 (40%), Gaps = 20/176 (11%)
Query: 149 VRAVTLNVYHVGDDSVNRYITSEPHTDYFSNLVSFFRKQCMDLNRLISETLKNPGPDSNS 208
V VTL VY VG+ N Y+++ PH+ + + + + L L+S + N S +
Sbjct: 269 VLLVTLLVYIVGEYLTNLYMSTLPHSSTVAIMYVYSWTGTVSLCNLVSSWILNTKTHSYA 328
Query: 209 TVTTAVDEIEDNLYYFSDIISAGIPDVERL----ITDSILMLLIFPVLLPS-----LRTV 259
VT E L + + A + ++ I S+ M+ + P++ S L +
Sbjct: 329 LVTVFKLYFELTLQVYVRNLYARLESPQQFVLVQIASSLTMMSVIPLITMSRTVFRLNVL 388
Query: 260 ADNDMQSGVV------TSLYLLCCILRIVKIKDLANTIVA-----ALYYPLESFTK 304
M S VV + Y+ C + + L +++ A YP SF+K
Sbjct: 389 MTKSMDSYVVYRKNMGRNFYVKCVASNVSMLSFLGWSLILHFGSNAPLYPYFSFSK 444
>SYL_CAUCR (Q9A217) Leucyl-tRNA synthetase (EC 6.1.1.4)
(Leucine--tRNA ligase) (LeuRS)
Length = 861
Score = 33.1 bits (74), Expect = 3.3
Identities = 19/72 (26%), Positives = 37/72 (51%), Gaps = 1/72 (1%)
Query: 397 LQTKELDESMLDGLGILPQRKQQKKLLLQALVGEASGEEQLFSSESSLTRDGIACGLDVY 456
L+ + ++++DGL L + + +L+ + +G + G F + DG+A GL+VY
Sbjct: 193 LRITDYADALIDGLKTLDRWPDKVRLMQENWIGRSKGLRFKFQFDGEAP-DGMAEGLEVY 251
Query: 457 LNKIKEQYGVSF 468
+ +G SF
Sbjct: 252 TTRPDTLFGASF 263
>OMBP_MANSE (P31420) Ommochrome-binding protein precursor (OBP)
(YCP)
Length = 274
Score = 31.6 bits (70), Expect = 9.6
Identities = 25/76 (32%), Positives = 38/76 (49%), Gaps = 9/76 (11%)
Query: 153 TLNVYHVGDDSVNRYITSEPHTDYFSNLVSFFRKQCMD--LNRLISETLKNPGPDSNSTV 210
T V +GD +VN + YFS+ V F+ D +N+LISET G DS +
Sbjct: 194 TNKVIDLGDYNVNAFTKDSKGKLYFSSPVGFYAVNEADRKMNKLISET----GEDS---I 246
Query: 211 TTAVDEIEDNLYYFSD 226
A + +DN+ Y ++
Sbjct: 247 WGAAFDKDDNIVYSNE 262
>MCR_SHEEP (Q9BDJ7) Mineralocorticoid receptor (MR) (Fragment)
Length = 258
Score = 31.6 bits (70), Expect = 9.6
Identities = 13/40 (32%), Positives = 23/40 (57%)
Query: 631 PKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLH 670
P+ + FPP + F+E+ + + F + MH ++ FV LH
Sbjct: 98 PQFLYFPPDLVFNEKKMHHSAMFELCQGMHHILLQFVRLH 137
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 100,421,944
Number of Sequences: 164201
Number of extensions: 4188504
Number of successful extensions: 9643
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 9638
Number of HSP's gapped (non-prelim): 6
length of query: 880
length of database: 59,974,054
effective HSP length: 119
effective length of query: 761
effective length of database: 40,434,135
effective search space: 30770376735
effective search space used: 30770376735
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Medicago: description of AC148819.3


