

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148817.2 - phase: 0
(608 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
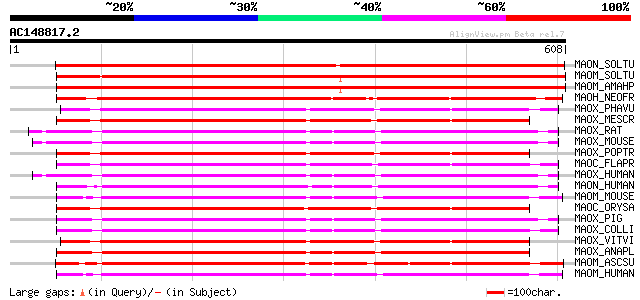
Score E
Sequences producing significant alignments: (bits) Value
MAON_SOLTU (P37225) NAD-dependent malic enzyme 59 kDa isoform, m... 924 0.0
MAOM_SOLTU (P37221) NAD-dependent malic enzyme 62 kDa isoform, m... 749 0.0
MAOM_AMAHP (P37224) NAD-dependent malic enzyme 65 kDa isoform, m... 727 0.0
MAOH_NEOFR (P78715) Malic enzyme, hydrogenosomal precursor (EC 1... 441 e-123
MAOX_PHAVU (P12628) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 427 e-119
MAOX_MESCR (P37223) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 423 e-118
MAOX_RAT (P13697) NADP-dependent malic enzyme (EC 1.1.1.40) (NAD... 420 e-117
MAOX_MOUSE (P06801) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 419 e-116
MAOX_POPTR (P34105) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 417 e-116
MAOC_FLAPR (P36444) NADP-dependent malic enzyme, chloroplast pre... 417 e-116
MAOX_HUMAN (P48163) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 414 e-115
MAON_HUMAN (Q16798) NADP-dependent malic enzyme, mitochondrial p... 414 e-115
MAOM_MOUSE (Q99KE1) NAD-dependent malic enzyme, mitochondrial pr... 413 e-115
MAOC_ORYSA (P43279) NADP-dependent malic enzyme, chloroplast pre... 412 e-114
MAOX_PIG (Q29558) NADP-dependent malic enzyme (EC 1.1.1.40) (NAD... 411 e-114
MAOX_COLLI (P40927) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 411 e-114
MAOX_VITVI (P51615) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 410 e-114
MAOX_ANAPL (P28227) NADP-dependent malic enzyme (EC 1.1.1.40) (N... 410 e-114
MAOM_ASCSU (P27443) NAD-dependent malic enzyme, mitochondrial pr... 409 e-113
MAOM_HUMAN (P23368) NAD-dependent malic enzyme, mitochondrial pr... 408 e-113
>MAON_SOLTU (P37225) NAD-dependent malic enzyme 59 kDa isoform,
mitochondrial precursor (EC 1.1.1.39) (NAD-ME)
Length = 601
Score = 924 bits (2388), Expect = 0.0
Identities = 452/559 (80%), Positives = 510/559 (90%), Gaps = 5/559 (0%)
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GFP+TERDRLGLRGLLPPR+ISF++QYDRFM S+RSLEKNT GQ D +VSL+KWRILNRL
Sbjct: 46 GFPMTERDRLGLRGLLPPRVISFEQQYDRFMESFRSLEKNTEGQPDSVVSLAKWRILNRL 105
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIY 170
HDRNETLYYR LIDNIK+FAPIIYTPTVGLVCQNY GL+RRPRGMYFSAKDKGEMMSMI+
Sbjct: 106 HDRNETLYYRVLIDNIKDFAPIIYTPTVGLVCQNYSGLFRRPRGMYFSAKDKGEMMSMIF 165
Query: 171 NWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTN 230
NWP+ QVDMIVLTDGSRILGLGDLGVQGIGIP+GKLDMYVAAAGINPQR+LP+MLDVGTN
Sbjct: 166 NWPSTQVDMIVLTDGSRILGLGDLGVQGIGIPIGKLDMYVAAAGINPQRVLPVMLDVGTN 225
Query: 231 NQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRY 290
NQKLL+D LYLGLRQPRLEGEEYLSIVDEF+EAVHARWPKA+VQFEDFQ KWAFETL RY
Sbjct: 226 NQKLLEDPLYLGLRQPRLEGEEYLSIVDEFVEAVHARWPKAVVQFEDFQAKWAFETLDRY 285
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R KFCMFNDDIQGTAGVALAGLLGTVRAQGRPL+DFA QKIV+VGAGSAGLGVL MA+QA
Sbjct: 286 RKKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLTDFANQKIVVVGAGSAGLGVLKMALQA 345
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLA--EGASIIEV 408
VSRM+G S A FFL+DKNGL+T +R ++DPAA+PFAK ++EGL EGA + EV
Sbjct: 346 VSRMTGPS---ADPHFFLLDKNGLITKDRKDIDPAALPFAKAHHEIEGLGLQEGAGLAEV 402
Query: 409 VKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGE 468
VKKVKPHVLLGLSGVGG+F+E+VL+AM+ES S +PAIFAMSNPT NAEC +DAF AGE
Sbjct: 403 VKKVKPHVLLGLSGVGGIFHEEVLRAMKESDSVRPAIFAMSNPTNNAECCPVDAFKLAGE 462
Query: 469 NIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLA 528
+IVFASGSPF NVDLGNG+ GHVNQANNMYLFPGIGLG+LLSGA +I+D ML+AA+ECLA
Sbjct: 463 DIVFASGSPFANVDLGNGKIGHVNQANNMYLFPGIGLGALLSGARNISDTMLEAAAECLA 522
Query: 529 SYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDT 588
SYM++++I +GILYPSID IR++TAEVGAAVLRAA+AE LAEGHG VG KEL+HMS+++T
Sbjct: 523 SYMSDDEINRGILYPSIDDIRDITAEVGAAVLRAAVAEDLAEGHGDVGVKELQHMSKEET 582
Query: 589 VEYVRGNMWYPEYCPLVHE 607
+E+VR NMWYP Y PLVHE
Sbjct: 583 IEHVRQNMWYPVYGPLVHE 601
>MAOM_SOLTU (P37221) NAD-dependent malic enzyme 62 kDa isoform,
mitochondrial precursor (EC 1.1.1.39) (NAD-ME)
Length = 626
Score = 749 bits (1935), Expect = 0.0
Identities = 366/563 (65%), Positives = 461/563 (81%), Gaps = 7/563 (1%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTH-GQSDKIVSLSKWRILNRL 110
F TERDRL +RGLLPP ++SF++Q RFM+ + LE G SD V L+KWRILNRL
Sbjct: 64 FSFTERDRLHIRGLLPPNVMSFEQQIARFMADLKRLEVQARDGPSDPYV-LAKWRILNRL 122
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIY 170
HDRNETLYY+ L++NI+E+API+YTPTVGLVCQ Y GL+RRPRGMYFSA+D+GEMMSM+Y
Sbjct: 123 HDRNETLYYKVLMENIEEYAPIVYTPTVGLVCQKYSGLFRRPRGMYFSAEDRGEMMSMVY 182
Query: 171 NWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTN 230
NWPA QVDMIV+TDGSRILGLGDLG+QGIGI +GKLD+YVAAAGINPQR+LP+M+DVGT+
Sbjct: 183 NWPADQVDMIVVTDGSRILGLGDLGIQGIGIAIGKLDLYVAAAGINPQRVLPVMIDVGTD 242
Query: 231 NQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRY 290
N+ LL D LYLGL+ RL+GEEY+ ++DEFMEAV RWP IVQFEDFQ KWAF+ L+RY
Sbjct: 243 NENLLKDPLYLGLQDHRLDGEEYIEVIDEFMEAVFTRWPHVIVQFEDFQSKWAFKLLQRY 302
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R+ + MFNDDIQGTAGVA+AGLLG VRAQGRP+ DF K KIV+ GAGSAG+GVLN A +
Sbjct: 303 RNNYRMFNDDIQGTAGVAIAGLLGAVRAQGRPMIDFPKMKIVVAGAGSAGIGVLNAARKT 362
Query: 351 VSRMSGCSETA---AKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLE--GLAEGASI 405
++RM G +E A A+SQF+++D GL+T R N+DP A PFA+ +++E GL+EGA++
Sbjct: 363 MARMLGNTEIAFESARSQFWVVDAKGLITEARENVDPDARPFARKIKEIERQGLSEGATL 422
Query: 406 IEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNH 465
EVV++VKP VLLGLS GG+F+++VL+A++ S ST+PAIF MSNPT NAECT +AF+
Sbjct: 423 AEVVREVKPDVLLGLSACGGLFSKEVLEALKHSTSTRPAIFPMSNPTRNAECTPEEAFSI 482
Query: 466 AGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASE 525
GENI+FASGSPF++VDLGNG GH NQANNM+LFPGIGLG+LLSG+ ++DGMLQAA+E
Sbjct: 483 LGENIIFASGSPFKDVDLGNGHVGHCNQANNMFLFPGIGLGTLLSGSRIVSDGMLQAAAE 542
Query: 526 CLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSE 585
CLA+Y+TEE++ KGI+YPSI IR++T EV AAV++ AI E LAEG+ + S+EL + E
Sbjct: 543 CLAAYITEEEVLKGIIYPSISRIRDITKEVAAAVVKEAIEEDLAEGYREMDSRELRKLDE 602
Query: 586 DDTVEYVRGNMWYPEYCPLVHEK 608
E+V NMW P+Y LV++K
Sbjct: 603 AQISEFVENNMWSPDYPTLVYKK 625
>MAOM_AMAHP (P37224) NAD-dependent malic enzyme 65 kDa isoform,
mitochondrial precursor (EC 1.1.1.39) (NAD-ME)
Length = 623
Score = 727 bits (1877), Expect = 0.0
Identities = 349/562 (62%), Positives = 454/562 (80%), Gaps = 5/562 (0%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F +TERDRL LRGLLPP +++ ++Q +RF + R LE T L+KWRILNRLH
Sbjct: 61 FSMTERDRLDLRGLLPPNVMTTEQQIERFTADLRVLELTTKDGPSDTYDLAKWRILNRLH 120
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNET++++ LI+NI+E+API+ TPTVGLVCQ + GLYRRPRGMYFS+ D+GEMMSM+YN
Sbjct: 121 DRNETMFFKVLIENIEEYAPIVSTPTVGLVCQKFSGLYRRPRGMYFSSDDRGEMMSMVYN 180
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WPA QVDMIV+TDGSRILGLGDLGV GIG+ +GKLD+YVAAAGINPQR+LP+M+DVGTNN
Sbjct: 181 WPAEQVDMIVVTDGSRILGLGDLGVHGIGVAIGKLDLYVAAAGINPQRVLPVMIDVGTNN 240
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRYR 291
+ LL + LYLGL++ RL+GEEYL+++DEFMEAV RWP IVQFED Q KWA L+RYR
Sbjct: 241 EDLLKNPLYLGLQKKRLDGEEYLAVMDEFMEAVFTRWPNVIVQFEDIQNKWALTLLQRYR 300
Query: 292 HKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAV 351
HK+ FN D+QGT+GVA+AGLLG VRAQGRP+ DF KQKIV+ GAGS+G+GVLN A + +
Sbjct: 301 HKYRTFNVDVQGTSGVAIAGLLGAVRAQGRPMIDFPKQKIVVAGAGSSGVGVLNAARKTM 360
Query: 352 SRMSGCSETA---AKSQFFLIDKNGLVTTERSNLDPAAVPFA--KNPRDLEGLAEGASII 406
+RM G E+A A+SQF+++D GL+T +R+NLDP PFA +N L+GL EGA ++
Sbjct: 361 ARMLGNDESAFDRARSQFWVVDDKGLITEKRANLDPEVQPFAWKENEISLQGLNEGAKLV 420
Query: 407 EVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHA 466
EVV++VKP VLLGLS GG+F+++VL+A+++S ST+PAIFAMSNPT NAECT +AF+
Sbjct: 421 EVVRQVKPDVLLGLSAYGGLFSKEVLEALKDSTSTRPAIFAMSNPTKNAECTPEEAFSIV 480
Query: 467 GENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASEC 526
G+++V+ASGSPF++VDLGNG+ GHVNQ NNMYLFPGIGLG LLSG+ I+D M QAA+E
Sbjct: 481 GDHVVYASGSPFKDVDLGNGKIGHVNQGNNMYLFPGIGLGVLLSGSRIISDSMFQAAAER 540
Query: 527 LASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSED 586
LA YMT+E++ G++YPSI IR++T EV AAV++ A+ E LAEG+ + ++EL+ ++E+
Sbjct: 541 LAGYMTDEEVINGVIYPSISRIRDITKEVAAAVIKEAVEEDLAEGYRDMDARELQKLNEE 600
Query: 587 DTVEYVRGNMWYPEYCPLVHEK 608
+EY+ NMW PEY LV++K
Sbjct: 601 QILEYIEKNMWNPEYPTLVYKK 622
>MAOH_NEOFR (P78715) Malic enzyme, hydrogenosomal precursor (EC
1.1.1.40) (ME)
Length = 592
Score = 441 bits (1134), Expect = e-123
Identities = 242/555 (43%), Positives = 343/555 (61%), Gaps = 29/555 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVS-LSKWRILNRL 110
F E+DRLG+RGL+PPR S + QY R ++ DKI L K+ LN L
Sbjct: 62 FTADEKDRLGIRGLVPPRPQSLEAQYKRCKTNL-----------DKISDPLEKFIYLNHL 110
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIY 170
+RNETLYY+ +++N E APIIYTP VG CQ + ++ + RGMYFS D+G+M ++
Sbjct: 111 QNRNETLYYKMILENFVELAPIIYTPVVGEACQKFHKIFTQTRGMYFSTADRGQMSAVAA 170
Query: 171 NWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTN 230
NWP VD+IV+TDGSRILGLGDLG G+ IP+GKL +YV GINP+ +LPI+LDVGTN
Sbjct: 171 NWPYDDVDVIVVTDGSRILGLGDLGAGGMQIPIGKLTLYVCGGGINPRNVLPIVLDVGTN 230
Query: 231 NQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRY 290
N++LL+D LYLG++ PRL+GEE+ + VDE++ A+ R+PKA++QFEDF M A + L +Y
Sbjct: 231 NKELLNDPLYLGMQHPRLQGEEFHAFVDEWVSAITDRFPKAVIQFEDFMMPNALDLLLKY 290
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+ + CMFNDDIQ T + LA +L T+RA+G +D K+ + +GAGS+G+GV V
Sbjct: 291 KDQICMFNDDIQSTGAITLASVLATMRARGGTFADIKKETFLCLGAGSSGVGVCETIVDC 350
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
+ G + A +QF++ D GL+ R +L P+ F + +++EG G + E++K
Sbjct: 351 IV-AEGATREEAYAQFYMFDHKGLLGKGRDDLLPSQQVFMR--KEIEG---GKTPAELLK 404
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
K+KP LLGLS +FN+++L + S KP IF +SNPT +ECTA +A N+
Sbjct: 405 KIKPTCLLGLSTCPKLFNKEMLSYV-ASYCEKPGIFPLSNPTSRSECTAEEAVEFTDGNL 463
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
+FASGSPF+ V+ G+ NQ NN Y FPGIGLG + S A + Q + +AS
Sbjct: 464 IFASGSPFDPVE-WKGKTIQTNQCNNSYSFPGIGLGLVSSRATRVPFETFQVCARVIASL 522
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVE 590
T E + G ++P +D +R V+ EVG V + A +A G D E
Sbjct: 523 GTPEMLATGKIFPDLDNLRAVSLEVGIEVAKMAERMNIATQFPPKGM---------DWRE 573
Query: 591 YVRGNMWYPEYCPLV 605
+++ NMW PEY +V
Sbjct: 574 WLKHNMWQPEYPHIV 588
>MAOX_PHAVU (P12628) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 589
Score = 427 bits (1097), Expect = e-119
Identities = 234/549 (42%), Positives = 330/549 (59%), Gaps = 35/549 (6%)
Query: 56 ERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNE 115
ERD LRGLLPP + + + Q R M + R E V L ++ L L +RNE
Sbjct: 69 ERDAHYLRGLLPPSVFNQELQEKRLMHNLRQYE----------VPLHRYMALMDLQERNE 118
Query: 116 TLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAP 175
L+Y+ LIDN+ E P++YTPTVG CQ Y ++RRP+G+Y S K+KG+++ ++ NWP
Sbjct: 119 RLFYKLLIDNVAELLPVVYTPTVGEACQKYGSIFRRPQGLYISLKEKGKILEVLKNWPEK 178
Query: 176 QVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLL 235
+ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LP+ +DVGTNN+KLL
Sbjct: 179 SIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSSCLPVTIDVGTNNEKLL 238
Query: 236 DDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKF 294
+D Y+GLRQ R G+EY + +DEFM AV + K +VQFEDF AF+ L++Y
Sbjct: 239 NDEFYIGLRQRRATGQEYATFLDEFMRAVKQNYGEKVLVQFEDFANHNAFDLLEKYSSSH 298
Query: 295 CMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRM 354
+FNDDIQGTA V LAGLL +++ G L+D + +GAG AG G+ + VS+
Sbjct: 299 LVFNDDIQGTASVVLAGLLASLKLVGGTLAD---HTFLFLGAGEAGTGIAELIAVEVSKQ 355
Query: 355 SGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVK 413
+ + + +L+D GL+ + R +L P+A ++GL +E VK +K
Sbjct: 356 TKAPVEETRKKIWLVDSKGLIVSSRLESLQQFKKPWAHEHEPVKGL------LEAVKAIK 409
Query: 414 PHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFA 473
P VL+G SG G F ++V++ M S++ KP I A+SNPT +ECTA +A+ + +FA
Sbjct: 410 PTVLIGSSGAGKTFTKEVVETM-ASLNEKPLILALSNPTSQSECTAEEAYTWSKGRAIFA 468
Query: 474 SGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTE 533
SGSPF+ V+ G+ QANN Y+FPG GLG ++SGA + D ML AASE LA+ ++E
Sbjct: 469 SGSPFDPVEY-EGKLFVPGQANNAYIFPGFGLGLIMSGAIRVRDEMLLAASEALAAQVSE 527
Query: 534 EDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSE-DDTVEYV 592
E+ KG++YP IR ++A + A V A GLA H+ D V+Y
Sbjct: 528 ENYDKGLIYPPFTNIRKISANIAAKVAAKAYDLGLA-----------SHLKRPKDLVKYA 576
Query: 593 RGNMWYPEY 601
M+ P Y
Sbjct: 577 ESCMYSPGY 585
>MAOX_MESCR (P37223) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 585
Score = 423 bits (1087), Expect = e-118
Identities = 229/520 (44%), Positives = 323/520 (62%), Gaps = 23/520 (4%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP ++S + Q +F+++ R + V L K+ + L
Sbjct: 61 FTEKERDAHFLRGLLPPVVLSQELQEKKFLTTLRQYQ----------VPLQKYMAMMDLQ 110
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ L+D+++E P++YTPTVG CQ Y ++RRP+G++ S KDKG ++ ++ N
Sbjct: 111 ERNEKLFYKLLVDHVEELLPLVYTPTVGEGCQKYGSIFRRPQGLFISLKDKGRILELLRN 170
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP ++ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI LDVGTNN
Sbjct: 171 WPEKKIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYSALGGVCPSACLPITLDVGTNN 230
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
QKLLDD Y+GL+Q R GEEY V EFM AV + K +VQFEDF AFE L++Y
Sbjct: 231 QKLLDDEFYIGLKQKRATGEEYAEFVQEFMSAVKQNYGEKILVQFEDFANHNAFELLEKY 290
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R +FNDDIQGTA V LAGL+ +++ G L+D K + +GAG AG G+ +
Sbjct: 291 RTTHLVFNDDIQGTASVVLAGLIASLKLLGGTLAD---HKFLFLGAGEAGTGIAELIALE 347
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+S+ + + + +L+D GLV + R L +P+A + ++I+ V
Sbjct: 348 MSKKTKAPVEQMRKKIWLVDSKGLVVSSRKETLQQFKLPWAHEHEPI------TTLIDAV 401
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
+ +KP VL+G SG G F ++V++AM +++ KP I A+SNPT +ECTA +A+ + +
Sbjct: 402 QAIKPTVLIGTSGKGKQFTKEVVEAM-ANINAKPLILALSNPTSQSECTAEEAYTWSQGH 460
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
+FASGSPF+ V+ GR QANN Y+FPG GLG ++ GA + D ML AASE LAS
Sbjct: 461 AIFASGSPFDPVEY-EGRTFVPGQANNAYIFPGFGLGLIMCGAIRVHDDMLLAASEALAS 519
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
+T E KG++YP IR ++A + A V A GLA
Sbjct: 520 QVTGEHFIKGLIYPPFKDIRKISAHIAAGVAAKAYELGLA 559
>MAOX_RAT (P13697) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME) (Malic enzyme 1)
Length = 572
Score = 420 bits (1079), Expect = e-117
Identities = 230/582 (39%), Positives = 344/582 (58%), Gaps = 35/582 (6%)
Query: 21 DRGCSRRRYLHRVWFINVVLIFFMILGSIRGFPLTERDRLGLRGLLPPRIISFQEQYDRF 80
D RRR+ H+ ++ L L F L ER +L + GLLPP I++ + Q R
Sbjct: 2 DPRAPRRRHTHQRGYL---LTRDPHLNKDLAFTLEERQQLKIHGLLPPCIVNQEIQVLRV 58
Query: 81 MSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNETLYYRALIDNIKEFAPIIYTPTVGL 140
+ ++ L + ++ +L L DRNE L+Y L+ N+++F PI+YTPTVGL
Sbjct: 59 IKNFERLNSD----------FDRYLLLMDLQDRNEKLFYSVLMSNVEKFMPIVYTPTVGL 108
Query: 141 VCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIG 200
CQ Y +R+PRG++ S DKG + S++ WP V IV+TDG RILGLGDLG G+G
Sbjct: 109 ACQQYSLAFRKPRGLFISIHDKGHIASVLNAWPEDVVKAIVVTDGERILGLGDLGCNGMG 168
Query: 201 IPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEF 260
IPVGKL +Y A G+NPQ+ LPI LDVGT N++LL D LY+GLR R+ G EY + +DEF
Sbjct: 169 IPVGKLALYTACGGVNPQQCLPITLDVGTENEELLKDPLYIGLRHRRVRGPEYDAFLDEF 228
Query: 261 MEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQ 319
MEA +++ ++QFEDF AF L +YR+K+C FNDDIQGTA VA+AGLL +R
Sbjct: 229 MEAASSKYGMNCLIQFEDFANLNAFRLLNKYRNKYCTFNDDIQGTASVAVAGLLAALRIT 288
Query: 320 GRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTER 379
LSD Q ++ GAG A LG+ ++ V A+ + G S+ A+ + +L+D GL+ R
Sbjct: 289 KNKLSD---QTVLFQGAGEAALGIAHLIVMAMEK-EGLSKEKARQKIWLVDSKGLIVKGR 344
Query: 380 SNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESV 439
++L FA +++ L +V+K+KP L+G++ +GG F EQ+LK M +
Sbjct: 345 ASLTEEKEVFAHEHEEMKNLE------AIVQKIKPTALIGVAAIGGAFTEQILKDM-AAF 397
Query: 440 STKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYL 499
+ +P IFA+SNPT AEC+A + + +FASGSPF+ V L +GR Q NN Y+
Sbjct: 398 NERPIIFALSNPTSKAECSAEECYKVTKGRAIFASGSPFDPVTLPDGRTLFPGQGNNSYV 457
Query: 500 FPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAV 559
FPG+ LG + G HI D + +E ++ ++++ +++G LYP ++ IR+V+ ++ +
Sbjct: 458 FPGVALGVVACGLRHINDSVFLTTAEVISQQVSDKHLEEGRLYPPLNTIRDVSLKIAVKI 517
Query: 560 LRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEY 601
++ A E +A + +KE E+V M+ Y
Sbjct: 518 VQDAYKEKMATVYPEPQNKE----------EFVSSQMYSTNY 549
>MAOX_MOUSE (P06801) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME) (Malic enzyme 1)
Length = 572
Score = 419 bits (1076), Expect = e-116
Identities = 230/577 (39%), Positives = 342/577 (58%), Gaps = 35/577 (6%)
Query: 26 RRRYLHRVWFINVVLIFFMILGSIRGFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYR 85
RRR+ H+ ++ L L F L ER +L + GLLPP IIS + Q R + ++
Sbjct: 7 RRRHTHQRGYL---LTRDPHLNKDLAFTLEERQQLNIHGLLPPCIISQELQVLRIIKNFE 63
Query: 86 SLEKNTHGQSDKIVSLSKWRILNRLHDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNY 145
L + ++ +L L DRNE L+Y L+ ++++F PI+YTPTVGL CQ Y
Sbjct: 64 RLNSD----------FDRYLLLMDLQDRNEKLFYSVLMSDVEKFMPIVYTPTVGLACQQY 113
Query: 146 CGLYRRPRGMYFSAKDKGEMMSMIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGK 205
+R+PRG++ S DKG + S++ WP V IV+TDG RILGLGDLG G+GIPVGK
Sbjct: 114 SLAFRKPRGLFISIHDKGHIASVLNAWPEDVVKAIVVTDGERILGLGDLGCNGMGIPVGK 173
Query: 206 LDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVH 265
L +Y A G+NPQ+ LPI LDVGT N++LL D LY+GLR R+ G EY + +DEFMEA
Sbjct: 174 LALYTACGGVNPQQCLPITLDVGTENEELLKDPLYIGLRHRRVRGPEYDAFLDEFMEAAS 233
Query: 266 ARW-PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLS 324
+++ ++QFEDF + AF L +YR+K+C FNDDIQGTA VA+AGLL +R LS
Sbjct: 234 SKYGMNCLIQFEDFANRNAFRLLNKYRNKYCTFNDDIQGTASVAVAGLLAALRITKNKLS 293
Query: 325 DFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDP 384
D Q ++ GAG A LG+ ++ V A+ + G S+ A+ + +L+D GL+ R++L
Sbjct: 294 D---QTVLFQGAGEAALGIAHLVVMAMEK-EGLSKENARKKIWLVDSKGLIVKGRASLTE 349
Query: 385 AAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPA 444
FA +++ L +V+K+KP L+G++ +GG F EQ+LK M + + +P
Sbjct: 350 EKEVFAHEHEEMKNLE------AIVQKIKPTALIGVAAIGGAFTEQILKDM-AAFNERPI 402
Query: 445 IFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIG 504
IFA+S+PT AEC+A + + +FASGSPF+ V L +GR Q NN Y+FPG+
Sbjct: 403 IFALSSPTSKAECSADECYKVTKGRAIFASGSPFDPVTLPDGRTLFPGQGNNSYVFPGVA 462
Query: 505 LGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAI 564
LG + G HI D + E ++ ++++ +Q+G LYP ++ IR V+ ++ +++ A
Sbjct: 463 LGVVACGLRHIDDKVFLTTREVISQQVSDKHLQEGRLYPPLNTIRGVSLKIAVKIVQDAY 522
Query: 565 AEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEY 601
E +A + +KE E+V M+ Y
Sbjct: 523 KEKMATVYPEPQNKE----------EFVSSQMYSTNY 549
>MAOX_POPTR (P34105) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 591
Score = 417 bits (1071), Expect = e-116
Identities = 225/520 (43%), Positives = 319/520 (61%), Gaps = 23/520 (4%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP IS Q Q + M++ R + + L K+ + L
Sbjct: 67 FTEKERDAHYLRGLLPPTTISQQLQEKKLMNTIRQYQ----------LPLQKYTAMMELE 116
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ LIDN++E P++YTPTVG CQ Y +++RP+G+Y S K+KG+++ ++ N
Sbjct: 117 ERNERLFYKLLIDNVEELLPVVYTPTVGEACQKYGSIFKRPQGLYISLKEKGKVLDVLKN 176
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +IV+TDG RILGLGDLG QGIGIPVGKL +Y A G+ P LP+ +DVGTNN
Sbjct: 177 WPQKSIQVIVVTDGERILGLGDLGCQGIGIPVGKLSLYTALGGVRPSACLPVTIDVGTNN 236
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
++LL D Y+GLRQ R G+EY ++ EFM AV + K ++QFEDF AF+ L +Y
Sbjct: 237 EQLLKDEFYIGLRQRRATGQEYSELLHEFMTAVKQNYGEKVLIQFEDFANHNAFDLLAKY 296
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+FNDDIQGTA V LAGL+ ++ G L+D + +GAG AG G+ +
Sbjct: 297 GTTHLVFNDDIQGTAAVVLAGLISALKLLGGSLAD---HTFLFLGAGEAGTGIAELIALE 353
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+SR S + + +L D GL+ + R +L P+A ++GL +EVV
Sbjct: 354 MSRRSKTPLEETRKKIWLTDSKGLIVSSRKESLQHFKKPWAHEHEPVKGL------LEVV 407
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
K +KP VL+G SGVG F ++V++AM S + KP I A+SNPT +ECTA +A+
Sbjct: 408 KAIKPIVLIGTSGVGKTFTKEVIEAM-ASFNEKPLILALSNPTSQSECTAQEAYTWTKGK 466
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
+FASGSPF+ V+ G+ Q+NN Y+FPG+GLG ++SGA + D ML AA+E LA
Sbjct: 467 AIFASGSPFDPVEY-EGKVFVPGQSNNAYIFPGLGLGLVISGAIRVHDDMLLAAAEALAG 525
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
+ EE + KG++YP + IR ++ ++ A V A GLA
Sbjct: 526 QIKEEYLAKGLIYPPLSNIRKISVQIAANVAAKAYELGLA 565
>MAOC_FLAPR (P36444) NADP-dependent malic enzyme, chloroplast
precursor (EC 1.1.1.40) (NADP-ME)
Length = 647
Score = 417 bits (1071), Expect = e-116
Identities = 227/552 (41%), Positives = 326/552 (58%), Gaps = 33/552 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP +++ Q + M + R E V L +++ + L
Sbjct: 123 FTEKERDAHYLRGLLPPVVVNHDLQVKKMMHNIRQYE----------VPLQRYQAMMDLQ 172
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ LI+NI+E PI+YTPTVG CQ Y +++ P+G+Y S KDKG+++ ++ N
Sbjct: 173 ERNERLFYKLLIENIEELLPIVYTPTVGEACQKYGTIFKNPQGLYISLKDKGKVLEILKN 232
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP ++ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A GI P LPI +DVGTNN
Sbjct: 233 WPQKKIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGIRPSACLPITIDVGTNN 292
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+K+L+D Y+GLRQ R G+EY +++EFM AV + K ++QFEDF AF+ L++Y
Sbjct: 293 EKMLNDEFYIGLRQRRASGKEYAELMNEFMSAVKQNYGEKVLIQFEDFANHNAFDLLEKY 352
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R +FNDDIQGTA V LAGL+ ++ G L+D K + +GAG AG G+ +
Sbjct: 353 RTTHLVFNDDIQGTASVVLAGLISALKLVGGSLAD---HKFLFLGAGEAGTGIAELIALE 409
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+S+ + + + +L+D GL+ R +L P+A + + ++ V
Sbjct: 410 ISKQTNAPLEETRKKIWLVDSKGLIVRSRLDSLQHFKKPWAHDHEPVN------KFLDAV 463
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
K +KP VL+G SG G F ++V++AM S + KP I A+SNPT +ECTA A+ +
Sbjct: 464 KAIKPTVLIGSSGAGQTFTKEVVEAM-SSFNEKPIILALSNPTSQSECTAEQAYTWSEGR 522
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
+FASGSPF V+ NG+ Q+NN Y+FPG GLG ++SGA + D ML AASE LA
Sbjct: 523 TIFASGSPFAPVEY-NGKVYVSGQSNNAYIFPGFGLGLIISGAIRVHDEMLLAASEALAE 581
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTV 589
+T+E G++YP IR ++A + A V A GLA ++ V
Sbjct: 582 QVTQEHFDNGLIYPPFTNIRKISAHIAAKVAAKAYELGLAS----------RLPQPENLV 631
Query: 590 EYVRGNMWYPEY 601
Y M+ P+Y
Sbjct: 632 AYAESCMYSPKY 643
>MAOX_HUMAN (P48163) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME) (Malic enzyme 1)
Length = 572
Score = 414 bits (1063), Expect = e-115
Identities = 227/577 (39%), Positives = 342/577 (58%), Gaps = 35/577 (6%)
Query: 26 RRRYLHRVWFINVVLIFFMILGSIRGFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYR 85
RRR+ H+ ++ L L F L ER +L + GLLPP S + Q R + ++
Sbjct: 7 RRRHTHQRGYL---LTRNPHLNKDLAFTLEERQQLNIHGLLPPSFNSQEIQVLRVVKNFE 63
Query: 86 SLEKNTHGQSDKIVSLSKWRILNRLHDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNY 145
L + ++ +L L DRNE L+YR L +I++F PI+YTPTVGL CQ Y
Sbjct: 64 HLNSD----------FDRYLLLMDLQDRNEKLFYRVLTSDIEKFMPIVYTPTVGLACQQY 113
Query: 146 CGLYRRPRGMYFSAKDKGEMMSMIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGK 205
++R+PRG++ + D+G + S++ WP + IV+TDG RILGLGDLG G+GIPVGK
Sbjct: 114 SLVFRKPRGLFITIHDRGHIASVLNAWPEDVIKAIVVTDGERILGLGDLGCNGMGIPVGK 173
Query: 206 LDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVH 265
L +Y A G+NPQ LP++LDVGT N++LL D LY+GLRQ R+ G EY +DEFMEAV
Sbjct: 174 LALYTACGGMNPQECLPVILDVGTENEELLKDPLYIGLRQRRVRGSEYDDFLDEFMEAVS 233
Query: 266 ARW-PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLS 324
+++ ++QFEDF AF L +YR+++C FNDDIQGTA VA+AGLL +R LS
Sbjct: 234 SKYGMNCLIQFEDFANVNAFRLLNKYRNQYCTFNDDIQGTASVAVAGLLAALRITKNKLS 293
Query: 325 DFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDP 384
D Q I+ GAG A LG+ ++ V A+ + G + A + +L+D GL+ R++L
Sbjct: 294 D---QTILFQGAGEAALGIAHLIVMALEK-EGLPKEKAIKKIWLVDSKGLIVKGRASLTQ 349
Query: 385 AAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPA 444
FA +++ L +V+++KP L+G++ +GG F+EQ+LK M + + +P
Sbjct: 350 EKEKFAHEHEEMKNLE------AIVQEIKPTALIGVAAIGGAFSEQILKDM-AAFNERPI 402
Query: 445 IFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIG 504
IFA+SNPT AEC+A + +FASGSPF+ V L NG+ + Q NN Y+FPG+
Sbjct: 403 IFALSNPTSKAECSAEQCYKITKGRAIFASGSPFDPVTLPNGQTLYPGQGNNSYVFPGVA 462
Query: 505 LGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAI 564
LG + G ITD + +E +A ++++ +++G LYP ++ IR+V+ ++ +++ A
Sbjct: 463 LGVVACGLRQITDNIFLTTAEVIAQQVSDKHLEEGRLYPPLNTIRDVSLKIAEKIVKDAY 522
Query: 565 AEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEY 601
E A + +KE +VR M+ +Y
Sbjct: 523 QEKTATVYPEPQNKE----------AFVRSQMYSTDY 549
>MAON_HUMAN (Q16798) NADP-dependent malic enzyme, mitochondrial
precursor (EC 1.1.1.40) (NADP-ME) (Malic enzyme 3)
Length = 604
Score = 414 bits (1063), Expect = e-115
Identities = 227/551 (41%), Positives = 331/551 (59%), Gaps = 32/551 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER +LG+ GL+PP +S Q R M Y QSD L K+ IL L
Sbjct: 65 FTLEERLQLGIHGLIPPCFLSQDVQLLRIMRYYE------RQQSD----LDKYIILMTLQ 114
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNE L+YR L ++++F PI+YTPTVGL CQ+Y +RRPRG++ + DKG + +M+ +
Sbjct: 115 DRNEKLFYRVLTSDVEKFMPIVYTPTVGLACQHYGLTFRRPRGLFITIHDKGHLATMLNS 174
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +V+TDG RILGLGDLG G+GIPVGKL +Y A G+NPQ+ LP++LDVGTNN
Sbjct: 175 WPEDNIKAVVVTDGERILGLGDLGCYGMGIPVGKLALYTACGGVNPQQCLPVLLDVGTNN 234
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWP-KAIVQFEDFQMKWAFETLKRY 290
++LL D LY+GL+ R+ G+ Y ++DEFM+AV ++ ++QFEDF AF L +Y
Sbjct: 235 EELLRDPLYIGLKHQRVHGKAYDDLLDEFMQAVTDKFGINCLIQFEDFANANAFRLLNKY 294
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R+K+CMFNDDIQGTA VA+AG+L +R LS+ V GAG A +G+ ++ V A
Sbjct: 295 RNKYCMFNDDIQGTASVAVAGILAALRITNNKLSNHV---FVFQGAGEAAMGIAHLLVMA 351
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
+ + G + A + +++D GL+ RS+L+ FA++ ++ S+ EVV+
Sbjct: 352 LEK-EGVPKAEATRKIWMVDSKGLIVKGRSHLNHEKEMFAQDHPEVN------SLEEVVR 404
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
VKP ++G++ + G F EQ+L+ M S +P IFA+SNPT AECTA +
Sbjct: 405 LVKPTAIIGVAAIAGAFTEQILRDM-ASFHERPIIFALSNPTSKAECTAEKCYRVTEGRG 463
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
+FASGSPF++V L +G+ Q NN Y+FPG+ LG + G HI D + +E +A
Sbjct: 464 IFASGSPFKSVTLEDGKTFIPGQGNNAYVFPGVALGVIAGGIRHIPDEIFLLTAEQIAQE 523
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVE 590
++E+ + +G LYP + IR+V+ + VL A LA + KE
Sbjct: 524 VSEQHLSQGRLYPPLSTIRDVSLRIAIKVLDYAYKHNLASYYPEPKDKE----------A 573
Query: 591 YVRGNMWYPEY 601
+VR ++ P+Y
Sbjct: 574 FVRSLVYTPDY 584
>MAOM_MOUSE (Q99KE1) NAD-dependent malic enzyme, mitochondrial
precursor (EC 1.1.1.38) (NAD-ME) (Malic enzyme 2)
Length = 589
Score = 413 bits (1061), Expect = e-115
Identities = 233/557 (41%), Positives = 337/557 (59%), Gaps = 33/557 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER LGL+GLLPP+I + Q RF +R+L+K T L K+ + +
Sbjct: 40 FTLQERQMLGLQGLLPPKIETQDIQALRF---HRNLKKMTS-------PLEKYIYIMGIQ 89
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+YR L D+I+ PI+YTPTVGL C Y ++RRP+G++ S D+G + S++ N
Sbjct: 90 ERNEKLFYRILQDDIESLMPIVYTPTVGLACCQYGHIFRRPKGLFISISDRGHVRSIVDN 149
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP V +V+TDG RILGLGDLGV G+GIPVGKL +Y A AGI P++ LP+ +DVGT+N
Sbjct: 150 WPENHVKAVVVTDGERILGLGDLGVYGMGIPVGKLCLYTACAGIQPEKCLPVCIDVGTDN 209
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPK-AIVQFEDFQMKWAFETLKRY 290
LL D Y+GL Q R + Y ++DEFM+A+ R+ + ++QFEDF AF L++Y
Sbjct: 210 MALLKDPFYMGLYQKRDRSQLYDDLMDEFMKAITDRYGRNTLIQFEDFGNHNAFRFLRKY 269
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+ K+C FNDDIQGTA VAL+GLL R +P+S+ KI+ +GAG A LG+ N+ V +
Sbjct: 270 QQKYCTFNDDIQGTAAVALSGLLAAQRVINKPVSE---HKILFLGAGEAALGIANLIVLS 326
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFA-KNPRDLEGLAEGASIIEV 408
+ SG SE A+ + ++ DK+GL+ R +++D P+A P + E A
Sbjct: 327 MVE-SGLSEEEAQRKIWMFDKSGLLVKGRTASIDSNQEPYAHAAPESIPATFEDA----- 380
Query: 409 VKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGE 468
V K+KP V++G++G G +F V+KAM S++ +P IFA+SNPT AECTA DA+
Sbjct: 381 VNKLKPSVIIGVAGAGPLFTHGVIKAM-ASINERPIIFALSNPTAQAECTAEDAYTLTEG 439
Query: 469 NIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLA 528
+FASGSPFE V L +GR Q NN Y+FPG+ L +L A HI+D + A++ L
Sbjct: 440 RCLFASGSPFEPVKLQDGRVFTPGQGNNAYIFPGVALAVILCEARHISDTVFLEAAKALT 499
Query: 529 SYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDT 588
+ +T+ ++ +G LYPS+ I+ V+A + + A +A + +D
Sbjct: 500 TQLTDAELAQGRLYPSLANIQEVSANIAIKLAEYLYANKMA----------FRYPEPEDK 549
Query: 589 VEYVRGNMWYPEYCPLV 605
YVR +W Y L+
Sbjct: 550 ARYVRERIWRSNYVSLL 566
>MAOC_ORYSA (P43279) NADP-dependent malic enzyme, chloroplast
precursor (EC 1.1.1.40) (NADP-ME)
Length = 638
Score = 412 bits (1059), Expect = e-114
Identities = 219/520 (42%), Positives = 321/520 (61%), Gaps = 23/520 (4%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP ++S Q + M + R V L ++ + L
Sbjct: 114 FSEKERDAHYLRGLLPPAVVSQDLQVKKIMHNLRQYS----------VPLQRYMAMMDLQ 163
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ LIDN++E P++YTPTVG CQ Y ++R+P+G+Y S KDKG+++ ++ N
Sbjct: 164 ERNERLFYKLLIDNVEELLPVVYTPTVGEACQKYGSIFRQPQGLYVSLKDKGKVLDVLRN 223
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI +DVGTNN
Sbjct: 224 WPERNIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSACLPITIDVGTNN 283
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
++LL+D Y+GLRQ R G+EY +++EFM AV + K ++QFEDF AF+ L +Y
Sbjct: 284 EQLLNDEFYIGLRQRRATGKEYHELMEEFMSAVKQIYGEKVLIQFEDFANHNAFDLLAKY 343
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+FNDDIQGTA V LAGLL +++ G L A+ + +GAG AG G+ +
Sbjct: 344 SKSHLVFNDDIQGTASVVLAGLLSSLKVVGGTL---AEHTYLFLGAGEAGTGIAELIALE 400
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+S+ + + + +L+D GL+ R +L P+A + ++++ V
Sbjct: 401 ISKQTKAPIEECRKKVWLLDSKGLIVNSRKESLQAFKKPWAHEHEPV------TTLLDAV 454
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
+ +KP VL+G SGVG F ++V++AM S + +P IF+++NPT ++ECTA +A+N +
Sbjct: 455 QSIKPTVLIGTSGVGKTFTKEVIEAM-ASFNERPVIFSLANPTSHSECTAEEAYNWSQGR 513
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
VFASGSPF+ V+ NG+ Q+NN Y+FPG GLG ++SGA + + ML AASE LA
Sbjct: 514 AVFASGSPFDPVEY-NGKIHVPGQSNNAYIFPGFGLGVVISGAVRVHEDMLLAASETLAD 572
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
T+E+ +KG ++P IR ++A + A V A GLA
Sbjct: 573 QATQENFEKGSIFPPFTNIRKISARIAATVAAKAYELGLA 612
>MAOX_PIG (Q29558) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME) (Malic enzyme 1) (Fragment)
Length = 557
Score = 411 bits (1057), Expect = e-114
Identities = 221/551 (40%), Positives = 331/551 (59%), Gaps = 32/551 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER +L + GLLPP IS Q R + ++ L + ++ +L L
Sbjct: 16 FTLEERQQLNIHGLLPPCFISQDIQVLRVIKNFERLNSD----------FDRYLLLMDLQ 65
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNE L+Y+ L+ +I++F PI+YTPTVGL CQ Y +R+PRG++ S D+G + S++
Sbjct: 66 DRNEKLFYKVLMSDIEKFMPIVYTPTVGLACQQYSLAFRKPRGLFISIHDRGHVASVLNA 125
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +V+TDG RILGLGDLG G+GIPVGKL +Y A G+NPQ LP++LDVGT N
Sbjct: 126 WPEDVIKAVVVTDGERILGLGDLGCNGMGIPVGKLALYTACGGVNPQECLPVILDVGTEN 185
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
++LL D LY+GLRQ R+ G EY +DEFMEAV +++ ++QFEDF AF LK+Y
Sbjct: 186 EELLKDPLYIGLRQRRVRGPEYDDFLDEFMEAVSSKYGMNCLIQFEDFANINAFRLLKKY 245
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
++++C FNDDIQGTA VA+AG+L +R LSD Q I+ GAG A LG+ ++ V A
Sbjct: 246 QNQYCTFNDDIQGTASVAVAGILAALRITKNKLSD---QTILFQGAGEAALGIAHLIVMA 302
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
+ + G + A + +L+D GL+ R+ L FA +++ L +V+
Sbjct: 303 MEK-EGVPKEKAIKKIWLVDSKGLIVKGRAALTNEKEEFAHEHEEMKNLE------AIVQ 355
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
+KP L+G++ +GG F+EQ+LK M + + +P IFA+SNPT AECTA +
Sbjct: 356 DIKPTALIGVAAIGGAFSEQILKDM-AAFNERPIIFALSNPTSKAECTAERGYTLTQGRA 414
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
+FASGSPF+ V L +G+ + Q NN Y+FPG+ L + G HITD + +E +A
Sbjct: 415 IFASGSPFDPVTLPSGQTLYPGQGNNSYVFPGVALAVVACGLRHITDKIFLTTAEVIAQQ 474
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVE 590
++++ +++G LYP ++ IR+V+ ++ ++R A E A + +KE
Sbjct: 475 VSDKHLEEGRLYPPLNTIRDVSLKIAEKIVRDAYQEKTATIYPEPSNKE----------A 524
Query: 591 YVRGNMWYPEY 601
+VR M+ +Y
Sbjct: 525 FVRSQMYSTDY 535
>MAOX_COLLI (P40927) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 557
Score = 411 bits (1057), Expect = e-114
Identities = 217/551 (39%), Positives = 328/551 (59%), Gaps = 32/551 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER +L + GLLPP + Q + ++ L + L ++ +L L
Sbjct: 19 FTLEERQQLNIHGLLPPCFLGQDAQVYSILKNFERLTSD----------LDRYILLMSLQ 68
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNE L+Y+ L +I+ F PI+YTPTVGL CQ+Y +RRPRG++ + D+G + +M+ +
Sbjct: 69 DRNEKLFYKVLTSDIERFMPIVYTPTVGLACQHYGLAFRRPRGLFITIHDRGHIATMLQS 128
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + IV+TDG RILGLGDLG G+GIPVGKL +Y A G+ P + LP+MLDVGT+N
Sbjct: 129 WPESVIKAIVVTDGERILGLGDLGCYGMGIPVGKLALYTACGGVKPHQCLPVMLDVGTDN 188
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+ LL D LY+GLR R+ G+ Y ++DEFMEAV +R+ ++QFEDF AF L +Y
Sbjct: 189 ETLLKDPLYIGLRHKRIRGQAYDDLLDEFMEAVTSRYGMNCLIQFEDFANANAFRLLHKY 248
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R+K+C FNDDIQGTA VA+AGLL +R LSD ++ GAG A LG+ N+ V A
Sbjct: 249 RNKYCTFNDDIQGTASVAVAGLLAALRITKNRLSD---HTVLFQGAGEAALGIANLIVMA 305
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
+ + G S+ A + +++D GL+ R++L P FA +++ L ++VK
Sbjct: 306 MQK-EGVSKEEAIKRIWMVDSKGLIVKGRASLTPEKEHFAHEHCEMKNLE------DIVK 358
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
+KP VL+G++ +GG F +Q+L+ M + + +P IFA+SNPT AECTA + +
Sbjct: 359 DIKPTVLIGVAAIGGAFTQQILQDM-AAFNKRPIIFALSNPTSKAECTAEQLYKYTEGRG 417
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
+FASGSPF+ V L +G+ + Q NN Y+FPG+ LG + G HI D + +E +A
Sbjct: 418 IFASGSPFDPVTLPSGQTLYPGQGNNSYVFPGVALGVISCGLKHIGDDVFLTTAEVIAQE 477
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVE 590
++EE++Q+G LYP + I+ V+ ++ + + A A + +D
Sbjct: 478 VSEENLQEGRLYPPLVTIQQVSLKIAVRIAKEAYRNNTAS----------TYPQPEDLEA 527
Query: 591 YVRGNMWYPEY 601
++R ++ +Y
Sbjct: 528 FIRSQVYSTDY 538
>MAOX_VITVI (P51615) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 591
Score = 410 bits (1055), Expect = e-114
Identities = 220/516 (42%), Positives = 317/516 (60%), Gaps = 23/516 (4%)
Query: 56 ERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNE 115
ERD L GLLPP + + + Q + M+S R + V L K+ + L +RNE
Sbjct: 71 ERDAHYLCGLLPPVVSTQELQERKLMNSIRQYQ----------VPLQKYMAMMDLQERNE 120
Query: 116 TLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAP 175
L+Y+ LIDN++E P++YTPTVG CQ Y ++RRP+G+Y S K+KG+++ ++ NWP
Sbjct: 121 RLFYKLLIDNVEELLPVVYTPTVGEACQKYGSIFRRPQGLYISLKEKGKILEVLKNWPER 180
Query: 176 QVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLL 235
++ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI +DVGTNN+KLL
Sbjct: 181 RIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSACLPITIDVGTNNEKLL 240
Query: 236 DDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKF 294
+ Y+GL+Q R G+EY + EFM V + K ++QFEDF AF+ L +Y
Sbjct: 241 ANEFYIGLKQRRATGKEYSEFLQEFMSPVKQNYGEKVLIQFEDFANHNAFDLLAKYGTTH 300
Query: 295 CMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRM 354
FNDDIQGTA V LAG++ +R G L+D K + +GAG AG G+ + +S+
Sbjct: 301 LAFNDDIQGTASVVLAGIVSALRLLGGTLAD---HKFLFLGAGEAGTGIAELIALEMSKQ 357
Query: 355 SGCSETAAKSQFFLIDKNGLVT-TERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVK 413
+ C + + +L+D GL+ + + +L P+A ++ L ++ VK +K
Sbjct: 358 TKCPIEETRKKIWLVDSKGLIVGSRKDSLQQFKKPWAHEHEPVKDL------LDAVKVIK 411
Query: 414 PHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFA 473
P VL+G SGVG F ++V++AM S + KP I A+SNPT +ECTA +A+ +FA
Sbjct: 412 PTVLIGSSGVGKAFTKEVIEAM-ASCNEKPLILALSNPTSQSECTAEEAYTWTQGRAIFA 470
Query: 474 SGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTE 533
SGSPF+ V+ NG+ QANN Y+FPG+G+G ++SGA + D ML AASE LA +T+
Sbjct: 471 SGSPFDPVEY-NGKTFVPGQANNAYIFPGLGMGLVISGAIRVHDEMLLAASEALARQVTQ 529
Query: 534 EDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
E+ KG++YP IR ++A + A V A GLA
Sbjct: 530 ENFDKGLIYPPFSNIRKISAHIAANVAAKAYELGLA 565
>MAOX_ANAPL (P28227) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME)
Length = 557
Score = 410 bits (1055), Expect = e-114
Identities = 213/519 (41%), Positives = 316/519 (60%), Gaps = 22/519 (4%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER +L + GLLPP + Q + ++ L + L ++ +L L
Sbjct: 19 FTLEERQQLNIHGLLPPCFLGQDVQVFSILKNFERLTSD----------LDRYILLMSLQ 68
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNE L+Y+ L +I+ F PI+YTPTVGL CQ Y +RRPRG++ + D+G + +M+ +
Sbjct: 69 DRNEKLFYKVLTSDIERFMPIVYTPTVGLACQQYGLAFRRPRGLFITIHDRGHIATMLKS 128
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + IV+TDG RILGLGDLG G+GIPVGKL +Y A G+ P LP+MLDVGT+N
Sbjct: 129 WPESVIKAIVVTDGERILGLGDLGCYGMGIPVGKLALYTACGGVKPHECLPVMLDVGTDN 188
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+ LL D LY+GLR R+ G+ Y ++DEFMEAV +R+ ++QFEDF AF L +Y
Sbjct: 189 EALLKDPLYIGLRHKRIRGQAYDDLLDEFMEAVTSRYGMNCLIQFEDFANANAFRLLHKY 248
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R+K+C FNDDIQGTA VA+AGLL +R LSD ++ GAG A LG+ N+ V A
Sbjct: 249 RNKYCTFNDDIQGTASVAVAGLLAALRITKNRLSD---HTVLFQGAGEAALGIANLIVMA 305
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
+ + G S+ AA + +++D GL+ R++L FA +++ L ++VK
Sbjct: 306 MEK-EGVSKEAAVKRIWMVDSKGLIVKGRASLTAEKTRFAHEHAEMKNLE------DIVK 358
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
+KP VL+G++ +GG F +++L+ M + + +P IFA+SNPT AECTA + +
Sbjct: 359 DIKPSVLIGVAAIGGAFTKEILQDM-AAFNKRPIIFALSNPTSKAECTAEQCYKYTEGRG 417
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
+FASGSPF+ V L NG+ + Q NN Y+FPG+ LG + G HI + + +E +A
Sbjct: 418 IFASGSPFDPVTLPNGKTLYPGQGNNSYVFPGVALGVIACGLKHIGEDVFLTTAEVIAEQ 477
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
++EE++Q+G LYP + I++V+ ++ + A A
Sbjct: 478 VSEENLQEGRLYPPLVTIQHVSLKIAVRIAEEAYRNNTA 516
>MAOM_ASCSU (P27443) NAD-dependent malic enzyme, mitochondrial
precursor (EC 1.1.1.38) (NAD-ME) (Fragment)
Length = 643
Score = 409 bits (1051), Expect = e-113
Identities = 226/559 (40%), Positives = 339/559 (60%), Gaps = 35/559 (6%)
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GF L ER LGL GLLPP ++ ++Q +YR + K +D L+++ L+ L
Sbjct: 91 GFSLYERQYLGLHGLLPPAFMTQEQQ------AYRVITKLREQPND----LARYIQLDGL 140
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKG--EMMSM 168
DRNE L+YR + D++KE PI+YTPTVGL CQN+ +YR+P+G+Y + D ++ +
Sbjct: 141 QDRNEKLFYRVVCDHVKELMPIVYTPTVGLACQNFGYIYRKPKGLYITINDNSVSKIYQI 200
Query: 169 IYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVG 228
+ NW V IV+TDG RILGLGDLG GIGIPVGKL +YVA G+ P+ LP++LDVG
Sbjct: 201 LSNWHEEDVRAIVVTDGERILGLGDLGAYGIGIPVGKLALYVALGGVQPKWCLPVLLDVG 260
Query: 229 TNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETL 287
TNN LL+D Y+GLR R+ G++Y +++D FM+A ++ K ++QFEDF AF L
Sbjct: 261 TNNMDLLNDPFYIGLRHKRVRGKDYDTLLDNFMKACTKKYGQKTLIQFEDFANPNAFRLL 320
Query: 288 KRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMA 347
+Y+ K+ MFNDDIQGTA V +AGLL R + +S ++K + GAG+A G+ M
Sbjct: 321 DKYQDKYTMFNDDIQGTASVIVAGLLTCTRVTKKLVS---QEKYLFFGAGAASTGIAEMI 377
Query: 348 VQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIE 407
V + G S+ A ++ +L+D +GLVT R ++P V FAK+ + E SI+E
Sbjct: 378 VHQMQN-EGISKEEACNRIYLMDIDGLVTKNRKEMNPRHVQFAKD------MPETTSILE 430
Query: 408 VVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAG 467
V++ +P L+G S V G FNE+V++AM E ++ +P IFA+SNPT AECTA +A+
Sbjct: 431 VIRAARPGALIGASTVRGAFNEEVIRAMAE-INERPIIFALSNPTSKAECTAEEAYTFTN 489
Query: 468 ENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECL 527
++ASGSPF N +L NG Q NN Y+FPG+ LG++L H+ + + A++ +
Sbjct: 490 GAALYASGSPFPNFEL-NGHTYKPGQGNNAYIFPGVALGTILFQIRHVDNDLFLLAAKKV 548
Query: 528 ASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDD 587
AS +TE+ ++ G +YP + IR ++ ++ + + G A + +D
Sbjct: 549 ASCVTEDSLKVGRVYPQLKEIREISIQIAVEMAKYCYKNGTAN----------LYPQPED 598
Query: 588 TVEYVRGNMWYPEYCPLVH 606
+YVR ++ EY L++
Sbjct: 599 LEKYVRAQVYNTEYEELIN 617
>MAOM_HUMAN (P23368) NAD-dependent malic enzyme, mitochondrial
precursor (EC 1.1.1.38) (NAD-ME) (Malic enzyme 2)
Length = 584
Score = 408 bits (1048), Expect = e-113
Identities = 229/557 (41%), Positives = 332/557 (59%), Gaps = 33/557 (5%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F L ER LGL+GLLPP+I + Q RF +R+L+K T L K+ + +
Sbjct: 40 FTLQERQMLGLQGLLPPKIETQDIQALRF---HRNLKKMTS-------PLEKYIYIMGIQ 89
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+YR L D+I+ PI+YTPTVGL C Y ++RRP+G++ S D+G + S++ N
Sbjct: 90 ERNEKLFYRILQDDIESLMPIVYTPTVGLACSQYGHIFRRPKGLFISISDRGHVRSIVDN 149
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP V +V+TDG RILGLGDLGV G+GIPVGKL +Y A AGI P R LP+ +DVGT+N
Sbjct: 150 WPENHVKAVVVTDGERILGLGDLGVYGMGIPVGKLCLYTACAGIRPDRCLPVCIDVGTDN 209
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPK-AIVQFEDFQMKWAFETLKRY 290
LL D Y+GL Q R ++Y ++DEFM+A+ R+ + ++QFEDF AF L++Y
Sbjct: 210 IALLKDPFYMGLYQKRDRTQQYDDLIDEFMKAITDRYGRNTLIQFEDFGNHNAFRFLRKY 269
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R K+C FNDDIQGTA VALAGLL + +P+S+ KI+ +GAG A LG+ N+ V +
Sbjct: 270 REKYCTFNDDIQGTAAVALAGLLAAQKVISKPISE---HKILFLGAGEAALGIANLIVMS 326
Query: 351 VSRMSGCSETAAKSQFFLIDKNG-LVTTERSNLDPAAVPFAKN-PRDLEGLAEGASIIEV 408
+ +G SE A+ + ++ DK G LV ++ +D PF + P + E A
Sbjct: 327 MVE-NGLSEQEAQKKIWMFDKYGLLVKGRKAKIDSYQEPFTHSAPESIPDTFEDA----- 380
Query: 409 VKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGE 468
V +KP ++G++G G +F V++AM S++ +P IFA+SNPT AECTA +A+
Sbjct: 381 VNILKPSTIIGVAGAGRLFTPDVIRAM-ASINERPVIFALSNPTAQAECTAEEAYTLTEG 439
Query: 469 NIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLA 528
+FASGSPF V L +GR Q NN+Y+FPG+ L +L HI+D + A++ L
Sbjct: 440 RCLFASGSPFGPVKLTDGRVFTPGQGNNVYIFPGVALAVILCNTRHISDSVFLEAAKALT 499
Query: 529 SYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDT 588
S +T+E++ +G LYP + I+ V+ + V A +A + +D
Sbjct: 500 SQLTDEELAQGRLYPPLANIQEVSINIAIKVTEYLYANKMA----------FRYPEPEDK 549
Query: 589 VEYVRGNMWYPEYCPLV 605
+YV+ W EY L+
Sbjct: 550 AKYVKERTWRSEYDSLL 566
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 72,240,985
Number of Sequences: 164201
Number of extensions: 3161043
Number of successful extensions: 7926
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 7668
Number of HSP's gapped (non-prelim): 49
length of query: 608
length of database: 59,974,054
effective HSP length: 116
effective length of query: 492
effective length of database: 40,926,738
effective search space: 20135955096
effective search space used: 20135955096
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC148817.2


