

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148655.10 + phase: 0
(250 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
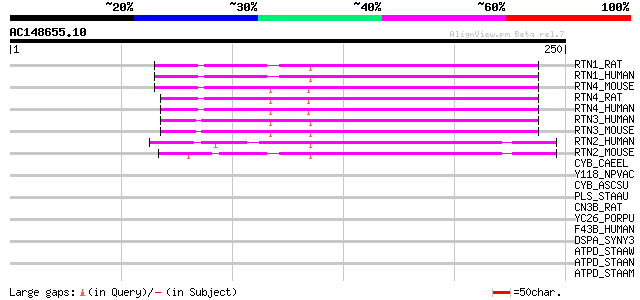
Score E
Sequences producing significant alignments: (bits) Value
RTN1_RAT (Q64548) Reticulon 1 (Neuroendocrine-specific protein) ... 79 8e-15
RTN1_HUMAN (Q16799) Reticulon 1 (Neuroendocrine-specific protein) 78 2e-14
RTN4_MOUSE (Q99P72) Reticulon 4 (Neurite outgrowth inhibitor) (N... 68 2e-11
RTN4_RAT (Q9JK11) Reticulon 4 (Neurite outgrowth inhibitor) (Nog... 66 7e-11
RTN4_HUMAN (Q9NQC3) Reticulon 4 (Neurite outgrowth inhibitor) (N... 65 2e-10
RTN3_HUMAN (O95197) Reticulon protein 3 (Neuroendocrine-specific... 60 5e-09
RTN3_MOUSE (Q9ES97) Reticulon protein 3 59 1e-08
RTN2_HUMAN (O75298) Reticulon protein 2 (Neuroendocrine-specific... 55 1e-07
RTN2_MOUSE (O70622) Reticulon protein 2 (Neuroendocrine-specific... 53 8e-07
CYB_CAEEL (P24890) Cytochrome b 32 1.1
Y118_NPVAC (P41671) Hypothetical 18.7 kDa protein in HE65-PK2 in... 30 4.2
CYB_ASCSU (P24878) Cytochrome b 30 4.2
PLS_STAAU (P80544) Surface protein precursor (Plasmin-sensitive ... 30 5.5
CN3B_RAT (Q63085) cGMP-inhibited 3',5'-cyclic phosphodiesterase ... 30 7.1
YC26_PORPU (P51392) Hypothetical sensor-like histidine kinase yc... 29 9.3
F43B_HUMAN (Q6ZT52) Protein FAM43B 29 9.3
DSPA_SYNY3 (P20169) Drug sensory protein A (EC 2.7.3.-) 29 9.3
ATPD_STAAW (P63659) ATP synthase delta chain (EC 3.6.3.14) 29 9.3
ATPD_STAAN (P99109) ATP synthase delta chain (EC 3.6.3.14) 29 9.3
ATPD_STAAM (P63658) ATP synthase delta chain (EC 3.6.3.14) 29 9.3
>RTN1_RAT (Q64548) Reticulon 1 (Neuroendocrine-specific protein)
(S-rex)
Length = 777
Score = 79.3 bits (194), Expect = 8e-15
Identities = 55/188 (29%), Positives = 91/188 (48%), Gaps = 22/188 (11%)
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFI 125
K D+L WR+ K T I G+ L + F L Q++++++V +L + AL+A S I
Sbjct: 589 KAIDLLYWRDIKQTGIVFGS--FLLLLFSLTQFSVVSVVAYLALAALSATI-----SFRI 641
Query: 126 HKSPLQIPH---------------VVIPQECVFEAASVLRIEINQVFAILREIGTGRDIK 170
+KS LQ + + QE + + L++ +N LR + +D+
Sbjct: 642 YKSVLQAVQKTDEGHPFKAYLELEITLSQEQIQKYTDCLQLYVNSTLKELRRLFLVQDLV 701
Query: 171 KFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQ 230
L +W ++ +G+ FN LTL + VS+FTLP+VY K++ QVD I
Sbjct: 702 DSLKFAVLMWLLTYVGALFNGLTLLLMAVVSMFTLPVVYVKHQAQVDQYLGLVRTHINTV 761
Query: 231 YAVLDAKV 238
A + AK+
Sbjct: 762 VAKIQAKI 769
>RTN1_HUMAN (Q16799) Reticulon 1 (Neuroendocrine-specific protein)
Length = 776
Score = 78.2 bits (191), Expect = 2e-14
Identities = 54/188 (28%), Positives = 90/188 (47%), Gaps = 22/188 (11%)
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFI 125
K D+L WR+ K T I G+ L + F L Q++++++V +L + AL+A S I
Sbjct: 588 KAIDLLYWRDIKQTGIVFGS--FLLLLFSLTQFSVVSVVAYLALAALSATI-----SFRI 640
Query: 126 HKSPLQIPH---------------VVIPQECVFEAASVLRIEINQVFAILREIGTGRDIK 170
+KS LQ + + QE + + L+ +N LR + +D+
Sbjct: 641 YKSVLQAVQKTDEGHPFKAYLELEITLSQEQIQKYTDCLQFYVNSTLKELRRLFLVQDLV 700
Query: 171 KFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQ 230
L +W ++ +G+ FN LTL + VS+FTLP+VY K++ Q+D I
Sbjct: 701 DSLKFAVLMWLLTYVGALFNGLTLLLMAVVSMFTLPVVYVKHQAQIDQYLGLVRTHINAV 760
Query: 231 YAVLDAKV 238
A + AK+
Sbjct: 761 VAKIQAKI 768
>RTN4_MOUSE (Q99P72) Reticulon 4 (Neurite outgrowth inhibitor) (Nogo
protein)
Length = 199
Score = 67.8 bits (164), Expect = 2e-11
Identities = 45/183 (24%), Positives = 87/183 (46%), Gaps = 12/183 (6%)
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALF---LWSNAS 122
K D+L WR+ K T + GA +L++ L ++++++ ++ + L+ ++
Sbjct: 11 KVVDLLYWRDIKKTGVVFGA--SLFLLLSLTVFSIVSVTAYIALALLSVTISFRIYKGVI 68
Query: 123 VFIHKSPLQIP-------HVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTV 175
I KS P V I +E V + ++ +N LR + D+ L
Sbjct: 69 QAIQKSDEGHPFRAYLESEVAISEELVQKYSNSALGHVNSTIKELRRLFLVDDLVDSLKF 128
Query: 176 IAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLD 235
+W + +G+ FN LTL + +SLF++P++YE+++ Q+D A +K A +
Sbjct: 129 AVLMWVFTYVGALFNGLTLLILALISLFSIPVIYERHQAQIDHYLGLANKSVKDAMAKIQ 188
Query: 236 AKV 238
AK+
Sbjct: 189 AKI 191
>RTN4_RAT (Q9JK11) Reticulon 4 (Neurite outgrowth inhibitor) (Nogo
protein) (Foocen) (Glut4 vesicle 20 kDa protein)
Length = 1163
Score = 66.2 bits (160), Expect = 7e-11
Identities = 44/180 (24%), Positives = 86/180 (47%), Gaps = 12/180 (6%)
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALF---LWSNASVFI 125
D+L WR+ K T + GA +L++ L ++++++ ++ + L+ ++ I
Sbjct: 978 DLLYWRDIKKTGVVFGA--SLFLLLSLTVFSIVSVTAYIALALLSVTISFRIYKGVIQAI 1035
Query: 126 HKSPLQIP-------HVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAG 178
KS P V I +E V + ++ +N LR + D+ L
Sbjct: 1036 QKSDEGHPFRAYLESEVAISEELVQKYSNSALGHVNSTIKELRRLFLVDDLVDSLKFAVL 1095
Query: 179 LWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKV 238
+W + +G+ FN LTL + +SLF++P++YE+++ Q+D A +K A + AK+
Sbjct: 1096 MWVFTYVGALFNGLTLLILALISLFSIPVIYERHQVQIDHYLGLANKSVKDAMAKIQAKI 1155
>RTN4_HUMAN (Q9NQC3) Reticulon 4 (Neurite outgrowth inhibitor) (Nogo
protein) (Foocen) (Neuroendocrine-specific protein) (NSP)
(Neuroendocrine specific protein C homolog) (RTN-x)
(Reticulon 5) (My043 protein)
Length = 1192
Score = 65.1 bits (157), Expect = 2e-10
Identities = 44/180 (24%), Positives = 86/180 (47%), Gaps = 12/180 (6%)
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALF---LWSNASVFI 125
D+L WR+ K T + GA +L++ L ++++++ ++ + L+ ++ I
Sbjct: 1007 DLLYWRDIKKTGVVFGA--SLFLLLSLTVFSIVSVTAYIALALLSVTISFRIYKGVIQAI 1064
Query: 126 HKSPLQIP-------HVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAG 178
KS P V I +E V + ++ +N LR + D+ L
Sbjct: 1065 QKSDEGHPFRAYLESEVAISEELVQKYSNSALGHVNCTIKELRRLFLVDDLVDSLKFAVL 1124
Query: 179 LWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKV 238
+W + +G+ FN LTL + +SLF++P++YE+++ Q+D A +K A + AK+
Sbjct: 1125 MWVFTYVGALFNGLTLLILALISLFSVPVIYERHQAQIDHYLGLANKNVKDAMAKIQAKI 1184
>RTN3_HUMAN (O95197) Reticulon protein 3 (Neuroendocrine-specific
protein-like 2) (NSP-like protein II) (NSPLII)
Length = 236
Score = 60.1 bits (144), Expect = 5e-09
Identities = 39/180 (21%), Positives = 83/180 (45%), Gaps = 12/180 (6%)
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALF---LWSNASVFI 125
D++ WR+ K T G T L + L +++I++V +L++ L+ ++ + +
Sbjct: 50 DLIFWRDVKKTGFVFG--TTLIMLLSLAAFSVISVVSYLILALLSVTISFRIYKSVIQAV 107
Query: 126 HKSPLQIPH-------VVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAG 178
KS P + + E + + IN+ ++ + D+ L +
Sbjct: 108 QKSEEGHPFKAYLDVDITLSSEAFHNYMNAAMVHINRALKLIIRLFLVEDLVDSLKLAVF 167
Query: 179 LWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKV 238
+W ++ +G+ FN +TL + + +F++P+VYEK + Q+D A + K + AK+
Sbjct: 168 MWLMTYVGAVFNGITLLILAELLIFSVPIVYEKYKTQIDHYVGIARDQTKSIVEKIQAKL 227
>RTN3_MOUSE (Q9ES97) Reticulon protein 3
Length = 237
Score = 58.9 bits (141), Expect = 1e-08
Identities = 38/180 (21%), Positives = 83/180 (46%), Gaps = 12/180 (6%)
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALF---LWSNASVFI 125
D++ WR+ K T G T L + L +++I++V +L++ L+ ++ + +
Sbjct: 51 DLIFWRDVKKTGFVFG--TTLIMLLSLAAFSVISVVSYLILALLSVTISFRVYKSVIQAV 108
Query: 126 HKSPLQIPH-------VVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAG 178
KS P + + E + + +N+ ++ + D+ L +
Sbjct: 109 QKSEEGHPFKAYLDVDITLSSEAFHNYMNAAMVHVNKALKLIIRLFLVEDLVDSLKLAVF 168
Query: 179 LWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKV 238
+W ++ +G+ FN +TL + + +F++P+VYEK + Q+D A + K + AK+
Sbjct: 169 MWLMTYVGAVFNGITLLILAELLVFSVPIVYEKYKTQIDHYVGIARDQTKSIVEKIQAKL 228
>RTN2_HUMAN (O75298) Reticulon protein 2 (Neuroendocrine-specific
protein-like 1) (NSP-like protein 1) (NSPLI)
Length = 545
Score = 55.5 bits (132), Expect = 1e-07
Identities = 44/199 (22%), Positives = 88/199 (44%), Gaps = 28/199 (14%)
Query: 64 GGKPADILLWRNKKCTAIALGAGTALWV-FFELMQYNLITLVCHLMILALAALFLWSNAS 122
G K AD+L W++ + + + T L V L+ ++++++ HL A L L S
Sbjct: 342 GSKVADLLYWKDTRTSGVVF---TGLMVSLLCLLHFSIVSVAAHL-----ALLLLCGTIS 393
Query: 123 VFIHKSPLQIPH---------------VVIPQECVFEAASVLRIEINQVFAILREIGTGR 167
+ +++ LQ H + + +E + + + LR
Sbjct: 394 LRVYRKVLQAVHRGDGANPFQAYLDVDLTLTREQTERLSHQITSRVVSAATQLRHFFLVE 453
Query: 168 DIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEI 227
D+ L + + ++ +G+ FN LTL + + LFT+PL+Y +++ Q+D +
Sbjct: 454 DLVDSLKLALLFYILTFVGAIFNGLTLLILGVIGLFTIPLLYRQHQAQIDQYVGL----V 509
Query: 228 KKQYAVLDAKVLSQIPIAG 246
Q + + AK+ ++IP G
Sbjct: 510 TNQLSHIKAKIRAKIPGTG 528
>RTN2_MOUSE (O70622) Reticulon protein 2 (Neuroendocrine-specific
protein-like 1) (NSP-like protein 1) (NSPLI)
Length = 471
Score = 52.8 bits (125), Expect = 8e-07
Identities = 43/195 (22%), Positives = 89/195 (45%), Gaps = 28/195 (14%)
Query: 68 ADILLWRNKKCT-AIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIH 126
AD+L W++ + + A+ G +L L+ ++++++ HL +L L A S+ ++
Sbjct: 273 ADLLYWKDTRTSGAVFTGLMASLLC---LLHFSIVSVAAHLALLGLCATI-----SLRVY 324
Query: 127 KSPLQIPH---------------VVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKK 171
+ LQ H + + +E + + + LR D+
Sbjct: 325 RKVLQAVHRGDGTNPFQAYLDMDLTLTREQTERLSQQIASHVVSTATQLRHFFLVEDLVD 384
Query: 172 FLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQY 231
L + + ++ +G+ FN LTL + V+LFT+PL+Y +++ Q+D + Q
Sbjct: 385 SLKLALLFYILTFVGAIFNGLTLVILGVVALFTVPLLYRQHQAQIDQYVGL----VTNQL 440
Query: 232 AVLDAKVLSQIPIAG 246
+ + AK+ ++IP G
Sbjct: 441 SHIKAKIRAKIPGTG 455
>CYB_CAEEL (P24890) Cytochrome b
Length = 370
Score = 32.3 bits (72), Expect = 1.1
Identities = 24/108 (22%), Positives = 49/108 (45%), Gaps = 11/108 (10%)
Query: 98 YNLITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVF 157
YN++ + + +L+L F +A +FI P+ P ++P+ A ++LR N+V
Sbjct: 227 YNIVIWLLFI-VLSLIYPFNLGDAEMFIEADPMMSPVHIVPEWYFLFAYAILRAIPNKVL 285
Query: 158 AILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTL 205
++ L I +F +++ + + LT F V +F +
Sbjct: 286 GVI----------ALLMSIVTFYFFALVNNYTSCLTKLNKFLVFMFII 323
>Y118_NPVAC (P41671) Hypothetical 18.7 kDa protein in HE65-PK2
intergenic region
Length = 157
Score = 30.4 bits (67), Expect = 4.2
Identities = 13/35 (37%), Positives = 21/35 (59%)
Query: 188 CFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEK 222
C NF T++ + YV LF + ++EK D +A+K
Sbjct: 41 CKNFNTMYNVMYVVLFAIFQLFEKGFDIAKTIADK 75
>CYB_ASCSU (P24878) Cytochrome b
Length = 365
Score = 30.4 bits (67), Expect = 4.2
Identities = 21/91 (23%), Positives = 41/91 (44%), Gaps = 3/91 (3%)
Query: 108 MILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGR 167
+ +L FL + +FI P+ P ++P+ A ++LR N+V ++ +
Sbjct: 231 IFFSLGYPFLLGDPEMFIESDPMMSPVHIVPEWYFLFAYAILRAIPNKVLGVVSLFASIL 290
Query: 168 DIKKFLTVIAGLWFISVIGSCFNFLTLFYIF 198
+ F+ V ++SV+ FL +IF
Sbjct: 291 VLVVFVLVNN---YVSVMSKLNKFLVFVFIF 318
>PLS_STAAU (P80544) Surface protein precursor (Plasmin-sensitive
surface protein) (230 kDa cell-wall protein)
Length = 1637
Score = 30.0 bits (66), Expect = 5.5
Identities = 15/35 (42%), Positives = 19/35 (53%)
Query: 5 DTESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPP 39
D++S D D SDSDS SD++ DHN P
Sbjct: 1559 DSDSDSDSDSDADSDSDSDSDSDADRDHNDKTDKP 1593
>CN3B_RAT (Q63085) cGMP-inhibited 3',5'-cyclic phosphodiesterase B (EC
3.1.4.17) (Cyclic GMP inhibited phosphodiesterase B)
(CGI-PDE B) (CGIPDE1)
Length = 1108
Score = 29.6 bits (65), Expect = 7.1
Identities = 15/37 (40%), Positives = 19/37 (50%)
Query: 5 DTESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIP 41
DTES DD +D D D DS+D+ +N P P
Sbjct: 1003 DTESDDDDDDDDDDDDDDDEELDSDDEETEDNLNPKP 1039
>YC26_PORPU (P51392) Hypothetical sensor-like histidine kinase ycf26
(EC 2.7.3.-)
Length = 656
Score = 29.3 bits (64), Expect = 9.3
Identities = 24/105 (22%), Positives = 42/105 (39%), Gaps = 14/105 (13%)
Query: 92 FFELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQ--IPHVVIPQECVFEAASVL 149
FF L +YN H +SN + + LQ + ++IP + +L
Sbjct: 138 FFSLSEYNCFRNENHY----------FSNTPIVNTNNRLQGEVIDIIIPLSKEKKLLGIL 187
Query: 150 RIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTL 194
I IN + RD+ + V +W + ++G+ FN T+
Sbjct: 188 NIGINSNPTLTTSSQLTRDVS--VAVFISIWLMVILGAAFNAFTI 230
>F43B_HUMAN (Q6ZT52) Protein FAM43B
Length = 329
Score = 29.3 bits (64), Expect = 9.3
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 1/52 (1%)
Query: 63 GGGKPADI-LLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALA 113
GG +PA LL R CTA WV+ ++ + L CH ++LA A
Sbjct: 133 GGRRPAHAYLLPRITYCTADGRHPRVFAWVYRHQARHKAVVLRCHAVLLARA 184
>DSPA_SYNY3 (P20169) Drug sensory protein A (EC 2.7.3.-)
Length = 663
Score = 29.3 bits (64), Expect = 9.3
Identities = 18/63 (28%), Positives = 30/63 (47%), Gaps = 2/63 (3%)
Query: 132 IPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNF 191
+ V IP + + VL I IN A + RD+ + V +W + ++G+ FN
Sbjct: 162 VTDVFIPLQYQGKFLGVLAIGINPNPAAVNSSNLTRDVT--IAVFISIWVMVILGAVFNA 219
Query: 192 LTL 194
LT+
Sbjct: 220 LTI 222
>ATPD_STAAW (P63659) ATP synthase delta chain (EC 3.6.3.14)
Length = 179
Score = 29.3 bits (64), Expect = 9.3
Identities = 22/82 (26%), Positives = 41/82 (49%), Gaps = 3/82 (3%)
Query: 169 IKKFLTVIAGLWFISVIGSCFN-FLTLFYIFYVSLF-TLPLVYEKNEDQVDALAEKAMIE 226
IK + V+A IS+I F F +L+ Y F T+ YE +++++D + + +
Sbjct: 72 IKNMMYVLADNRHISLIADVFKAFQSLYNGHYNQDFATIESTYELSQEELDKIVKLVTQQ 131
Query: 227 IKKQYAVLDAKVLSQIPIAGFK 248
K ++D K+ + I GF+
Sbjct: 132 TKLSKVIVDTKINPDL-IGGFR 152
>ATPD_STAAN (P99109) ATP synthase delta chain (EC 3.6.3.14)
Length = 179
Score = 29.3 bits (64), Expect = 9.3
Identities = 22/82 (26%), Positives = 41/82 (49%), Gaps = 3/82 (3%)
Query: 169 IKKFLTVIAGLWFISVIGSCFN-FLTLFYIFYVSLF-TLPLVYEKNEDQVDALAEKAMIE 226
IK + V+A IS+I F F +L+ Y F T+ YE +++++D + + +
Sbjct: 72 IKNMMYVLADNRHISLIADVFKAFQSLYNGHYNQDFATIESTYELSQEELDKIVKLVTQQ 131
Query: 227 IKKQYAVLDAKVLSQIPIAGFK 248
K ++D K+ + I GF+
Sbjct: 132 TKLSKVIVDTKINPDL-IGGFR 152
>ATPD_STAAM (P63658) ATP synthase delta chain (EC 3.6.3.14)
Length = 179
Score = 29.3 bits (64), Expect = 9.3
Identities = 22/82 (26%), Positives = 41/82 (49%), Gaps = 3/82 (3%)
Query: 169 IKKFLTVIAGLWFISVIGSCFN-FLTLFYIFYVSLF-TLPLVYEKNEDQVDALAEKAMIE 226
IK + V+A IS+I F F +L+ Y F T+ YE +++++D + + +
Sbjct: 72 IKNMMYVLADNRHISLIADVFKAFQSLYNGHYNQDFATIESTYELSQEELDKIVKLVTQQ 131
Query: 227 IKKQYAVLDAKVLSQIPIAGFK 248
K ++D K+ + I GF+
Sbjct: 132 TKLSKVIVDTKINPDL-IGGFR 152
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,772,730
Number of Sequences: 164201
Number of extensions: 1197034
Number of successful extensions: 4906
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 4754
Number of HSP's gapped (non-prelim): 111
length of query: 250
length of database: 59,974,054
effective HSP length: 108
effective length of query: 142
effective length of database: 42,240,346
effective search space: 5998129132
effective search space used: 5998129132
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC148655.10


