

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
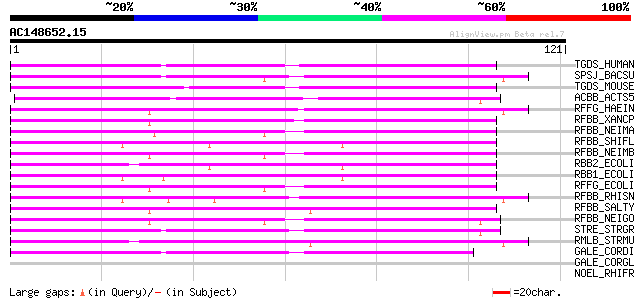
Score E
Sequences producing significant alignments: (bits) Value
TGDS_HUMAN (O95455) dTDP-D-glucose 4,6-dehydratase (EC 4.2.1.46) 72 2e-13
SPSJ_BACSU (P39630) Spore coat polysaccharide biosynthesis prote... 72 2e-13
TGDS_MOUSE (Q8VDR7) dTDP-D-glucose 4,6-dehydratase (EC 4.2.1.46) 70 8e-13
ACBB_ACTS5 (Q9ZAE8) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 64 5e-11
RFFG_HAEIN (P44914) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 64 6e-11
RFBB_XANCP (P55295) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 63 1e-10
RFBB_NEIMA (Q9S642) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 60 7e-10
RFBB_SHIFL (P37777) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 59 1e-09
RFBB_NEIMB (P55294) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 59 1e-09
RBB2_ECOLI (P55293) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 59 2e-09
RBB1_ECOLI (P37759) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 57 6e-09
RFFG_ECOLI (P27830) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 56 1e-08
RFBB_RHISN (P55462) Probable dTDP-glucose 4,6-dehydratase (EC 4.... 56 1e-08
RFBB_SALTY (P26391) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 56 2e-08
RFBB_NEIGO (P37761) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 54 5e-08
STRE_STRGR (P29782) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 53 1e-07
RMLB_STRMU (P95780) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46) 51 5e-07
GALE_CORDI (P33119) UDP-glucose 4-epimerase (EC 5.1.3.2) (Galact... 47 6e-06
GALE_CORGL (Q45291) UDP-glucose 4-epimerase (EC 5.1.3.2) (Galact... 39 0.002
NOEL_RHIFR (O85713) GDP-mannose 4,6-dehydratase (EC 4.2.1.47) (G... 38 0.005
>TGDS_HUMAN (O95455) dTDP-D-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 350
Score = 72.4 bits (176), Expect = 2e-13
Identities = 43/106 (40%), Positives = 58/106 (54%), Gaps = 4/106 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAAQTHVD S+ +FEFT + S + +V+ FI+VSTD+ YG + +
Sbjct: 96 LHFAAQTHVDLSFVRAFEFTYVNVYGTHVLVSA-AHEARVEKFIYVSTDEVYGGSLDKEF 154
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
E+S TNPY++ K AE V SY Y PV+ TR VY
Sbjct: 155 ---DESSPKQPTNPYASSKAAAECFVQSYWEQYKFPVVITRSSNVY 197
>SPSJ_BACSU (P39630) Spore coat polysaccharide biosynthesis protein
spsJ
Length = 315
Score = 72.4 bits (176), Expect = 2e-13
Identities = 44/116 (37%), Positives = 63/116 (53%), Gaps = 7/116 (6%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGE--TDEN 58
+HFAA++HVD S + F T+ M V K + K IH+STD+ YG+ D+
Sbjct: 79 IHFAAESHVDRSISQAEPFITTNVMGTYRLAEA-VLKGKAKKLIHISTDEVYGDLKADDP 137
Query: 59 AVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY-PFSHERK 113
A E + L NPYSA K +++LV+SY +++ LP I TR Y P+ H K
Sbjct: 138 AFT---ETTPLSPNNPYSASKASSDLLVLSYVKTHKLPAIITRCSNNYGPYQHSEK 190
>TGDS_MOUSE (Q8VDR7) dTDP-D-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 355
Score = 70.1 bits (170), Expect = 8e-13
Identities = 42/106 (39%), Positives = 56/106 (52%), Gaps = 4/106 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAAQTHVD S+ +FEFT + + V+ FI+VSTD+ YG + +
Sbjct: 96 LHFAAQTHVDLSFVRAFEFTYVNVYGTHVLVNAAYEAG-VEKFIYVSTDEVYGGSLDQEF 154
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
E+S TNPY++ K AE V SY Y PV+ TR VY
Sbjct: 155 ---DESSPKQPTNPYASSKAAAECFVQSYWERYKFPVVITRSSNVY 197
>ACBB_ACTS5 (Q9ZAE8) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 341
Score = 64.3 bits (155), Expect = 5e-11
Identities = 36/111 (32%), Positives = 60/111 (53%), Gaps = 9/111 (8%)
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVV 61
HFAA+THVD S S F ++ + + ++ + F+HVSTD+ YG D +
Sbjct: 83 HFAAETHVDRSVVASGPFVASNLVGTQVLLDAAL-RHHIGRFLHVSTDEVYGSIDTGSWA 141
Query: 62 GNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR-----GDRVYP 107
H L +PY+A K G+++L ++Y +++G+ V+ TR G R +P
Sbjct: 142 EGH---PLAPNSPYAASKAGSDLLALAYHQTHGMDVVVTRCSNNYGPRQFP 189
>RFFG_HAEIN (P44914) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 338
Score = 63.9 bits (154), Expect = 6e-11
Identities = 40/122 (32%), Positives = 61/122 (49%), Gaps = 10/122 (8%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S + +F QT+ + + + +L +K F H+STD+ Y
Sbjct: 79 MHLAAESHVDRSISGAADFVQTNIVGTYTLLEVAKNYWHTLDEAKKTTFRFHHISTDEVY 138
Query: 53 GETDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY-PFSHE 111
G+ + E S ++PYSA K + LV ++ R+YGLPVI T Y + H
Sbjct: 139 GDLSLSEPAFT-EQSPYHPSSPYSASKAASNHLVQAWHRTYGLPVIITNSSNNYGAYQHA 197
Query: 112 RK 113
K
Sbjct: 198 EK 199
>RFBB_XANCP (P55295) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 351
Score = 63.2 bits (152), Expect = 1e-10
Identities = 37/114 (32%), Positives = 60/114 (52%), Gaps = 10/114 (8%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
++FAA++HVD S E F QT+ + ++ +++L ++ F+HVSTD+ Y
Sbjct: 78 LNFAAESHVDRSIEGPGAFIQTNVVGTLALLEAVRDYWKALPDTRRDAFRFLHVSTDEVY 137
Query: 53 GETDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
G E E + +PYSA K ++ LV ++ +YGLPV+TT Y
Sbjct: 138 GTLGETGKFT--ETTPYAPNSPYSASKAASDHLVRAFHHTYGLPVLTTNCSNNY 189
>RFBB_NEIMA (Q9S642) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 341
Score = 60.5 bits (145), Expect = 7e-10
Identities = 38/117 (32%), Positives = 61/117 (51%), Gaps = 15/117 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSF--------RSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S ++ EF QT+ + + + + K++ F H+STD+ Y
Sbjct: 79 MHLAAESHVDRSIGSAGEFIQTNIVGTFNLLEAARAYRQQMPSEKHEAFRFHHISTDEVY 138
Query: 53 GE---TDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
G+ TD+ E + ++PYSA K ++ LV ++ R+YGLP I T Y
Sbjct: 139 GDLSGTDDLFT----ETAPYAPSSPYSASKASSDHLVRAWLRTYGLPTIVTNCSNNY 191
>RFBB_SHIFL (P37777) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 361
Score = 59.3 bits (142), Expect = 1e-09
Identities = 40/121 (33%), Positives = 60/121 (49%), Gaps = 15/121 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTS----FMELMSFRSLQVSKNQVKG----FIHVSTDKFY 52
MH AA++HVD S F +T+ ++ L + R+ + N K F H+STD+ Y
Sbjct: 78 MHLAAESHVDRSITGPAAFIETNIVGTYVLLEAARNYWSALNDEKKKSFRFHHISTDEVY 137
Query: 53 GETDENAVVGNHEASQLLE-------TNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRV 105
G+ N+EA L ++PYSA K ++ LV ++ R+YGLP I T
Sbjct: 138 GDLPHPDEANNNEALPLFTETTAYAPSSPYSASKASSDHLVRAWKRTYGLPTIVTNCSNN 197
Query: 106 Y 106
Y
Sbjct: 198 Y 198
>RFBB_NEIMB (P55294) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 355
Score = 59.3 bits (142), Expect = 1e-09
Identities = 37/117 (31%), Positives = 62/117 (52%), Gaps = 15/117 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S ++ EF QT+ + + ++ + +++ F H+STD+ Y
Sbjct: 79 MHLAAESHVDRSIGSAGEFIQTNIVGTFNLLEAARAYWQQMPSEQHEAFRFHHISTDEVY 138
Query: 53 GE---TDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
G+ TD+ E + ++PYSA K ++ LV ++ R+YGLP I T Y
Sbjct: 139 GDLGGTDDLFT----ETAPYAPSSPYSASKASSDHLVRAWLRTYGLPTIVTNCSNNY 191
>RBB2_ECOLI (P55293) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 361
Score = 58.9 bits (141), Expect = 2e-09
Identities = 38/123 (30%), Positives = 61/123 (48%), Gaps = 19/123 (15%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKG----------FIHVSTDK 50
MH AA++HVD S F +T+ + ++ L+ ++N G F H+STD+
Sbjct: 78 MHLAAESHVDRSITGPAAFIETNIVG--TYVLLEAARNYWSGLDDEKKKNFRFHHISTDE 135
Query: 51 FYGETDENAVVGNHEASQLLE-------TNPYSAMKIGAEMLVMSYGRSYGLPVITTRGD 103
YG+ V ++E QL ++PYSA K ++ LV ++ R+YGLP I +
Sbjct: 136 VYGDLPHPDEVNSNETLQLFTETTAYAPSSPYSASKASSDHLVRAWKRTYGLPTIVSNCS 195
Query: 104 RVY 106
Y
Sbjct: 196 NNY 198
>RBB1_ECOLI (P37759) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 361
Score = 57.4 bits (137), Expect = 6e-09
Identities = 40/121 (33%), Positives = 58/121 (47%), Gaps = 15/121 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTS----FMELMSFRS----LQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S F +T+ ++ L + R+ L K F H+STD+ Y
Sbjct: 78 MHLAAESHVDRSITGPAAFIETNIVGTYVLLEAARNYWSALDSDKKNSFRFHHISTDEVY 137
Query: 53 GETDENAVVGNHEASQLLE-------TNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRV 105
G+ V N E L ++PYSA K ++ LV ++ R+YGLP I T
Sbjct: 138 GDLPHPDEVNNTEELPLFTETTAYAPSSPYSASKASSDHLVRAWKRTYGLPTIVTNCSNN 197
Query: 106 Y 106
Y
Sbjct: 198 Y 198
>RFFG_ECOLI (P27830) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 355
Score = 56.2 bits (134), Expect = 1e-08
Identities = 36/117 (30%), Positives = 58/117 (48%), Gaps = 15/117 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S + F +T+ + + + +L K F H+STD+ Y
Sbjct: 79 MHLAAESHVDRSIDGPAAFIETNIVGTYTLLEAARAYWNALTEDKKSAFRFHHISTDEVY 138
Query: 53 GE---TDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
G+ TD+ E + ++PYSA K ++ LV ++ R+YGLP + T Y
Sbjct: 139 GDLHSTDDFFT----ETTPYAPSSPYSASKASSDHLVRAWLRTYGLPTLITNCSNNY 191
>RFBB_RHISN (P55462) Probable dTDP-glucose 4,6-dehydratase (EC
4.2.1.46)
Length = 350
Score = 56.2 bits (134), Expect = 1e-08
Identities = 40/122 (32%), Positives = 62/122 (50%), Gaps = 11/122 (9%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTS----FMELMSFRSL--QVSKNQVKGF--IHVSTDKFY 52
+H AA++HVD S + +F QT+ F L + R +S+N+ F +HVSTD+ Y
Sbjct: 77 IHLAAESHVDRSITGADDFVQTNVNGTFTMLETARQYWSNLSQNRKAFFKMLHVSTDEVY 136
Query: 53 GETDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY-PFSHE 111
G + E S ++PYSA K ++ ++ R+YGLPV+ + Y PF
Sbjct: 137 GSLGDRGQF--EEVSPYDPSSPYSASKAASDHFATAWQRTYGLPVVISNCSNNYGPFHFP 194
Query: 112 RK 113
K
Sbjct: 195 EK 196
>RFBB_SALTY (P26391) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 361
Score = 55.8 bits (133), Expect = 2e-08
Identities = 37/121 (30%), Positives = 56/121 (45%), Gaps = 15/121 (12%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD S F +T+ + + + +L K F H+STD+ Y
Sbjct: 78 MHLAAESHVDRSITGPAAFIETNIVGTYALLEVARKYWSALGEDKKNNFRFHHISTDEVY 137
Query: 53 GETDENAVVGNH-------EASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRV 105
G+ V N E + ++PYSA K ++ LV ++ R+YGLP I T
Sbjct: 138 GDLPHPDEVENSVTLPLFTETTAYAPSSPYSASKASSDHLVRAWRRTYGLPTIVTNCSNN 197
Query: 106 Y 106
Y
Sbjct: 198 Y 198
>RFBB_NEIGO (P37761) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 346
Score = 54.3 bits (129), Expect = 5e-08
Identities = 37/123 (30%), Positives = 62/123 (50%), Gaps = 20/123 (16%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMS--------FRSLQVSKNQVKGFIHVSTDKFY 52
MH AA++HVD + ++ EF +T+ + ++ + K + F H+STD+ Y
Sbjct: 84 MHLAAESHVDRAIGSAGEFIRTNIVGTFDLLEAARAYWQQMPSEKREAFRFHHISTDEVY 143
Query: 53 GE---TDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR-----GDR 104
G+ TD+ E + ++PYSA K A+ LV ++ R+Y LP I + G R
Sbjct: 144 GDLHGTDDLFT----ETTPYAPSSPYSASKAAADHLVRAWQRTYRLPSIVSNCSNNYGPR 199
Query: 105 VYP 107
+P
Sbjct: 200 QFP 202
>STRE_STRGR (P29782) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 328
Score = 52.8 bits (125), Expect = 1e-07
Identities = 34/112 (30%), Positives = 57/112 (50%), Gaps = 9/112 (8%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+H AA++HVD S ++ F +T+ + +++ V F+ VSTD+ YG + +
Sbjct: 80 VHLAAESHVDRSLLDASVFVRTNVHGTQTLLDA-ATRHGVASFVQVSTDEVYGSLEHGSW 138
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR-----GDRVYP 107
E L +PYSA K ++L +++ S+GL V TR G R +P
Sbjct: 139 T---EDEPLRPNSPYSASKASGDLLALAHHVSHGLDVRVTRCSNNYGPRQFP 187
>RMLB_STRMU (P95780) dTDP-glucose 4,6-dehydratase (EC 4.2.1.46)
Length = 348
Score = 50.8 bits (120), Expect = 5e-07
Identities = 32/123 (26%), Positives = 63/123 (51%), Gaps = 12/123 (9%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+H+AA++H DNS ++ F T+F+ ++ L+ ++ F HVSTD+ YG+
Sbjct: 80 VHYAAESHNDNSLKDPSPFIYTNFVG--TYILLEAARKYDIRFHHVSTDEVYGDLPLRED 137
Query: 61 VGNH---------EASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY-PFSH 110
+ H ++ ++PYS+ K ++++V ++ RS+G+ + Y P+ H
Sbjct: 138 LPGHGEGPGEKFTAETKYNPSSPYSSTKAASDLIVKAWVRSFGVKATISNCSNNYGPYQH 197
Query: 111 ERK 113
K
Sbjct: 198 IEK 200
>GALE_CORDI (P33119) UDP-glucose 4-epimerase (EC 5.1.3.2)
(Galactowaldenase) (UDP-galactose 4-epimerase)
Length = 328
Score = 47.4 bits (111), Expect = 6e-06
Identities = 28/101 (27%), Positives = 48/101 (46%), Gaps = 4/101 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAA++ V S E E+ Q + + ++ + +N V+ + ST YGE + +
Sbjct: 69 LHFAARSLVGESVEKPDEYWQHNMVTTLALLDA-MKRNNVRNIVFSSTAATYGEPETVPI 127
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR 101
E + TNPY A K+ + + SY +YG + R
Sbjct: 128 T---EDAPTHPTNPYGATKLSIDYAITSYAHAYGFAATSLR 165
>GALE_CORGL (Q45291) UDP-glucose 4-epimerase (EC 5.1.3.2)
(Galactowaldenase) (UDP-galactose 4-epimerase)
Length = 329
Score = 38.9 bits (89), Expect = 0.002
Identities = 28/101 (27%), Positives = 45/101 (43%), Gaps = 4/101 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAA++ V S E E+ + + ++ + + V + ST YGE D V
Sbjct: 69 VHFAARSLVGESVEKPNEYWHDNVVTALTLLDA-MRAHGVNNLVFSSTAATYGEPD---V 124
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR 101
V E TN Y A K+ + + SY ++GL + R
Sbjct: 125 VPITEDMPTQPTNAYGATKLSIDYAITSYAAAFGLAATSLR 165
>NOEL_RHIFR (O85713) GDP-mannose 4,6-dehydratase (EC 4.2.1.47)
(GDP-D-mannose dehydratase)
Length = 351
Score = 37.7 bits (86), Expect = 0.005
Identities = 29/100 (29%), Positives = 51/100 (51%), Gaps = 13/100 (13%)
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIH------VSTDKFYGET 55
+ AAQ+HV S+E E+T + + + R L+ + + G IH ST + YG
Sbjct: 87 NLAAQSHVQVSFETP-EYTANADA-IGTLRMLEAIR--ILGLIHRTRFYQASTSELYGLA 142
Query: 56 DENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
E + +E + +PY+A K+ A +V++Y +YG+
Sbjct: 143 QE---IPQNEKTPFYPRSPYAAAKLYAYWIVVNYREAYGM 179
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,873,549
Number of Sequences: 164201
Number of extensions: 419345
Number of successful extensions: 1213
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 1151
Number of HSP's gapped (non-prelim): 60
length of query: 121
length of database: 59,974,054
effective HSP length: 97
effective length of query: 24
effective length of database: 44,046,557
effective search space: 1057117368
effective search space used: 1057117368
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148652.15


