

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.13 - phase: 0 /pseudo
(395 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
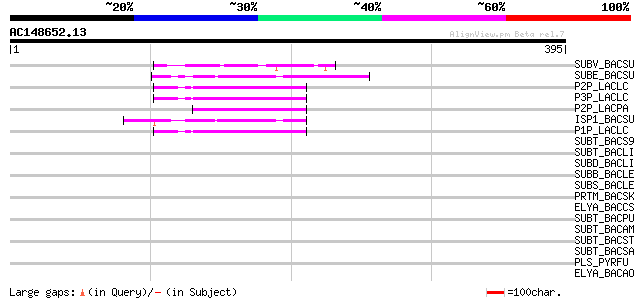
Score E
Sequences producing significant alignments: (bits) Value
SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (... 49 2e-05
SUBE_BACSU (P16396) Minor extracellular protease epr precursor (... 47 1e-04
P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96) ... 45 3e-04
P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)... 45 4e-04
P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96) ... 45 4e-04
ISP1_BACSU (P11018) Major intracellular serine protease precurso... 45 4e-04
P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-) (W... 44 7e-04
SUBT_BACS9 (P28842) Subtilisin precursor (EC 3.4.21.62) 43 0.001
SUBT_BACLI (P00780) Subtilisin Carlsberg precursor (EC 3.4.21.62) 42 0.002
SUBD_BACLI (P00781) Subtilisin DY (EC 3.4.21.62) 42 0.002
SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline prote... 41 0.006
SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline... 40 0.010
PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-) 40 0.010
ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-) 40 0.010
SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline mesenter... 40 0.014
SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62) (Su... 39 0.018
SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62) 39 0.023
SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC ... 39 0.023
PLS_PYRFU (P72186) Pyrolysin precursor (EC 3.4.21.-) 39 0.023
ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-) 39 0.023
>SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (EC
3.4.21.-)
Length = 806
Score = 48.9 bits (115), Expect = 2e-05
Identities = 40/137 (29%), Positives = 65/137 (47%), Gaps = 28/137 (20%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFD 162
H TH A T A GT +G P + + AY++ G +G + + ++A +
Sbjct: 233 HGTHVAGTVA------------ANGTIKGVAPDATLLAYRVLGPGGSG--TTENVIAGVE 278
Query: 163 DAISDGVDVITVSLGPEHASDFLNDP---IAIGSFHAMEKGILTTQAAGNFGPIPSSVCS 219
A+ DG DV+ +SLG + LN+P + AM +G++ + GN GP +V
Sbjct: 279 RAVQDGADVMNLSLG-----NSLNNPDWATSTALDWAMSEGVVAVTSNGNSGPNGWTV-- 331
Query: 220 GAPW----LVSVAATSI 232
G+P +SV AT +
Sbjct: 332 GSPGTSREAISVGATQL 348
>SUBE_BACSU (P16396) Minor extracellular protease epr precursor (EC
3.4.21.-)
Length = 645
Score = 46.6 bits (109), Expect = 1e-04
Identities = 44/156 (28%), Positives = 69/156 (44%), Gaps = 16/156 (10%)
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAF 161
GH TH A G + + YG G P ++I A K NG ++L
Sbjct: 171 GHGTHVAGIIGAKH----NGYGID-----GIAPEAQIYAVKALDQ--NGSGDLQSLLQGI 219
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFG-PIPSSVCSG 220
D +I++ +D++ +SLG S L+D + A E+G+L A+GN G P + +
Sbjct: 220 DWSIANRMDIVNMSLGTTSDSKILHDAVN----KAYEQGVLLVAASGNDGNGKPVNYPAA 275
Query: 221 APWLVSVAATSIDRQFIDKVILGNGKTFVGKSINIT 256
+V+V+AT+ Q G+ F NIT
Sbjct: 276 YSSVVAVSATNEKNQLASFSTTGDEVEFSAPGTNIT 311
>P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 45.1 bits (105), Expect = 3e-04
Identities = 33/110 (30%), Positives = 48/110 (43%), Gaps = 6/110 (5%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAIL-AAF 161
H H A G D AK G P +++ A K+ + D +G A L +A
Sbjct: 281 HGMHVAGIIGANGTGDDP----AKSVV-GVAPEAQLLAMKVFTNSDTSATTGSATLVSAI 335
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFG 211
+D+ G DV+ +SLG + + L DP +A E G +AGN G
Sbjct: 336 EDSAKIGADVLNMSLGSDSGNQTLEDPELAAVQNANESGTAAVISAGNSG 385
>P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
Length = 1902
Score = 44.7 bits (104), Expect = 4e-04
Identities = 32/110 (29%), Positives = 48/110 (43%), Gaps = 6/110 (5%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDA-ILAAF 161
H H A G D AK G P +++ A K+ + D +G A +++A
Sbjct: 281 HGMHVAGIIGANGTGDDP----AKSVV-GVAPEAQLLAMKVFSNSDTSAKTGSATVVSAI 335
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFG 211
+D+ G DV+ +SLG + L DP +A E G +AGN G
Sbjct: 336 EDSAKIGADVLNMSLGSNSGNQTLEDPELAAVQNANESGTAAVISAGNSG 385
>P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 44.7 bits (104), Expect = 4e-04
Identities = 26/82 (31%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 131 GGVPSSRIAAYKICGDIDNGRCSGDAIL-AAFDDAISDGVDVITVSLGPEHASDFLNDPI 189
G P +++ A K+ + D +G A L +A +D+ G DV+ +SLG + + L DP
Sbjct: 304 GVAPEAQLLAMKVFTNSDTSATTGSATLVSAIEDSAKIGADVLNMSLGSDSGNQTLEDPE 363
Query: 190 AIGSFHAMEKGILTTQAAGNFG 211
+A E G +AGN G
Sbjct: 364 IAAVQNANESGTAAVISAGNSG 385
>ISP1_BACSU (P11018) Major intracellular serine protease precursor
(EC 3.4.21.-) (ISP-1)
Length = 319
Score = 44.7 bits (104), Expect = 4e-04
Identities = 36/134 (26%), Positives = 59/134 (43%), Gaps = 18/134 (13%)
Query: 82 NKIIGARFYGDGDVSARYSF----GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSR 137
N+IIG + + D D + GH TH A T + + G G P +
Sbjct: 62 NQIIGGKNFTDDDGGKEDAISDYNGHGTHVAGTIAAND---------SNGGIAGVAPEAS 112
Query: 138 IAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAM 197
+ K+ G +NG + I+ + A+ VD+I++SLG L + + +A+
Sbjct: 113 LLIVKVLGG-ENGSGQYEWIINGINYAVEQKVDIISMSLGGPSDVPELKEAVK----NAV 167
Query: 198 EKGILTTQAAGNFG 211
+ G+L AAGN G
Sbjct: 168 KNGVLVVCAAGNEG 181
>P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-)
(Wall-associated serine proteinase)
Length = 1902
Score = 43.9 bits (102), Expect = 7e-04
Identities = 32/110 (29%), Positives = 48/110 (43%), Gaps = 6/110 (5%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAIL-AAF 161
H H A G D AK G P +++ A K+ + D +G + L +A
Sbjct: 281 HGMHVAGIIGANGTGDDP----AKSVV-GVAPEAQLLAMKVFTNSDTSATTGSSTLVSAI 335
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFG 211
+D+ G DV+ +SLG + + L DP +A E G +AGN G
Sbjct: 336 EDSAKIGADVLNMSLGSDSGNQTLEDPELAAVQNANESGTAAVISAGNSG 385
>SUBT_BACS9 (P28842) Subtilisin precursor (EC 3.4.21.62)
Length = 420
Score = 43.1 bits (100), Expect = 0.001
Identities = 39/137 (28%), Positives = 59/137 (42%), Gaps = 15/137 (10%)
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCS--GDAILA 159
GH TH A +A YG A P + + AYK+ GD +G AI
Sbjct: 181 GHGTHVAGSALADGGTGNGVYGVA--------PDADLWAYKVLGDDGSGYADDIAAAIRH 232
Query: 160 AFDDAISDGVDV-ITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVC 218
A D A + V I +SLG S + + + ++ KG+L AAGN GP S+
Sbjct: 233 AGDQATALNTKVVINMSLGSSGESSLITNAVN----YSYNKGVLIIAAAGNSGPYQGSIG 288
Query: 219 SGAPWLVSVAATSIDRQ 235
+ +VA +++ +
Sbjct: 289 YPGALVNAVAVAALENK 305
>SUBT_BACLI (P00780) Subtilisin Carlsberg precursor (EC 3.4.21.62)
Length = 379
Score = 42.4 bits (98), Expect = 0.002
Identities = 36/139 (25%), Positives = 59/139 (41%), Gaps = 15/139 (10%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
++G + G+ GH TH A T + G A PS + A K+
Sbjct: 149 VVGGASFVAGEAYNTDGNGHGTHVAGTVAALD-NTTGVLGVA--------PSVSLYAVKV 199
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
+G SG I++ + A ++G+DVI +SLG S + + +A +G++
Sbjct: 200 LNSSGSGTYSG--IVSGIEWATTNGMDVINMSLGGPSGSTAMKQAVD----NAYARGVVV 253
Query: 204 TQAAGNFGPIPSSVCSGAP 222
AAGN G ++ G P
Sbjct: 254 VAAAGNSGSSGNTNTIGYP 272
>SUBD_BACLI (P00781) Subtilisin DY (EC 3.4.21.62)
Length = 274
Score = 42.4 bits (98), Expect = 0.002
Identities = 39/140 (27%), Positives = 56/140 (39%), Gaps = 15/140 (10%)
Query: 83 KIIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYK 142
K++G + G+ GH TH A T + G A P+ + A K
Sbjct: 43 KVVGGASFVSGESYNTDGNGHGTHVAGTVAALD-NTTGVLGVA--------PNVSLYAIK 93
Query: 143 ICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGIL 202
+ +G S AI++ + A +G+DVI +SLG S L + A GI+
Sbjct: 94 VLNSSGSGTYS--AIVSGIEWATQNGLDVINMSLGGPSGSTALKQAVD----KAYASGIV 147
Query: 203 TTQAAGNFGPIPSSVCSGAP 222
AAGN G S G P
Sbjct: 148 VVAAAGNSGSSGSQNTIGYP 167
>SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 40.8 bits (94), Expect = 0.006
Identities = 45/172 (26%), Positives = 70/172 (40%), Gaps = 15/172 (8%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
I G + G+ S + GH TH A T + G A PS+ + A K+
Sbjct: 43 IRGGASFVPGEPSTQDGNGHGTHVAGTIAALN-NSIGVLGVA--------PSAELYAVKV 93
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
G +GR + +I + A ++G+ V +SLG S L A+ S A +G+L
Sbjct: 94 LGA--DGRGAISSIAQGLEWAGNNGMHVANLSLGSPSPSATLEQ--AVNS--ATSRGVLV 147
Query: 204 TQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A+GN G S + ++V AT + G G V +N+
Sbjct: 148 VAASGNSGASSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNV 199
>SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 40.0 bits (92), Expect = 0.010
Identities = 45/172 (26%), Positives = 69/172 (39%), Gaps = 15/172 (8%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
I G + G+ S + GH TH A T + G A PS+ + A K+
Sbjct: 43 IRGGASFVPGEPSTQDGNGHGTHVAGTIAALN-NSIGVLGVA--------PSAELYAVKV 93
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
G +G S +I + A ++G+ V +SLG S L A+ S A +G+L
Sbjct: 94 LGASGSGSVS--SIAQGLEWAGNNGMHVANLSLGSPSPSATLEQ--AVNS--ATSRGVLV 147
Query: 204 TQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A+GN G S + ++V AT + G G V +N+
Sbjct: 148 VAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNV 199
>PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-)
Length = 269
Score = 40.0 bits (92), Expect = 0.010
Identities = 45/172 (26%), Positives = 69/172 (39%), Gaps = 15/172 (8%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
I G + G+ S + GH TH A T + G A PS+ + A K+
Sbjct: 43 IRGGASFVPGEPSTQDGNGHGTHVAGTIAALN-NSIGVLGVA--------PSAELYAVKV 93
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
G +G S +I + A ++G+ V +SLG S L A+ S A +G+L
Sbjct: 94 LGASGSGSVS--SIAQGLEWAGNNGMHVANLSLGSPSPSATLEQ--AVNS--ATSRGVLV 147
Query: 204 TQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A+GN G S + ++V AT + G G V +N+
Sbjct: 148 VAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNV 199
>ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 40.0 bits (92), Expect = 0.010
Identities = 45/172 (26%), Positives = 69/172 (39%), Gaps = 15/172 (8%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
I G + G+ S + GH TH A T + G A PS+ + A K+
Sbjct: 154 IRGGASFVPGEPSTQDGNGHGTHVAGTIAALN-NSIGVLGVA--------PSAELYAVKV 204
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
G +G S +I + A ++G+ V +SLG S L A+ S A +G+L
Sbjct: 205 LGASGSGSVS--SIAQGLEWAGNNGMHVANLSLGSPSPSATLEQ--AVNS--ATSRGVLV 258
Query: 204 TQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A+GN G S + ++V AT + G G V +N+
Sbjct: 259 VAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNV 310
>SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline
mesentericopeptidase)
Length = 275
Score = 39.7 bits (91), Expect = 0.014
Identities = 37/120 (30%), Positives = 51/120 (41%), Gaps = 15/120 (12%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFD 162
H TH A T + G A PSS + A K+ +G+ S I+ +
Sbjct: 64 HGTHVAGTIAALN-NSIGVLGVA--------PSSALYAVKVLDSTGSGQYSW--IINGIE 112
Query: 163 DAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
AIS+ +DVI +SLG S L + A+ GI+ AAGN G S+ G P
Sbjct: 113 WAISNNMDVINMSLGGPTGSTALKTVVD----KAVSSGIVVAAAAGNEGSSGSTSTVGYP 168
>SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62)
(Subtilisin Novo) (Subtilisin DFE) (Alkaline protease)
Length = 382
Score = 39.3 bits (90), Expect = 0.018
Identities = 36/120 (30%), Positives = 52/120 (43%), Gaps = 15/120 (12%)
Query: 103 HRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFD 162
H TH A T + G A PS+ + A K+ G +G+ S I+ +
Sbjct: 171 HGTHVAGTVAALN-NSIGVLGVA--------PSASLYAVKVLGADGSGQYSW--IINGIE 219
Query: 163 DAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
AI++ +DVI +SLG S L + A+ G++ AAGN G SS G P
Sbjct: 220 WAIANNMDVINMSLGGPSGSAALKAAVD----KAVASGVVVVAAAGNEGTSGSSSTVGYP 275
>SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62)
Length = 381
Score = 38.9 bits (89), Expect = 0.023
Identities = 31/92 (33%), Positives = 44/92 (47%), Gaps = 6/92 (6%)
Query: 131 GGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIA 190
G PS+ + A K+ +G+ S I+ + AIS+ +DVI +SLG S L +
Sbjct: 189 GVSPSASLYAVKVLDSTGSGQYSW--IINGIEWAISNNMDVINMSLGGPSGSTALKTVVD 246
Query: 191 IGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
A+ GI+ AAGN G SS G P
Sbjct: 247 ----KAVSSGIVVAAAAGNEGSSGSSSTVGYP 274
>SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC
3.4.21.62)
Length = 381
Score = 38.9 bits (89), Expect = 0.023
Identities = 31/92 (33%), Positives = 44/92 (47%), Gaps = 6/92 (6%)
Query: 131 GGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIA 190
G PS+ + A K+ +G+ S I+ + AIS+ +DVI +SLG S L +
Sbjct: 189 GVSPSASLYAVKVLDSTGSGQYSW--IINGIEWAISNNMDVINMSLGGPSGSTALKTVVD 246
Query: 191 IGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
A+ GI+ AAGN G SS G P
Sbjct: 247 ----KAVSSGIVVAAAAGNEGSSGSSSTVGYP 274
>PLS_PYRFU (P72186) Pyrolysin precursor (EC 3.4.21.-)
Length = 1398
Score = 38.9 bits (89), Expect = 0.023
Identities = 42/151 (27%), Positives = 65/151 (42%), Gaps = 26/151 (17%)
Query: 102 GHRTHTASTAGGREVEDVSF----------------YGF-----AKGTARGGVPSSRIAA 140
GH TH A T G + + ++ YG+ T +G P ++I A
Sbjct: 364 GHGTHVAGTVAGYDSNNDAWDWLSMYSGEWEVFSRLYGWDYTNVTTDTVQGVAPGAQIMA 423
Query: 141 YKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEK- 199
++ +GR S I+ A + G DVI++SLG DP ++ EK
Sbjct: 424 IRVLRS--DGRGSMWDIIEGMTYAATHGADVISMSLGGNAPYLDGTDPESVAVDELTEKY 481
Query: 200 GILTTQAAGNFGPIPSSVCSGAPWLVSVAAT 230
G++ AAGN GP + V G+P + + A T
Sbjct: 482 GVVFVIAAGNEGPGINIV--GSPGVATKAIT 510
>ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 38.9 bits (89), Expect = 0.023
Identities = 44/172 (25%), Positives = 69/172 (39%), Gaps = 15/172 (8%)
Query: 84 IIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKI 143
I G + G+ S + GH TH A T + G A P++ + A K+
Sbjct: 154 IRGGASFVPGEPSTQDGNGHGTHVAGTIAALN-NSIGVLGVA--------PNAELYAVKV 204
Query: 144 CGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILT 203
G +G S +I + A ++G+ V +SLG S L A+ S A +G+L
Sbjct: 205 LGASGSGSVS--SIAQGLEWAGNNGMHVANLSLGSPSPSATLEQ--AVNS--ATSRGVLV 258
Query: 204 TQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A+GN G S + ++V AT + G G V +N+
Sbjct: 259 VAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNV 310
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,679,101
Number of Sequences: 164201
Number of extensions: 2259423
Number of successful extensions: 4372
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 4349
Number of HSP's gapped (non-prelim): 61
length of query: 395
length of database: 59,974,054
effective HSP length: 112
effective length of query: 283
effective length of database: 41,583,542
effective search space: 11768142386
effective search space used: 11768142386
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148652.13


