

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.15 - phase: 0
(174 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
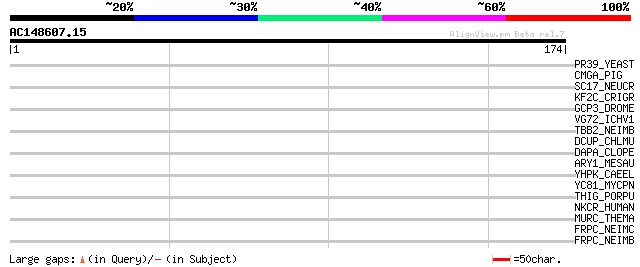
Score E
Sequences producing significant alignments: (bits) Value
PR39_YEAST (P39682) Pre-mRNA processing protein PRP39 39 0.008
CMGA_PIG (P04404) Chromogranin A precursor (CgA) [Contains: Panc... 31 1.3
SC17_NEUCR (Q9P6A5) Probable vesicular-fusion protein sec17 homolog 30 3.6
KF2C_CRIGR (P70096) Kinesin-like protein KIF2C (Mitotic centrome... 30 3.6
GCP3_DROME (Q9XYP8) Gamma-tubulin complex component 3 homolog (G... 29 4.8
VG72_ICHV1 (Q00103) Hypothetical gene 72 protein 29 6.2
TBB2_NEIMB (Q06988) Transferrin-binding protein 2 precursor (TBP-2) 29 6.2
DCUP_CHLMU (Q9PLH7) Uroporphyrinogen decarboxylase (EC 4.1.1.37)... 29 6.2
DAPA_CLOPE (Q8XJ56) Dihydrodipicolinate synthase (EC 4.2.1.52) (... 29 6.2
ARY1_MESAU (P50292) Arylamine N-acetyltransferase 1 (EC 2.3.1.5)... 29 6.2
YHPK_CAEEL (O62518) Hypothetical protein ZK795.3 in chromosome IV 28 8.1
YC81_MYCPN (P75496) Hypothetical lipoprotein MPN281 precursor (A... 28 8.1
THIG_PORPU (P51361) Thiazole biosynthesis protein thiG 28 8.1
NKCR_HUMAN (P30414) NK-tumor recognition protein (Natural-killer... 28 8.1
MURC_THEMA (Q9WY73) UDP-N-acetylmuramate--L-alanine ligase (EC 6... 28 8.1
FRPC_NEIMC (P55127) Iron-regulated protein frpC 28 8.1
FRPC_NEIMB (Q9JYV5) Iron-regulated protein frpC 28 8.1
>PR39_YEAST (P39682) Pre-mRNA processing protein PRP39
Length = 629
Score = 38.5 bits (88), Expect = 0.008
Identities = 31/141 (21%), Positives = 53/141 (36%), Gaps = 51/141 (36%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
Q++W + IY I+E P Q R+F S+K+ + + L R D
Sbjct: 174 QKNWHNVQRIYEYIIEVPLHQYARFFTSYKKFLNEKNLKTTRNID--------------- 218
Query: 62 GVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPY 121
I +R K + ++I FE+ I++P+
Sbjct: 219 -------------------------------IVLR-----KTQTTVNEIWQFESKIKQPF 242
Query: 122 FHVRPLNIGELENWHNYLDFI 142
F++ + +LENW YL F+
Sbjct: 243 FNLGQVLNDDLENWSRYLKFV 263
>CMGA_PIG (P04404) Chromogranin A precursor (CgA) [Contains:
Pancreastatin; Parastatin; WE-14] (Fragment)
Length = 446
Score = 31.2 bits (69), Expect = 1.3
Identities = 25/78 (32%), Positives = 34/78 (43%), Gaps = 2/78 (2%)
Query: 31 KELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELE 90
+E AS PL+ L + A A EG QG A+ P+ EAE E
Sbjct: 169 EEAASTHPLASLPSKKRPGAQAEEDHEGPSQGPVDREKGPSAEQGPQAEREEEEEAEAGE 228
Query: 91 KYIAIREEMYKKAKEFDS 108
K A+ EE +++ FDS
Sbjct: 229 K--AVPEEEGPRSEAFDS 244
>SC17_NEUCR (Q9P6A5) Probable vesicular-fusion protein sec17 homolog
Length = 292
Score = 29.6 bits (65), Expect = 3.6
Identities = 32/151 (21%), Positives = 64/151 (42%), Gaps = 7/151 (4%)
Query: 9 AMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAA---AVAGVVSEGID-QGVE 64
A I+T L+ PN + ++FK + P + +R + A +AG + +
Sbjct: 64 AKIFTEKLKEPNDAANAMLDAFKVYRKDAPDNAVRCVEVAIKQYTMAGNFRRAASHKENQ 123
Query: 65 GEVHPDGADNSPKPASAGLTEAE--ELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYF 122
EV+ + N P+ A T AE E + +A+ +++ K + + F I + +
Sbjct: 124 AEVYENELQNKPEAIKAYTTAAEWYENDGAVALANKLWLKVADLSALAGDFFAAIEK-FE 182
Query: 123 HVRPLNIGELENWHNYLDFIEREGDLSKVTK 153
V ++G ++ ++ + G S TK
Sbjct: 183 KVAEASLGNNLMRYSVKEYFLKAGLCSLATK 213
>KF2C_CRIGR (P70096) Kinesin-like protein KIF2C (Mitotic
centromere-associated kinesin) (MCAK) (Kinesin-like
protein 6)
Length = 718
Score = 29.6 bits (65), Expect = 3.6
Identities = 19/64 (29%), Positives = 30/64 (46%), Gaps = 8/64 (12%)
Query: 59 IDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREE--------MYKKAKEFDSKI 110
+ + E +VHP + +S PA +E+EK REE K+A+E+DS
Sbjct: 162 VSEEAEEQVHPTRSTSSANPARRKSCIVKEMEKMKNKREEKRAQNSEIRIKRAQEYDSSF 221
Query: 111 IGFE 114
+E
Sbjct: 222 PNWE 225
>GCP3_DROME (Q9XYP8) Gamma-tubulin complex component 3 homolog
(Gamma ring complex protein 91) (dGrip91) (d91p)
Length = 917
Score = 29.3 bits (64), Expect = 4.8
Identities = 17/54 (31%), Positives = 28/54 (51%)
Query: 8 LAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
L ++Y I+E Q Y + FKEL + + LS++ E V G+ ++ I Q
Sbjct: 769 LEVVYENIIELEKWQSSFYKDCFKELNARKELSKIVEKSEKKGVYGLTNKMILQ 822
>VG72_ICHV1 (Q00103) Hypothetical gene 72 protein
Length = 1350
Score = 28.9 bits (63), Expect = 6.2
Identities = 38/129 (29%), Positives = 54/129 (41%), Gaps = 20/129 (15%)
Query: 44 TADEAAAVAGVVSEG------IDQGVEGEVHPDGADNSPKPASAG--LTEAEELEKYIAI 95
T A +AGV++EG + V+GE D + P G + +LE IA
Sbjct: 409 TGKSAQTIAGVLTEGGVAPYPLIMDVDGE--RDLPIDLPNQIITGRVIESLGDLEGDIAA 466
Query: 96 REEMYKKAKEFDSKIIGFETTIRRPYFHVRPLNIGELENWHNYLDFIEREGDL---SKVT 152
R+E+ KKA E RP + + I E W Y + GDL S+
Sbjct: 467 RKEVIKKAVE------ALRVYSYRPLMQLSVIGIFFDEGW-IYSVLEKPTGDLKTMSQTL 519
Query: 153 KLLSYKGTF 161
++LS GTF
Sbjct: 520 RVLSTTGTF 528
>TBB2_NEIMB (Q06988) Transferrin-binding protein 2 precursor (TBP-2)
Length = 599
Score = 28.9 bits (63), Expect = 6.2
Identities = 22/85 (25%), Positives = 41/85 (47%), Gaps = 7/85 (8%)
Query: 84 TEAEELEKYIAIREEMYKKAKEFDSKIIGFE--TTIRRPYFHVRPLNIGELENW-----H 136
+E +ELEK E + K ++ S+++G+ T +R Y ++ NI N
Sbjct: 107 SERDELEKKRGSSELIESKWEDGQSRVVGYTNFTYVRSGYVYLNKNNIDIKNNIVLFGPD 166
Query: 137 NYLDFIEREGDLSKVTKLLSYKGTF 161
YL + +E ++ ++YKGT+
Sbjct: 167 GYLYYKGKEPSKELPSEKITYKGTW 191
>DCUP_CHLMU (Q9PLH7) Uroporphyrinogen decarboxylase (EC 4.1.1.37)
(URO-D) (UPD)
Length = 334
Score = 28.9 bits (63), Expect = 6.2
Identities = 16/46 (34%), Positives = 25/46 (53%), Gaps = 3/46 (6%)
Query: 21 QQLDRYFNSFKELASNRPLSELRTADEA---AAVAGVVSEGIDQGV 63
+Q+ RY ++EL +NRPL EL E+ A + G G+D +
Sbjct: 22 RQVGRYMPQYQELKTNRPLKELFLDTESIIEATLLGPSLLGVDAAI 67
>DAPA_CLOPE (Q8XJ56) Dihydrodipicolinate synthase (EC 4.2.1.52)
(DHDPS)
Length = 291
Score = 28.9 bits (63), Expect = 6.2
Identities = 15/48 (31%), Positives = 26/48 (53%)
Query: 119 RPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFS 166
R ++ PL + +L + +N + E GDLS+V K+ G +I+S
Sbjct: 137 RTGMNITPLMLDKLADLNNVVAIKEASGDLSQVAKMAELCGDRIAIYS 184
>ARY1_MESAU (P50292) Arylamine N-acetyltransferase 1 (EC 2.3.1.5)
(Arylamide acetylase 1) (Arylamine N-acetyltransferase,
monomorphic) (MNAT) (N-acetyltransferase type 1) (NAT-1)
(AT-1)
Length = 290
Score = 28.9 bits (63), Expect = 6.2
Identities = 22/70 (31%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Query: 71 GADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPY-FHVRPLNI 129
G D PA LTE E IR E + +EF + + + T R+ Y F ++P I
Sbjct: 140 GKDQPQVPAIFRLTEENETWYLDQIRREQHVPNQEFVNSDLLEKNTYRKIYSFTLQPRTI 199
Query: 130 GELENWHNYL 139
+ E + YL
Sbjct: 200 EDFEYANTYL 209
>YHPK_CAEEL (O62518) Hypothetical protein ZK795.3 in chromosome IV
Length = 292
Score = 28.5 bits (62), Expect = 8.1
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 12/91 (13%)
Query: 76 PKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRP--LNIGELE 133
PKP S + E YI+ R +YK E D ++ E T P F ++P + +G LE
Sbjct: 205 PKPDSKRIITFSNSEDYISFRHHVYK--TENDGEV---ELTEAGPRFELKPYQIKLGTLE 259
Query: 134 NWHNYLDFIEREGDLSKVTKLLSYKGTFCSI 164
L E E L T + K TF SI
Sbjct: 260 T----LAAAEDEWVLRSYTN-TARKRTFLSI 285
>YC81_MYCPN (P75496) Hypothetical lipoprotein MPN281 precursor
(A65_orf377)
Length = 377
Score = 28.5 bits (62), Expect = 8.1
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 2/30 (6%)
Query: 133 ENWHNYLDFIER--EGDLSKVTKLLSYKGT 160
ENWH++LDF R + KV+ + + KGT
Sbjct: 288 ENWHDFLDFSTRVAKSFTEKVSNITNKKGT 317
>THIG_PORPU (P51361) Thiazole biosynthesis protein thiG
Length = 277
Score = 28.5 bits (62), Expect = 8.1
Identities = 13/37 (35%), Positives = 21/37 (56%)
Query: 78 PASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFE 114
P +AG AEE + ++ +E+ KK +FD+ I E
Sbjct: 85 PNTAGCESAEEAVRIASLGQEIVKKLGQFDNNFIKLE 121
>NKCR_HUMAN (P30414) NK-tumor recognition protein (Natural-killer
cells cyclophilin-related protein) (NK-TR protein)
Length = 1462
Score = 28.5 bits (62), Expect = 8.1
Identities = 19/82 (23%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 29 SFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAE- 87
+ K A++RP +++R D + + ++ + H +G+D+S +S+ + +E
Sbjct: 155 NLKTDAASRPYADVRVIDCGVLATKSIKDVFEKKRKKPTHSEGSDSSSNSSSSSESSSES 214
Query: 88 ELEKYIAIREEMYKKAKEFDSK 109
ELE + R + ++ K SK
Sbjct: 215 ELEHERSRRRKHKRRPKVKRSK 236
>MURC_THEMA (Q9WY73) UDP-N-acetylmuramate--L-alanine ligase (EC
6.3.2.8) (UDP-N-acetylmuramoyl-L-alanine synthetase)
Length = 457
Score = 28.5 bits (62), Expect = 8.1
Identities = 15/47 (31%), Positives = 25/47 (52%)
Query: 16 LENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQG 62
LEN L RY ++F++++ N L DE + G V+ G+ +G
Sbjct: 183 LENYGNSLTRYRSAFEKISRNTDLVVTFAEDELTSHLGDVTFGVKKG 229
>FRPC_NEIMC (P55127) Iron-regulated protein frpC
Length = 1829
Score = 28.5 bits (62), Expect = 8.1
Identities = 19/72 (26%), Positives = 35/72 (48%), Gaps = 1/72 (1%)
Query: 10 MIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHP 69
+IY I++N +Q +++ + KEL+ + AD +A A V E + Q + E +
Sbjct: 290 IIYNDIVDNTSQGIEKGVKAIKELSEKMKNAASDLADGSAEKAKQVVEDLAQAAK-EAYE 348
Query: 70 DGADNSPKPASA 81
+ + K A A
Sbjct: 349 NAKSTAEKAAQA 360
>FRPC_NEIMB (Q9JYV5) Iron-regulated protein frpC
Length = 1829
Score = 28.5 bits (62), Expect = 8.1
Identities = 19/72 (26%), Positives = 35/72 (48%), Gaps = 1/72 (1%)
Query: 10 MIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHP 69
+IY I++N +Q +++ + KEL+ + AD +A A V E + Q + E +
Sbjct: 290 IIYNDIVDNTSQGIEKGVKAIKELSEKMKNAASDLADGSAEKAKQVVEDLAQAAK-EAYE 348
Query: 70 DGADNSPKPASA 81
+ + K A A
Sbjct: 349 NAKSTAEKAAQA 360
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,941,253
Number of Sequences: 164201
Number of extensions: 889062
Number of successful extensions: 2892
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2886
Number of HSP's gapped (non-prelim): 18
length of query: 174
length of database: 59,974,054
effective HSP length: 103
effective length of query: 71
effective length of database: 43,061,351
effective search space: 3057355921
effective search space used: 3057355921
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148607.15


