

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.6 + phase: 0
(243 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
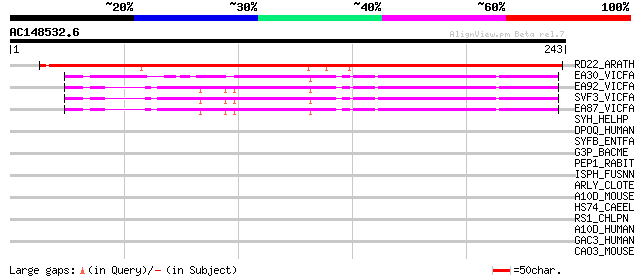
RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor 244 2e-64
EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor 116 6e-26
EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor 115 1e-25
SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1) 111 1e-24
EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor 111 1e-24
SYH_HELHP (Q7VF68) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 38 0.019
DPOQ_HUMAN (O75417) DNA polymerase theta (EC 2.7.7.7) (DNA polym... 33 0.81
SYFB_ENTFA (Q836J5) Phenylalanyl-tRNA synthetase beta chain (EC ... 32 1.8
G3P_BACME (P23722) Glyceraldehyde-3-phosphate dehydrogenase (EC ... 32 1.8
PEP1_RABIT (P28712) Pepsin II-1 precursor (EC 3.4.23.1) (Pepsin A) 31 2.4
ISPH_FUSNN (Q8RI52) 4-hydroxy-3-methylbut-2-enyl diphosphate red... 31 2.4
ARLY_CLOTE (P59616) Argininosuccinate lyase (EC 4.3.2.1) (Argino... 31 2.4
A10D_MOUSE (Q8K2X1) Potential phospholipid-transporting ATPase V... 31 2.4
HS74_CAEEL (P20163) Heat shock 70 kDa protein D precursor 30 4.0
RS1_CHLPN (Q9Z8M3) 30S ribosomal protein S1 30 5.3
A10D_HUMAN (Q9P241) Potential phospholipid-transporting ATPase V... 30 6.9
GAC3_HUMAN (Q99928) Gamma-aminobutyric-acid receptor gamma-3 sub... 29 9.0
CAO3_MOUSE (Q9EPL9) Acyl-coenzyme A oxidase 3, peroxisomal (EC 1... 29 9.0
>RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor
Length = 392
Score = 244 bits (622), Expect = 2e-64
Identities = 128/237 (54%), Positives = 157/237 (66%), Gaps = 9/237 (3%)
Query: 14 FIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGA-----TFLPREVANSIP 68
F+Y Y A TQLHD N LFF EKDL G ++N++F FLPR A ++P
Sbjct: 156 FVY-NYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGGKTAFLPRGEAETVP 214
Query: 69 FSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG 128
F S K L FS++ GS E+E +K+ I CE + GEEK C TSLESMVDF+ SKLG
Sbjct: 215 FGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCATSLESMVDFSVSKLG 274
Query: 129 N-NVEAVSTE-ANKESDKQQY-IIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVY 185
+V AVSTE A K + Q+Y I A GVKKLS++K VVCH YP+AVFYCHK T VY
Sbjct: 275 KYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSDDKSVVCHKQKYPFAVFYCHKAMMTTVY 334
Query: 186 YLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFS 242
+P+EG +G + K A+CH +TS W+P HLAF VLKV+PGTVPVCHFL HV+WFS
Sbjct: 335 AVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVCHFLPETHVVWFS 391
>EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor
Length = 268
Score = 116 bits (290), Expect = 6e-26
Identities = 74/217 (34%), Positives = 119/217 (54%), Gaps = 22/217 (10%)
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFSIK 84
+L Y L FFE DLH G + T + + + PF+ ++ + + + SI
Sbjct: 61 ELKQYSTL---FFEHDLHPGKNFILGNTNSVGSIIR-------PFTKSR-QGVTD--SIW 107
Query: 85 QGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNN-VEAVSTEANKESD 143
+ E ++L+ +C +PT E K CV+SL+SM+D S G+ ++A+S+ D
Sbjct: 108 LANKEKQSLED---FCYSPTAIAEHKHCVSSLKSMIDQVISHFGSTKIKAISSNFAPYQD 164
Query: 144 KQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAIC 203
QY++ + VKK+ +N V+CH L++ VF CH++R T Y + + DGTK K +C
Sbjct: 165 --QYVV-EDVKKVGDNA-VMCHRLNFEKVVFNCHQVRDTTAYVVSLVASDGTKTKALTVC 220
Query: 204 HNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
H+DT +P+ L + L+V PGTVPVCHF+ + W
Sbjct: 221 HHDTRGMNPE-LLYEALEVTPGTVPVCHFIGNKAAAW 256
>EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor
Length = 268
Score = 115 bits (287), Expect = 1e-25
Identities = 80/223 (35%), Positives = 120/223 (52%), Gaps = 34/223 (15%)
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFS-I 83
+L Y L FFE DLH PR+ N I ++N V +I+ F+
Sbjct: 61 ELKQYSTL---FFEHDLH-----------------PRK--NFILGNTNSVGSIIRPFTKS 98
Query: 84 KQGSAESENL----KQAI-GYCETPTIEGEEKSCVTSLESMVDFTTSKLGNN-VEAVSTE 137
+QG +S L KQ+ +C +PT E K CV+SL+SM+D S G+ ++A+S+
Sbjct: 99 RQGVTDSIWLANKEKQSFEDFCYSPTAIAEHKHCVSSLKSMIDQVISHFGSTKIKAISSN 158
Query: 138 ANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKV 197
D QY++ + VKK+ +N V+CH L++ VF CH++R T Y + + DGTK
Sbjct: 159 FAPYQD--QYVV-EDVKKVGDNA-VMCHRLNFEKVVFNCHQVRETTAYVVSLVASDGTKT 214
Query: 198 KTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
K +CH+DT +P+ L + L+V PGTVPVCHF+ + W
Sbjct: 215 KALTVCHHDTRGMNPE-LLYEALEVTPGTVPVCHFIGNKAAAW 256
>SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1)
Length = 268
Score = 111 bits (278), Expect = 1e-24
Identities = 79/223 (35%), Positives = 120/223 (53%), Gaps = 34/223 (15%)
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFS-I 83
+L Y L FFE DLH PR+ N I ++N V +I+ F+
Sbjct: 61 ELKQYSTL---FFEHDLH-----------------PRK--NFILGNTNSVGSIIRPFTKS 98
Query: 84 KQGSAESENL----KQAI-GYCETPTIEGEEKSCVTSLESMVDFTTSKLGNN-VEAVSTE 137
+QG +S L KQ++ +C +PT E K CV+SL+SM+D S G+ ++A+S+
Sbjct: 99 RQGVTDSIWLANKEKQSLEDFCYSPTAIAEHKHCVSSLKSMIDQVISHFGSTKIKAISSN 158
Query: 138 ANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKV 197
D QY++ + VKK+ +N V+CH L++ VF CH++R T Y + + DGTK
Sbjct: 159 FAPYQD--QYVV-EDVKKVGDNA-VMCHRLNFEKVVFNCHQVRDTTAYVVSLVASDGTKT 214
Query: 198 KTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
K +CH+DT +P+ L + L+V GTVPVCHF+ + W
Sbjct: 215 KALTVCHHDTRGMNPE-LLYEALEVTLGTVPVCHFIGNKAAAW 256
>EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor
Length = 268
Score = 111 bits (278), Expect = 1e-24
Identities = 79/223 (35%), Positives = 120/223 (53%), Gaps = 34/223 (15%)
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFS-I 83
+L Y L FFE DLH PR+ N I ++N V +I+ F+
Sbjct: 61 ELKQYSTL---FFEHDLH-----------------PRK--NFILGNTNSVGSIIRPFTKS 98
Query: 84 KQGSAESENL----KQAI-GYCETPTIEGEEKSCVTSLESMVDFTTSKLGNN-VEAVSTE 137
+QG +S L KQ++ +C +PT E K CV+SL+SM+D S G+ ++A+S+
Sbjct: 99 RQGVTDSIWLANKEKQSLEDFCYSPTAIAEHKHCVSSLKSMIDQVISHFGSTKIKAISSN 158
Query: 138 ANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKV 197
D QY++ + VKK+ +N V+CH L++ VF CH++R T Y + + DGTK
Sbjct: 159 FAPYQD--QYVV-EDVKKVGDNA-VMCHRLNFEKVVFNCHQVRDTTAYVVSLVASDGTKT 214
Query: 198 KTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
K +CH+DT +P+ L + L+V GTVPVCHF+ + W
Sbjct: 215 KALTVCHHDTRGMNPE-LLYEALEVTLGTVPVCHFIGNKAAAW 256
>SYH_HELHP (Q7VF68) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 442
Score = 38.1 bits (87), Expect = 0.019
Identities = 21/64 (32%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Query: 38 EKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAI 97
EKD++ ++ + K G + E+ + ++N+VEN+L Y SIKQ ESE+ + I
Sbjct: 180 EKDINVALRIIDKLDKIGKERVSEELQKELGLNNNQVENLLEYTSIKQ-QGESESFFRQI 238
Query: 98 GYCE 101
+ E
Sbjct: 239 SFME 242
>DPOQ_HUMAN (O75417) DNA polymerase theta (EC 2.7.7.7) (DNA
polymerase eta)
Length = 1762
Score = 32.7 bits (73), Expect = 0.81
Identities = 29/113 (25%), Positives = 54/113 (47%), Gaps = 8/113 (7%)
Query: 38 EKDLHNGTKLNMQFTKTGATF-----LPREVAN-SIPFSSNKVENI-LNYFSIKQGSAES 90
EK GT+ N F +GA+F L R + S P + K++++ +N+ S+ + + E
Sbjct: 788 EKSKLTGTRQNHSFIWSGASFDLSPGLQRILDKVSSPLENEKLKSMTINFSSLNRKNTEL 847
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESD 143
++ I ET ++G S ++S ++ + N+ E S KES+
Sbjct: 848 NEEQEVISNLETKQVQGISFSSNNEVKSKIEMLENN-ANHDETSSLLPRKESN 899
>SYFB_ENTFA (Q836J5) Phenylalanyl-tRNA synthetase beta chain (EC
6.1.1.20) (Phenylalanine--tRNA ligase beta chain)
(PheRS)
Length = 807
Score = 31.6 bits (70), Expect = 1.8
Identities = 38/155 (24%), Positives = 63/155 (40%), Gaps = 36/155 (23%)
Query: 60 PREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLES- 118
P+E+A+ + + +VE + + E LK+ + GE K CV S
Sbjct: 19 PQELADKMSVTGIEVEGV---------AVPEEGLKKIV--------VGEVKECVPHPNSD 61
Query: 119 -----MVDFTTSKLGN------NVEA-----VSTEANKESDKQQYIIAKGVKKLSENKIV 162
VD +L NV+A V+ ++ + Q+ I KG + + +
Sbjct: 62 HLSICQVDIGEEELSQIVCGAPNVKAGIKVIVALPGSRIAGNQK--IKKGKMRGEVSNGM 119
Query: 163 VCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKV 197
+C L Y+ K A +YYLP E ++GT V
Sbjct: 120 ICSLEELGYSDNVVPKAYAEGIYYLPQEAVNGTPV 154
>G3P_BACME (P23722) Glyceraldehyde-3-phosphate dehydrogenase (EC
1.2.1.12) (GAPDH)
Length = 334
Score = 31.6 bits (70), Expect = 1.8
Identities = 23/74 (31%), Positives = 37/74 (49%), Gaps = 10/74 (13%)
Query: 88 AESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKL--GNNVEAV------STEAN 139
A +LK +GY E P + G+ + S S +D ++ + GN V+ + S +N
Sbjct: 263 AAEGDLKGILGYSEEPLVSGDYNGNINS--STIDALSTMVMEGNMVKVISWYDNESGYSN 320
Query: 140 KESDKQQYIIAKGV 153
+ D QYI AKG+
Sbjct: 321 RVVDLAQYIAAKGL 334
>PEP1_RABIT (P28712) Pepsin II-1 precursor (EC 3.4.23.1) (Pepsin A)
Length = 387
Score = 31.2 bits (69), Expect = 2.4
Identities = 30/105 (28%), Positives = 42/105 (39%), Gaps = 16/105 (15%)
Query: 104 TIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVV 163
TI GE +C S +++VD TS L A+S K Q I L EN I
Sbjct: 259 TINGETIACADSCQAVVDTGTSLLAGPTSAIS--------KIQSYIGASKNLLGENIISC 310
Query: 164 CHLLSYPYAVFYCHKLR---ATKVYYLP-----MEGIDGTKVKTA 200
+ S P VF + ++ Y L + G DG + T+
Sbjct: 311 SAIDSLPDIVFTINNVQYPLPASAYILKEDDDCLSGFDGMNLDTS 355
>ISPH_FUSNN (Q8RI52) 4-hydroxy-3-methylbut-2-enyl diphosphate
reductase (EC 1.17.1.2)
Length = 827
Score = 31.2 bits (69), Expect = 2.4
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIA 150
E++ Q Y + P GE + + + D+ K+G+ VE + T ++E D Q+YI A
Sbjct: 317 ESMDQNFSYLDVP---GERTAVRVRTDELKDY---KVGDTVEVLITGLSEEDDDQEYITA 370
>ARLY_CLOTE (P59616) Argininosuccinate lyase (EC 4.3.2.1)
(Arginosuccinase) (ASAL)
Length = 438
Score = 31.2 bits (69), Expect = 2.4
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 111 SCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIV 162
SC+ +E M+ K GN + AV ++ Y++ KGV +KI+
Sbjct: 335 SCIKVMEGMLSSLKVKKGNTLRAVKMGFLNATESADYLVLKGVPFRDAHKII 386
>A10D_MOUSE (Q8K2X1) Potential phospholipid-transporting ATPase VD
(EC 3.6.3.1) (ATPVD)
Length = 1416
Score = 31.2 bits (69), Expect = 2.4
Identities = 17/71 (23%), Positives = 36/71 (49%), Gaps = 6/71 (8%)
Query: 116 LESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVV--CHLLSYPYAV 173
++S VDF K+ + ++ + ++ + QY+ + L+ENK+V C + +
Sbjct: 403 IQSDVDFYNEKMDSTIQCRALNITEDLGQIQYLFSDKTGTLTENKMVFRRCSVAGFD--- 459
Query: 174 FYCHKLRATKV 184
YCH+ A ++
Sbjct: 460 -YCHEENAKRL 469
>HS74_CAEEL (P20163) Heat shock 70 kDa protein D precursor
Length = 657
Score = 30.4 bits (67), Expect = 4.0
Identities = 33/126 (26%), Positives = 50/126 (39%), Gaps = 13/126 (10%)
Query: 39 KDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIG 98
+D G K + T P ++ I N + ++ ES N +A
Sbjct: 516 EDKGTGNKNKLTITNDHNRLSPEDIERMI----NDADKFAADDQAQKEKVESRNELEAYA 571
Query: 99 YCETPTIEGEEK-------SCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQ-QYIIA 150
Y I +EK S+ES V+ LG+N +A STE NKE K+ + ++
Sbjct: 572 YQMKTQIADKEKLGGKLTDEDKVSIESAVERAIEWLGSNQDA-STEENKEQKKELESVVQ 630
Query: 151 KGVKKL 156
V KL
Sbjct: 631 PIVSKL 636
>RS1_CHLPN (Q9Z8M3) 30S ribosomal protein S1
Length = 580
Score = 30.0 bits (66), Expect = 5.3
Identities = 18/34 (52%), Positives = 21/34 (60%), Gaps = 4/34 (11%)
Query: 126 KLGNNVEAVSTEANKESDKQQYIIAKGVKKLSEN 159
K GN+VEAV +KES K I GVK+LS N
Sbjct: 438 KKGNSVEAVILSVDKESKK----ITLGVKQLSSN 467
>A10D_HUMAN (Q9P241) Potential phospholipid-transporting ATPase VD
(EC 3.6.3.1) (ATPVD)
Length = 1426
Score = 29.6 bits (65), Expect = 6.9
Identities = 18/71 (25%), Positives = 36/71 (50%), Gaps = 6/71 (8%)
Query: 116 LESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVV--CHLLSYPYAV 173
++S VDF K+ + V+ + ++ + QY+ + L+ENK+V C + +
Sbjct: 403 IQSDVDFYNEKMDSIVQCRALNIAEDLGQIQYLFSDKTGTLTENKMVFRRCSVAGFD--- 459
Query: 174 FYCHKLRATKV 184
YCH+ A ++
Sbjct: 460 -YCHEENARRL 469
>GAC3_HUMAN (Q99928) Gamma-aminobutyric-acid receptor gamma-3
subunit precursor (GABA(A) receptor)
Length = 467
Score = 29.3 bits (64), Expect = 9.0
Identities = 23/83 (27%), Positives = 37/83 (43%), Gaps = 12/83 (14%)
Query: 134 VSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGID 193
+ST A K + Y+ A + + VC L + + Y AT YY
Sbjct: 299 LSTIARKSLPRVSYVTAMDLF------VTVCFLFVFAALMEY-----ATLNYYSSCRKPT 347
Query: 194 GTKVKTAAICHNDTSQWSPKHLA 216
TK KT ++ H D+S+W P+ ++
Sbjct: 348 TTK-KTTSLLHPDSSRWIPERIS 369
>CAO3_MOUSE (Q9EPL9) Acyl-coenzyme A oxidase 3, peroxisomal (EC
1.3.3.6) (Pristanoyl-CoA oxidase)
Length = 700
Score = 29.3 bits (64), Expect = 9.0
Identities = 22/84 (26%), Positives = 42/84 (49%), Gaps = 7/84 (8%)
Query: 73 KVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSC-----VTSLESMVDFTTSKL 127
K+ ++N + S ++ + + + T + G EK + SLE F ++L
Sbjct: 102 KILVLINCLGMYDWSLANKCVLHMLVFGTTVFVSGSEKHFKYLEKIYSLEIFGCFALTEL 161
Query: 128 --GNNVEAVSTEANKESDKQQYII 149
G+N +A+ T A+ ESD Q++I+
Sbjct: 162 SHGSNTKAMRTTAHYESDTQEFIL 185
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.133 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,021,862
Number of Sequences: 164201
Number of extensions: 1071039
Number of successful extensions: 2605
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 2586
Number of HSP's gapped (non-prelim): 19
length of query: 243
length of database: 59,974,054
effective HSP length: 107
effective length of query: 136
effective length of database: 42,404,547
effective search space: 5767018392
effective search space used: 5767018392
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC148532.6


