

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
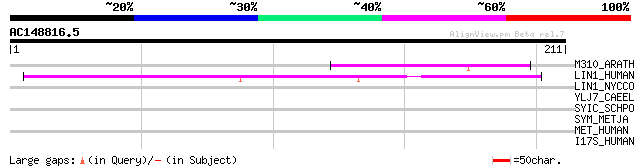
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 51 2e-06
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 48 1e-05
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 41 0.002
YLJ7_CAEEL (P34370) Hypothetical protein C50C3.7 in chromosome III 34 0.29
SYIC_SCHPO (O13651) Isoleucyl-tRNA synthetase, cytoplasmic (EC 6... 32 1.4
SYM_METJA (Q58659) Methionyl-tRNA synthetase (EC 6.1.1.10) (Meth... 29 7.0
MET_HUMAN (P08581) Hepatocyte growth factor receptor precursor (... 29 7.0
I17S_HUMAN (Q9NRM6) Interleukin-17B receptor precursor (IL-17B r... 29 9.2
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 50.8 bits (120), Expect = 2e-06
Identities = 24/77 (31%), Positives = 45/77 (58%), Gaps = 1/77 (1%)
Query: 123 SMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKE-NGGVRVKD 181
++P++ +S ++ + K++ F W +KISWV W+ +C+ KE +GG+ +D
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 182 IRVMNISLLAKWRWRLI 198
+ N +LLAK +R+I
Sbjct: 62 LGWFNQALLAKQSFRII 78
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 48.1 bits (113), Expect = 1e-05
Identities = 39/201 (19%), Positives = 86/201 (42%), Gaps = 9/201 (4%)
Query: 6 YADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNC 65
+ADD + + + + + ++ F VSG KIN KS + + L
Sbjct: 699 FADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPF 758
Query: 66 SAGSITFKYLGLPVGANMRSM--STWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
+ S KYLG+ + +++ + ++PL+ I N W S+ GRI ++ +
Sbjct: 759 TIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILP 818
Query: 124 MPIFYLSF--LKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVKD 181
I+ + +K+ + + + + F+W +K + + ++ Q+ + GG+ + D
Sbjct: 819 KVIYRFNAIPIKLPMTFFTELEKTTLKFIW-----NQKRAHIAKSTLSQKNKAGGITLPD 873
Query: 182 IRVMNISLLAKWRWRLIDGRE 202
++ + + K W R+
Sbjct: 874 FKLYYKATVTKTAWYWYQNRD 894
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 41.2 bits (95), Expect = 0.002
Identities = 41/201 (20%), Positives = 86/201 (42%), Gaps = 9/201 (4%)
Query: 6 YADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNC 65
+ADD + + + + + +++ + VSG KIN KS + D +
Sbjct: 699 FADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKTVKDSIPF 758
Query: 66 SAGSITFKYLGLPVGANMRSM--STWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
+ KYLG+ + +++ + +E L + I +N W S+ GRI ++ +
Sbjct: 759 TVVPKKMKYLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSILP 818
Query: 124 MPIFYLSF--LKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVKD 181
I+ + +K + +K + +I F+W +K + + + + GG+ + D
Sbjct: 819 KAIYNFNAIPIKAPLSYFKDLEKIILHFIW-----NQKKPQIAKTLLSNKNKAGGITLPD 873
Query: 182 IRVMNISLLAKWRWRLIDGRE 202
+R+ S++ K W RE
Sbjct: 874 LRLYYKSIVIKTAWYWHKNRE 894
>YLJ7_CAEEL (P34370) Hypothetical protein C50C3.7 in chromosome III
Length = 398
Score = 33.9 bits (76), Expect = 0.29
Identities = 27/69 (39%), Positives = 36/69 (52%), Gaps = 5/69 (7%)
Query: 94 ETIGGRLNTWSTRYISF---GGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRI-VRIQR*F 149
ETIGG + TW+T S+ GR+VLL + + K L+G KRI R QR
Sbjct: 48 ETIGGAVLTWATTIASWMNTNGRMVLLAKTFQATNQVLIFGRKQLIGQIKRIDYRFQRNT 107
Query: 150 LWGGVGGGK 158
+ GG+ G K
Sbjct: 108 M-GGLTGHK 115
>SYIC_SCHPO (O13651) Isoleucyl-tRNA synthetase, cytoplasmic (EC
6.1.1.5) (Isoleucine--tRNA ligase) (IleRS)
Length = 1064
Score = 31.6 bits (70), Expect = 1.4
Identities = 27/105 (25%), Positives = 41/105 (38%), Gaps = 21/105 (20%)
Query: 16 ASVQNLWTMKAILRGFQM-------VSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAG 68
+++++ T A L+G+ + GL + +GI S+D MAM D N
Sbjct: 56 STIKDSVTRYACLKGYHVERRFGWDTHGLPVEHEIDKKLGITGSDDVMAMGIDKYNAECR 115
Query: 69 SITFKYLGLPVGANMRSMSTWEPLVETIGGRL---NTWSTRYISF 110
I Y S W VE +G + N + T Y SF
Sbjct: 116 KIVMTY-----------ASEWRATVERLGRWIDFDNDYKTLYPSF 149
>SYM_METJA (Q58659) Methionyl-tRNA synthetase (EC 6.1.1.10)
(Methionine--tRNA ligase) (MetRS)
Length = 651
Score = 29.3 bits (64), Expect = 7.0
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 4/46 (8%)
Query: 100 LNTWS-TRYISFGGRIVLLNSVLN---SMPIFYLSFLKMLVGVWKR 141
L+ W +R IS+G I N V+ PI Y+SF KML +WK+
Sbjct: 222 LHDWDISRDISWGVPIPGTNQVMYVWLEAPIGYISFTKMLGEIWKK 267
>MET_HUMAN (P08581) Hepatocyte growth factor receptor precursor (EC
2.7.1.112) (Met proto-oncogene tyrosine kinase) (c-met)
(HGF receptor) (HGF-SF receptor)
Length = 1390
Score = 29.3 bits (64), Expect = 7.0
Identities = 14/33 (42%), Positives = 21/33 (63%)
Query: 92 LVETIGGRLNTWSTRYISFGGRIVLLNSVLNSM 124
L+ G LN+ ++R+IS GG+ L SV NS+
Sbjct: 674 LLTLTGNYLNSGNSRHISIGGKTCTLKSVSNSI 706
>I17S_HUMAN (Q9NRM6) Interleukin-17B receptor precursor (IL-17B
receptor) (IL-17 receptor homolog 1) (IL-17Rh1)
(IL17Rh1) (Cytokine receptor CRL4) (UNQ2501/PRO19612)
Length = 502
Score = 28.9 bits (63), Expect = 9.2
Identities = 20/66 (30%), Positives = 28/66 (42%), Gaps = 9/66 (13%)
Query: 22 WTMKA-----ILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLG 76
W ++A +L+ ++ K NF S V N +E F S G TF Y+G
Sbjct: 70 WVLRADASIRLLKATKICVTGKSNFQSYSCVRCNYTEAFQTQT----RPSGGKWTFSYIG 125
Query: 77 LPVGAN 82
PV N
Sbjct: 126 FPVELN 131
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,203,948
Number of Sequences: 164201
Number of extensions: 919138
Number of successful extensions: 2377
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2372
Number of HSP's gapped (non-prelim): 10
length of query: 211
length of database: 59,974,054
effective HSP length: 105
effective length of query: 106
effective length of database: 42,732,949
effective search space: 4529692594
effective search space used: 4529692594
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148816.5


