

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.5 - phase: 0
(122 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
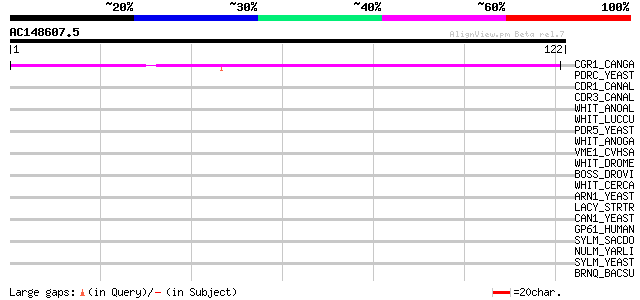
Score E
Sequences producing significant alignments: (bits) Value
CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1 (Pleom... 41 5e-04
PDRC_YEAST (Q02785) ATP-dependent permease PDR12 38 0.005
CDR1_CANAL (P43071) Multidrug resistance protein CDR1 38 0.005
CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3 36 0.013
WHIT_ANOAL (Q16928) White protein 36 0.017
WHIT_LUCCU (Q05360) White protein 35 0.023
PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin 33 0.087
WHIT_ANOGA (Q27256) White protein 33 0.11
VME1_CVHSA (P59596) E1 glycoprotein (Matrix glycoprotein) (Membr... 32 0.19
WHIT_DROME (P10090) White protein 32 0.25
BOSS_DROVI (Q24738) Bride of sevenless protein precursor 32 0.25
WHIT_CERCA (Q17320) White protein 32 0.33
ARN1_YEAST (P38731) Siderophore iron transporter ARN1 (Ferrichro... 31 0.43
LACY_STRTR (P23936) Lactose permease (Lactose-proton symporter) ... 30 1.3
CAN1_YEAST (P04817) Arginine permease 29 1.6
GP61_HUMAN (Q9BZJ8) Probable G protein-coupled receptor GPR61 (B... 28 2.8
SYLM_SACDO (P13503) Leucyl-tRNA synthetase, mitochondrial precur... 28 3.6
NULM_YARLI (Q9B6D4) NADH-ubiquinone oxidoreductase chain 4L (EC ... 28 3.6
SYLM_YEAST (P11325) Leucyl-tRNA synthetase, mitochondrial precur... 28 4.8
BRNQ_BACSU (P94499) Branched-chain amino acid transport system c... 28 4.8
>CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1
(Pleomorphic drug resistance homolog)
Length = 1542
Score = 40.8 bits (94), Expect = 5e-04
Identities = 30/122 (24%), Positives = 60/122 (48%), Gaps = 3/122 (2%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGA-LFYGCIVILFIGVA 59
+K ++ V +F++ + MA I ++F + + + S A + GA +F+ + F +
Sbjct: 515 IKNSASVTLFQVFGNSAMAFILGSMFYKIQ--KGSSADTFYFRGAAMFFAILFNAFSSLL 572
Query: 60 ELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIG 119
E+ + P+ K R Y + P A A + I +IP V ++ I+ Y+++ F G
Sbjct: 573 EIFSLYEARPITEKHRTYSLYHPSADAFASVISEIPPKIVTAILFNIIFYFLVNFRRDAG 632
Query: 120 RY 121
R+
Sbjct: 633 RF 634
>PDRC_YEAST (Q02785) ATP-dependent permease PDR12
Length = 1511
Score = 37.7 bits (86), Expect = 0.005
Identities = 26/101 (25%), Positives = 44/101 (42%), Gaps = 1/101 (0%)
Query: 12 LCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGA-LFYGCIVILFIGVAELSMVVSRLPV 70
L I A+I ++F + + S G G LFY + +AE+ S PV
Sbjct: 515 LSSFLIKALIIGSMFHKIDDKSQSTTAGAYSRGGMLFYVLLFASVTSLAEIGNSFSSRPV 574
Query: 71 FYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYV 111
K + Y + A +L I + P FV + + ++TY++
Sbjct: 575 IVKHKSYSMYHLSAESLQEIITEFPTKFVAIVILCLITYWI 615
>CDR1_CANAL (P43071) Multidrug resistance protein CDR1
Length = 1501
Score = 37.7 bits (86), Expect = 0.005
Identities = 27/123 (21%), Positives = 53/123 (42%), Gaps = 7/123 (5%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYV--GALFYGCIVILFIGV 58
MK + + IF + +M +I ++F S G Y A+F+ + F +
Sbjct: 508 MKGDPSIPIFSVFGQLVMGLILSSVFYNL-----SQTTGSFYYRGAAMFFAVLFNAFSSL 562
Query: 59 AELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYI 118
E+ + P+ K + Y + P A AL + I ++P+ + + Y+++ F
Sbjct: 563 LEIMSLFEARPIVEKHKKYALYRPSADALASIISELPVKLAMSMSFNFVFYFMVNFRRNP 622
Query: 119 GRY 121
GR+
Sbjct: 623 GRF 625
>CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3
Length = 1501
Score = 36.2 bits (82), Expect = 0.013
Identities = 21/97 (21%), Positives = 44/97 (44%), Gaps = 3/97 (3%)
Query: 18 MAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGY 77
MA+I ++F + + S + ++Y + + V E+ + + K R Y
Sbjct: 516 MALILSSVFYNLQPNSSSFYYR---TSVMYYALLFNAYSSVLEIYNMYEGRAIVQKHREY 572
Query: 78 LFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+PP A A+ + I PL V ++ ++ Y+++ F
Sbjct: 573 ALYPPMADAIGSIISDFPLKVVCSVLFNLILYFMVNF 609
>WHIT_ANOAL (Q16928) White protein
Length = 709
Score = 35.8 bits (81), Expect = 0.017
Identities = 29/112 (25%), Positives = 54/112 (47%), Gaps = 4/112 (3%)
Query: 11 KLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGV-AELSMVVSRLP 69
+L Q A++A + +I+ + +D V + + G+LF + F V A +++ + LP
Sbjct: 456 RLLQTAMVASLIGSIYFGQVLDQDGVMNIN---GSLFLFLTNMTFQNVFAVINVFSAELP 512
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
VF +++ + Y L I ++PL V+ +TY +IG I Y
Sbjct: 513 VFLREKRSRLYRVDTYFLGKTIAELPLFIAVPFVFTSITYPMIGLKAAISHY 564
>WHIT_LUCCU (Q05360) White protein
Length = 677
Score = 35.4 bits (80), Expect = 0.023
Identities = 29/112 (25%), Positives = 54/112 (47%), Gaps = 4/112 (3%)
Query: 11 KLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGV-AELSMVVSRLP 69
+L Q ++A++ IFL M + V + + GA+F + F V A +++ S LP
Sbjct: 425 RLIQTTMVAVLIGLIFLNQPMTQVGVMNIN---GAIFLFLTNMTFQNVFAVINVFTSELP 481
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
VF ++ + Y L + ++PL V +++ + Y +IG P I +
Sbjct: 482 VFMRETRSRLYRCDTYFLGKTLAELPLFLVVPFLFIAIAYPMIGLRPGITHF 533
>PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin
Length = 1511
Score = 33.5 bits (75), Expect = 0.087
Identities = 23/114 (20%), Positives = 50/114 (43%), Gaps = 1/114 (0%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
++ N +F + MA+I ++F + M + + A+F+ + F + E
Sbjct: 516 LRNNIGFTLFMILGNCSMALILGSMFFKI-MKKGDTSTFYFRGSAMFFAILFNAFSSLLE 574
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+ + P+ K R Y + P A A + + +IP + + I+ Y+++ F
Sbjct: 575 IFSLYEARPITEKHRTYSLYHPSADAFASVLSEIPSKLIIAVCFNIIFYFLVDF 628
>WHIT_ANOGA (Q27256) White protein
Length = 695
Score = 33.1 bits (74), Expect = 0.11
Identities = 27/104 (25%), Positives = 52/104 (49%), Gaps = 4/104 (3%)
Query: 11 KLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGV-AELSMVVSRLP 69
+L Q A++A + +I+ + +D V + + G+LF + F V A +++ + LP
Sbjct: 443 RLLQTAMVATLIGSIYFGQVLDQDGVMNIN---GSLFLFLTNMTFQNVFAVINVFSAELP 499
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIG 113
VF +++ + Y L I ++PL V+ +TY +IG
Sbjct: 500 VFLREKRSRLYRVDTYFLGKTIAELPLFIAVPFVFTSITYPMIG 543
>VME1_CVHSA (P59596) E1 glycoprotein (Matrix glycoprotein) (Membrane
glycoprotein)
Length = 221
Score = 32.3 bits (72), Expect = 0.19
Identities = 32/111 (28%), Positives = 50/111 (44%), Gaps = 11/111 (9%)
Query: 3 RNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS 62
RN F+YI KL L ++ + + F+ ++R + G I A+ CIV G+ LS
Sbjct: 41 RNRFLYIIKLVFLWLLWPVTLACFVLAAVYRINWVTGGI---AIAMACIV----GLMWLS 93
Query: 63 MVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIG 113
V+ +F + R F P L L +PL V ++ + VIG
Sbjct: 94 YFVASFRLFARTRSMWSFNPETNIL----LNVPLRGTIVTRPLMESELVIG 140
>WHIT_DROME (P10090) White protein
Length = 687
Score = 32.0 bits (71), Expect = 0.25
Identities = 27/104 (25%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Query: 11 KLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGV-AELSMVVSRLP 69
+L Q ++A++ IFL ++ + V + + GA+F + F V A +++ S LP
Sbjct: 434 RLIQTTMVAILIGLIFLGQQLTQVGVMNIN---GAIFLFLTNMTFQNVFATINVFTSELP 490
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIG 113
VF ++ + Y L I ++PL V+ + Y +IG
Sbjct: 491 VFMREARSRLYRCDTYFLGKTIAELPLFLTVPLVFTAIAYPMIG 534
>BOSS_DROVI (Q24738) Bride of sevenless protein precursor
Length = 893
Score = 32.0 bits (71), Expect = 0.25
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 6/69 (8%)
Query: 12 LCQLAIMAMIAMTIFLRTEMH--RDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLP 69
+C L++ + M++ L MH +SV+ +IY G +G + F+ + L VS +P
Sbjct: 654 ICVLSVFVQVGMSVQLLVVMHLASESVSCENIYYGRWLWGLLAYDFLLLCSL---VSLVP 710
Query: 70 VFYK-QRGY 77
Y+ QR Y
Sbjct: 711 FIYRSQRNY 719
>WHIT_CERCA (Q17320) White protein
Length = 679
Score = 31.6 bits (70), Expect = 0.33
Identities = 26/112 (23%), Positives = 53/112 (47%), Gaps = 4/112 (3%)
Query: 11 KLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIG-VAELSMVVSRLP 69
+L Q ++A++ IFL ++ + V + + GA+F + F A +++ + LP
Sbjct: 426 RLLQTTMVAVLIGLIFLGQQLTQVGVMNIN---GAIFLFLTNMTFQNSFATITVFTTELP 482
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
VF ++ + Y L I ++PL V ++ + Y +IG P + +
Sbjct: 483 VFMRETRSRLYRCDTYFLGKTIAELPLFLVVPFLFTAIAYPLIGLRPGVDHF 534
>ARN1_YEAST (P38731) Siderophore iron transporter ARN1 (Ferrichrome
permease)
Length = 627
Score = 31.2 bits (69), Expect = 0.43
Identities = 21/76 (27%), Positives = 33/76 (42%), Gaps = 7/76 (9%)
Query: 16 AIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQR 75
A++ + TI A G I+ A + G I+IL I +++ S + RL
Sbjct: 146 AVILYVVGTIIQSQAYDVQRYAAGAIFYNAGYVGVILILLIILSDFSSLKWRLL------ 199
Query: 76 GYLFFPPWAYALPAWI 91
Y F P W + + WI
Sbjct: 200 -YQFVPTWPFIINTWI 214
>LACY_STRTR (P23936) Lactose permease (Lactose-proton symporter)
(Lactose transport protein)
Length = 634
Score = 29.6 bits (65), Expect = 1.3
Identities = 12/36 (33%), Positives = 18/36 (49%)
Query: 46 LFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFP 81
LFYGCI ++ G+ S+ + LP+ F P
Sbjct: 316 LFYGCIAVMLGGIGIFSIAGTSLPIILTAAELFFIP 351
>CAN1_YEAST (P04817) Arginine permease
Length = 590
Score = 29.3 bits (64), Expect = 1.6
Identities = 26/101 (25%), Positives = 39/101 (37%), Gaps = 19/101 (18%)
Query: 41 IYVGALFYGCIVIL-----FIGVAELSMVVSRL---PVFYKQRGYLFFPPWAYA------ 86
+++G+L Y L FI V V S+ P F GY+++ WA
Sbjct: 127 LFMGSLAYSVTQSLGEMATFIPVTSSFTVFSQRFLSPAFGAANGYMYWFSWAITFALELS 186
Query: 87 -----LPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRYE 122
+ W K+PL WVI+T + Y G +E
Sbjct: 187 VVGQVIQFWTYKVPLAAWISIFWVIITIMNLFPVKYYGEFE 227
>GP61_HUMAN (Q9BZJ8) Probable G protein-coupled receptor GPR61
(Biogenic amine receptor-like G-protein-coupled
receptor)
Length = 417
Score = 28.5 bits (62), Expect = 2.8
Identities = 25/102 (24%), Positives = 45/102 (43%), Gaps = 11/102 (10%)
Query: 6 FVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALF--YGCIVILFIGVAELSM 63
FV++F LC + ++A + T M ++ ++ ALF C + LF+ V +S+
Sbjct: 76 FVFVFHLCLVDLLAAL-------TLMPLAMLSSPALFDHALFGEVACRLYLFLSVCFVSL 128
Query: 64 VVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWV 105
+ + +R Y P Y + + + V V VWV
Sbjct: 129 AILSVSAINVERYYYVVHPMRYEVRMTLGLV--ASVLVGVWV 168
>SYLM_SACDO (P13503) Leucyl-tRNA synthetase, mitochondrial precursor
(EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS)
Length = 894
Score = 28.1 bits (61), Expect = 3.6
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 3/40 (7%)
Query: 53 ILFIGVAELSMVVSRLPVFYKQRGYLFFPP--W-AYALPA 89
+L IG + ++ L FYKQRGY P W A+ LPA
Sbjct: 61 VLHIGHLRVYVISDSLNRFYKQRGYNVIHPMGWDAFGLPA 100
>NULM_YARLI (Q9B6D4) NADH-ubiquinone oxidoreductase chain 4L (EC
1.6.5.3)
Length = 89
Score = 28.1 bits (61), Expect = 3.6
Identities = 22/73 (30%), Positives = 39/73 (53%), Gaps = 3/73 (4%)
Query: 6 FVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVV 65
FV+ + LA + + M + + + R+SV DI G+LF IVI+ + E ++ +
Sbjct: 15 FVFNRRNIILAFICLETMLLGINLILLRNSVLFDDIS-GSLF--AIVIIILAGVESAIGL 71
Query: 66 SRLPVFYKQRGYL 78
S L +Y+ RG +
Sbjct: 72 SLLVSYYRLRGVI 84
>SYLM_YEAST (P11325) Leucyl-tRNA synthetase, mitochondrial precursor
(EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS)
Length = 894
Score = 27.7 bits (60), Expect = 4.8
Identities = 23/75 (30%), Positives = 35/75 (46%), Gaps = 4/75 (5%)
Query: 33 RDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPP--W-AYALPA 89
+D++ G Y+ F L IG + ++ L FYKQ+GY P W A+ LPA
Sbjct: 41 QDTLNSGSKYILCQFPYPSGALHIGHLRVYVISDSLNRFYKQKGYNVIHPMGWDAFGLPA 100
Query: 90 WILKIPLTFVEVAVW 104
I + + A+W
Sbjct: 101 ENAAIERS-INPAIW 114
>BRNQ_BACSU (P94499) Branched-chain amino acid transport system
carrier protein brnQ (Branched-chain amino acid uptake
carrier brnQ)
Length = 440
Score = 27.7 bits (60), Expect = 4.8
Identities = 25/88 (28%), Positives = 42/88 (47%), Gaps = 9/88 (10%)
Query: 15 LAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSM---VVSR---- 67
L M IA+++ T +H ++Y G+L + I+ LF G+ + VV+R
Sbjct: 348 LLTMYPIAISLIFLTFLHSVFKGKTEVYQGSLLFAFIISLFDGLKAAGIKIEVVNRIFTQ 407
Query: 68 -LPVFYKQRGYLFFPPWAYALPAWILKI 94
LP++ G+L P A + +IL I
Sbjct: 408 ILPMYNIGLGWL-IPAIAGGICGYILSI 434
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.336 0.149 0.486
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,360,048
Number of Sequences: 164201
Number of extensions: 472924
Number of successful extensions: 1562
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 1550
Number of HSP's gapped (non-prelim): 28
length of query: 122
length of database: 59,974,054
effective HSP length: 98
effective length of query: 24
effective length of database: 43,882,356
effective search space: 1053176544
effective search space used: 1053176544
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148607.5


