

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
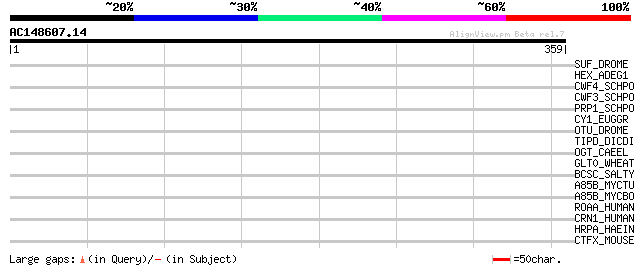
Query= AC148607.14 - phase: 0
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SUF_DROME (P25991) Suppressor of forked protein 38 0.035
HEX_ADEG1 (P42671) Hexon protein (Late protein 2) 36 0.13
CWF4_SCHPO (P87312) Cell cycle control protein cwf4 36 0.13
CWF3_SCHPO (Q9P7R9) Cell cycle control protein cwf3 35 0.38
PRP1_SCHPO (Q12381) Pre-mRNA splicing factor prp1 32 2.5
CY1_EUGGR (P20114) Cytochrome c1, heme protein 32 2.5
OTU_DROME (P10383) Ovarian tumor locus protein 31 4.2
TIPD_DICDI (O15736) Protein tipD 31 5.5
OGT_CAEEL (O18158) UDP-N-acetylglucosamine--peptide N-acetylgluc... 31 5.5
GLT0_WHEAT (P10387) Glutenin, high molecular weight subunit DY10... 31 5.5
BCSC_SALTY (Q8ZLB8) Cellulose synthase operon protein C 31 5.5
A85B_MYCTU (P31952) Antigen 85-B precursor (85B) (Extracellular ... 31 5.5
A85B_MYCBO (P12942) Antigen 85-B precursor (85B) (Extracellular ... 31 5.5
ROAA_HUMAN (Q99729) Heterogeneous nuclear ribonucleoprotein A/B ... 30 7.2
CRN1_HUMAN (Q9BZJ0) Crooked neck-like protein 1 (Crooked neck ho... 30 7.2
HRPA_HAEIN (P45018) ATP-dependent helicase hrpA homolog 30 9.4
CTFX_MOUSE (Q9D5U8) Protein C20orf152 homolog 30 9.4
>SUF_DROME (P25991) Suppressor of forked protein
Length = 733
Score = 38.1 bits (87), Expect = 0.035
Identities = 52/257 (20%), Positives = 99/257 (38%), Gaps = 48/257 (18%)
Query: 19 ERCVIACANYPEYWIRYVLCMEASESM--------------DLANNVLARASQVFVKRQP 64
E+C++ ++P W + ++ S + D N+L R+ + R
Sbjct: 282 EQCLLVLTHHPAVWHQASQFLDTSARVLTEKGDVQAAKIFADECANILERSINGVLNRNA 341
Query: 65 EIHLFCARFKEQAGDIVGARAAYQLVHT---------EISPGLLEAIIRHANMEHRLGKL 115
++ A F+E R Y+ VHT +I P L+ +++ R +
Sbjct: 342 LLYFAYADFEE-------GRLKYEKVHTMYNKLLQLPDIDPTLV--YVQYMKFARRAEGI 392
Query: 116 EDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKP 175
+ A S++++A + + H +F + Y S + E A I GL+ S
Sbjct: 393 KSARSIFKKAREDVRSRYH------IFVAAALMEYYCSKDKEIAFRIFELGLKRFGGSPE 446
Query: 176 LLEALLHFEAIQPQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLF 235
+ + + + + + F L + G S + E+ N FLEF +
Sbjct: 447 YVMCYIDYLSHLNEDNNTRVLFERVL----------SSGGLSPHKSVEVWNRFLEFESNI 496
Query: 236 GDVQSIKRAEDRHAKLF 252
GD+ SI + E R + +F
Sbjct: 497 GDLSSIVKVERRRSAVF 513
>HEX_ADEG1 (P42671) Hexon protein (Late protein 2)
Length = 942
Score = 36.2 bits (82), Expect = 0.13
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 8/55 (14%)
Query: 283 AQSPAQSVAGAYPNGPNQWPNYG-VQPQTWPATTQAQGQQWPAGYTQQQVNWFFP 336
+ +P + G+ P+ N N G + P++WP T QG+ WPA NW +P
Sbjct: 790 SSTPVVNDTGSQPSQDNVRNNSGFIAPRSWPVWTAQQGEAWPA-------NWPYP 837
>CWF4_SCHPO (P87312) Cell cycle control protein cwf4
Length = 674
Score = 36.2 bits (82), Expect = 0.13
Identities = 40/183 (21%), Positives = 81/183 (43%), Gaps = 22/183 (12%)
Query: 17 LYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCARF--- 73
++ER + + Y W++Y+ C + +++ A N+ RA V + P + ++
Sbjct: 92 VFERALDVDSTYIPLWLKYIECEMKNRNINHARNLFDRA----VTQLPRVDKLWYKYVYM 147
Query: 74 KEQAGDIVGARAAYQ-LVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGK 132
+E G+I G R ++ + E + IR ME R + E A +YE+ + +
Sbjct: 148 EEMLGNITGCRQVFERWLKWEPDENCWMSYIR---MERRYHENERARGIYERFVVV---- 200
Query: 133 EHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLE---NASLSKPLLEALLHFEAIQPQ 189
H + L +++RF GN+ R++ + ++ L++ A FE Q +
Sbjct: 201 -HPEVTNWL--RWARFEE-ECGNAANVRQVYLAAIDALGQEFLNERFFIAFAKFEIRQKE 256
Query: 190 PKR 192
+R
Sbjct: 257 YER 259
>CWF3_SCHPO (Q9P7R9) Cell cycle control protein cwf3
Length = 790
Score = 34.7 bits (78), Expect = 0.38
Identities = 38/158 (24%), Positives = 66/158 (41%), Gaps = 16/158 (10%)
Query: 57 QVFVKRQPEIHLFCARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLE 116
QV + + +I ++ +E G I R Y V E+ + ++ +AN+ E
Sbjct: 473 QVRLHKSSKIWMYYLDLEESVGTIETTRKLYDRVF-ELKIATPQVVVNYANLLEENAYFE 531
Query: 117 DAFSLYEQAIAIEKGKEHSQTLPMLFAQY----SRFVYLASG-NSEKAREILVGGLENA- 170
D+F +YE+ +A+ + P+ F + ++FV G + E+ R++ LE
Sbjct: 532 DSFKIYERGVAL-------FSYPVAFELWNLYLTKFVKRYQGTHMERTRDLFEQALEGCP 584
Query: 171 -SLSKPLLEALLHFEAIQPQPKRVDIDFLESLVVKFIT 207
SK + FE + KR I LE K T
Sbjct: 585 PEFSKSIYLLYADFEEKFGKAKR-SISILEKAADKVKT 621
>PRP1_SCHPO (Q12381) Pre-mRNA splicing factor prp1
Length = 906
Score = 32.0 bits (71), Expect = 2.5
Identities = 40/176 (22%), Positives = 65/176 (36%), Gaps = 10/176 (5%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+ + + E+ V +C W+ Y + + A N+L RA + + EI L
Sbjct: 561 ESVCSILEKAVESCPKAEILWLLYAKERKNVNDIAGARNILGRAFE-YNSNSEEIWLAAV 619
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
R + + RA L I G + ++E L + + A L E A+ I
Sbjct: 620 RIEFVNNE--NERARKLLARARIESGTERIWTKSISLERILDEKDRALQLLENALKI--- 674
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQ 187
H L M+ Q ++ E AR+ + G + S PL L E Q
Sbjct: 675 YPHYDKLYMMKGQ----IFEDKEQIELARDAYLAGTKVCPYSIPLWLLLAKLEEKQ 726
>CY1_EUGGR (P20114) Cytochrome c1, heme protein
Length = 243
Score = 32.0 bits (71), Expect = 2.5
Identities = 13/40 (32%), Positives = 26/40 (64%)
Query: 237 DVQSIKRAEDRHAKLFLPNRGLSELKKRHAEDFLASDKTK 276
D +S++R ++ + ++F P LS +K RH E F++ ++ K
Sbjct: 21 DWRSVRRGKEVYEQVFAPCHSLSFIKYRHFEAFMSKEEVK 60
>OTU_DROME (P10383) Ovarian tumor locus protein
Length = 853
Score = 31.2 bits (69), Expect = 4.2
Identities = 20/61 (32%), Positives = 29/61 (46%), Gaps = 11/61 (18%)
Query: 274 KTKVSRAY---SAQSPAQSVAGAYPNGPNQWPNY--------GVQPQTWPATTQAQGQQW 322
+TK SR S+ S Q V+G+ P Q+ NY G P WPA+ A +++
Sbjct: 495 RTKASRVQPQNSSSSQNQEVSGSAAPPPTQYMNYVPMIPSRPGHLPPPWPASPMAIAEEF 554
Query: 323 P 323
P
Sbjct: 555 P 555
>TIPD_DICDI (O15736) Protein tipD
Length = 612
Score = 30.8 bits (68), Expect = 5.5
Identities = 28/115 (24%), Positives = 52/115 (44%), Gaps = 6/115 (5%)
Query: 155 NSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKRVDIDFLESLVVKFITPNPENPG 214
N++ A IL+ +N L L+ + E I+ ++ D+D ++ L + + EN
Sbjct: 151 NADNASSILLLNDKNKDLQNELMSKEIEIERIRSTIQQ-DLDSIKRL--EMVVIEKEN-- 205
Query: 215 VASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKLFLPNRGLSELKKRHAEDF 269
S R+ELS++ EFL+ V +++ + +L + K A DF
Sbjct: 206 -VSQIIRDELSSLQTEFLHNESKVVKLEQENSSLVERWLRKKNEEASKMNEANDF 259
>OGT_CAEEL (O18158) UDP-N-acetylglucosamine--peptide
N-acetylglucosaminyltransferase (EC 2.4.1.-) (O-GlcNAc)
(OGT)
Length = 1151
Score = 30.8 bits (68), Expect = 5.5
Identities = 24/83 (28%), Positives = 38/83 (44%), Gaps = 3/83 (3%)
Query: 64 PEIHLFCARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYE 123
P+ + A ++ G +V A Y E+ P ++ AN++ GK+EDA LY
Sbjct: 397 PDAYCNLANALKEKGSVVEAEQMYMKA-LELCPTHADSQNNLANIKREQGKIEDATRLYL 455
Query: 124 QAIAI--EKGKEHSQTLPMLFAQ 144
+A+ I E HS +L Q
Sbjct: 456 KALEIYPEFAAAHSNLASILQQQ 478
>GLT0_WHEAT (P10387) Glutenin, high molecular weight subunit DY10
precursor
Length = 648
Score = 30.8 bits (68), Expect = 5.5
Identities = 16/45 (35%), Positives = 19/45 (41%), Gaps = 1/45 (2%)
Query: 284 QSPAQSVAGAYPNGPNQWPNYGVQPQTWPATTQAQGQQWPAGYTQ 328
Q P Q G YP P Q G QP+ W + Q Q +P Q
Sbjct: 228 QQPGQWQQGYYPTSPQQL-GQGQQPRQWQQSGQGQQGHYPTSLQQ 271
>BCSC_SALTY (Q8ZLB8) Cellulose synthase operon protein C
Length = 1143
Score = 30.8 bits (68), Expect = 5.5
Identities = 47/227 (20%), Positives = 94/227 (40%), Gaps = 23/227 (10%)
Query: 39 MEASESMDLANNVL----ARASQVFVKRQP---EIHLFCARFKEQAGDIVGARAAYQLVH 91
+++++ ++ AN + R ++ +++QP I L A + +Q GD ARAAY V
Sbjct: 557 LQSNQVLESANRLRDSGKEREAEALLRQQPPSTRIALTLADWAQQRGDNAAARAAYDAVL 616
Query: 92 TEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYL 151
PG ++A++ ++ G D + Q A+ SQ + + L
Sbjct: 617 AR-EPGNVDAMLGRVEIDIAQG---DNAAARAQLAALPA----SQITSINMQRRVALAQL 668
Query: 152 ASGNSEKAREILVGGLENASLSKPLLEALL------HFEAIQPQPKRVDIDFLESLVVKF 205
G+ A A P +E+ + F+A +P+R + +++V
Sbjct: 669 QLGDITAAARTFNRITPQAKAQPPSMESAMVLRDAAAFQAQTGEPQRALETYKDAMVAAA 728
Query: 206 ITP-NPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL 251
ITP P++ + R + + +L+ + D + R +D + L
Sbjct: 729 ITPVRPQDNDTFTRLTRNDEKDDWLK-RGVRSDAAELYRQQDLNVTL 774
>A85B_MYCTU (P31952) Antigen 85-B precursor (85B) (Extracellular
alpha-antigen) (Antigen 85 complex B) (Ag85B) (Mycolyl
transferase 85B) (EC 2.3.1.-) (Fibronectin-binding
protein B) (30 kDa extracellular protein)
Length = 325
Score = 30.8 bits (68), Expect = 5.5
Identities = 17/55 (30%), Positives = 25/55 (44%), Gaps = 1/55 (1%)
Query: 254 PNR-GLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGVQ 307
PN G + + E+F+ S K AY+A +V PNG + W +G Q
Sbjct: 256 PNELGGANIPAEFLENFVRSSNLKFQDAYNAAGGHNAVFNFPPNGTHSWEYWGAQ 310
>A85B_MYCBO (P12942) Antigen 85-B precursor (85B) (Extracellular
alpha-antigen) (Antigen 85 complex B) (Ag85B) (Mycolyl
transferase 85B) (EC 2.3.1.-) (Fibronectin-binding
protein B)
Length = 325
Score = 30.8 bits (68), Expect = 5.5
Identities = 17/55 (30%), Positives = 25/55 (44%), Gaps = 1/55 (1%)
Query: 254 PNR-GLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGVQ 307
PN G + + E+F+ S K AY+A +V PNG + W +G Q
Sbjct: 256 PNELGGANIPAEFLENFVRSSNLKFQDAYNAAGGHNAVFNFPPNGTHSWEYWGAQ 310
>ROAA_HUMAN (Q99729) Heterogeneous nuclear ribonucleoprotein A/B
(hnRNP A/B) (APOBEC-1 binding protein 1) (ABBP-1)
Length = 331
Score = 30.4 bits (67), Expect = 7.2
Identities = 49/227 (21%), Positives = 84/227 (36%), Gaps = 51/227 (22%)
Query: 152 ASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKRVD-----------IDFLES 200
AS N E A ++ VGGL + K L + F + ++D I F ++
Sbjct: 60 ASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDA 119
Query: 201 LVVKFITPNPE--------NPGVASATEREELSNIFL--------------EFLNLFGDV 238
V+ + E +P A A +++ + IF+ E+ FG++
Sbjct: 120 ASVEKVLDQKEHRLDGRVIDPKKAMAMKKDPVKKIFVGGLNPESPTEEKIREYFGEFGEI 179
Query: 239 QSIKRAED-----RHAKLFLPNRGLSELKKRHAEDF--LASDKTKVSRAYSAQSPAQSVA 291
++I+ D R +F+ + +KK + F ++ K ++ A + Q
Sbjct: 180 EAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKCEIKVAQPKEVYQQQQY 239
Query: 292 GAYPNGPNQWPNYGV--------QPQTWPATTQAQGQQWPAGYTQQQ 330
G+ G N G Q Q+W Q G W GY QQ
Sbjct: 240 GSGGRGNRNRGNRGSGGGGGGGGQSQSW---NQGYGNYWNQGYGYQQ 283
>CRN1_HUMAN (Q9BZJ0) Crooked neck-like protein 1 (Crooked neck
homolog) (hCrn) (CGI-201) (MSTP021)
Length = 848
Score = 30.4 bits (67), Expect = 7.2
Identities = 24/87 (27%), Positives = 36/87 (40%), Gaps = 3/87 (3%)
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
VD+ +YER V+ + WI+Y E A V RA + F + HL+
Sbjct: 359 VDRARTIYERFVLVHPDVKN-WIKYARFEEKHAYFAHARKVYERAVEFFGDEHMDEHLYV 417
Query: 71 --ARFKEQAGDIVGARAAYQLVHTEIS 95
A+F+E + R Y+ IS
Sbjct: 418 AFAKFEENQKEFERVRVIYKYALDRIS 444
Score = 30.0 bits (66), Expect = 9.4
Identities = 28/140 (20%), Positives = 61/140 (43%), Gaps = 14/140 (10%)
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
V+ +++R + ++W +Y E ++ A V R +++ QPE +
Sbjct: 292 VNHARNIWDRAITTLPRVNQFWYKYTYMEEMLGNVAGARQVFER----WMEWQPEEQAWH 347
Query: 71 A--RFKEQAGDIVGARAAYQ---LVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQA 125
+ F+ + ++ AR Y+ LVH ++ I++A E + A +YE+A
Sbjct: 348 SYINFELRYKEVDRARTIYERFVLVHPDVKNW-----IKYARFEEKHAYFAHARKVYERA 402
Query: 126 IAIEKGKEHSQTLPMLFAQY 145
+ + + L + FA++
Sbjct: 403 VEFFGDEHMDEHLYVAFAKF 422
>HRPA_HAEIN (P45018) ATP-dependent helicase hrpA homolog
Length = 1304
Score = 30.0 bits (66), Expect = 9.4
Identities = 16/58 (27%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Query: 74 KEQAGDIVGARAAYQLVHTEISPGLLEAI-IRHANMEHRLGKLEDAFSLYEQAIAIEK 130
+E + I +A YQ +HT + GLL I ++ A + LG F+++ ++ +K
Sbjct: 612 REMSLPINSEKAEYQQIHTALLSGLLSHIGLKEAEKQQYLGARNAHFAIFPNSVLFKK 669
>CTFX_MOUSE (Q9D5U8) Protein C20orf152 homolog
Length = 556
Score = 30.0 bits (66), Expect = 9.4
Identities = 31/92 (33%), Positives = 43/92 (46%), Gaps = 12/92 (13%)
Query: 221 REELSNIFLEFLNL--FGDVQSIKRAEDRHAKLFLPNRGLSELKKRHAEDFLASDK---- 274
R++L + +E LN F VQ+ K+ H LFL L + K+H FLA K
Sbjct: 469 RKDLLRLLVEPLNKSPFIPVQTKKKEIYNHKSLFLDLCSLEKKVKQHYPIFLAPQKYLPP 528
Query: 275 TKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGV 306
+V +A SA P Q + P Q+ N GV
Sbjct: 529 LRVVQAISA--PRQKIQELLP----QYKNAGV 554
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,831,440
Number of Sequences: 164201
Number of extensions: 1660984
Number of successful extensions: 4127
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4119
Number of HSP's gapped (non-prelim): 21
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC148607.14


