

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
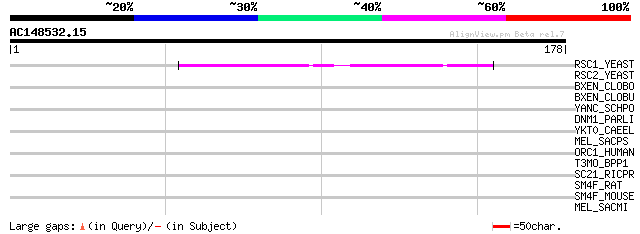
Score E
Sequences producing significant alignments: (bits) Value
RSC1_YEAST (P53236) Chromatin structure remodeling complex prote... 42 6e-04
RSC2_YEAST (Q06488) Chromatin structure remodeling complex prote... 40 0.002
BXEN_CLOBO (P46082) Botulinum neurotoxin type E, nontoxic component 32 0.59
BXEN_CLOBU (Q06366) Botulinum neurotoxin type E, nontoxic component 32 0.77
YANC_SCHPO (Q10077) Hypothetical protein C3H1.12c in chromosome I 32 1.0
DNM1_PARLI (Q27746) DNA (cytosine-5)-methyltransferase PliMCI (E... 32 1.0
YKT0_CAEEL (P34321) Hypothetical protein C07A9.10 in chromosome III 30 2.3
MEL_SACPS (Q03647) Alpha-galactosidase precursor (EC 3.2.1.22) (... 30 3.8
ORC1_HUMAN (Q13415) Origin recognition complex subunit 1 (Replic... 29 5.0
T3MO_BPP1 (P08763) Type III restriction-modification system EcoP... 29 6.6
SC21_RICPR (Q9ZEB4) SCO2-like protein RP031 29 6.6
SM4F_RAT (Q9Z143) Semaphorin 4F precursor (Semaphorin W) (Sema W) 28 8.6
SM4F_MOUSE (Q9Z123) Semaphorin 4F precursor (Semaphorin W) (Sema W) 28 8.6
MEL_SACMI (Q11129) Alpha-galactosidase precursor (EC 3.2.1.22) (... 28 8.6
>RSC1_YEAST (P53236) Chromatin structure remodeling complex protein
RSC1 (Remodel the structure of chromatin complex subunit
RSC1)
Length = 928
Score = 42.4 bits (98), Expect = 6e-04
Identities = 27/101 (26%), Positives = 46/101 (44%), Gaps = 7/101 (6%)
Query: 55 NDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLA 114
ND +P +G+I ++W T + + W+FRP + + + Y NE+ G
Sbjct: 382 NDINKPIVGQIFRLWSTTDGNKWLNACWYFRPEQTVHRVDRL-FYKNEVM-----KTGQY 435
Query: 115 TIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRY 155
HP++ I GKC ++ ++ R S KV +RY
Sbjct: 436 RDHPIQDIKGKCYVIHFTRFQRGDP-STKVNGPQFVCEFRY 475
>RSC2_YEAST (Q06488) Chromatin structure remodeling complex protein
RSC2 (Remodel the structure of chromatin complex subunit
RSC2)
Length = 889
Score = 40.4 bits (93), Expect = 0.002
Identities = 25/98 (25%), Positives = 51/98 (51%), Gaps = 10/98 (10%)
Query: 44 DYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNEL 103
D++L N +ND +P +G+I ++W+TP ++ + W++RP + + + Y NE+
Sbjct: 414 DWALLRN---QNDPQKPIVGQIFRLWKTPDGKQWLNACWYYRPEQTVHRVDRL-FYKNEV 469
Query: 104 FFACGDGDGLATIHPLESIAGKCNIVCISKDSR-NPQL 140
G H + ++ GKC ++ ++ R NP +
Sbjct: 470 M-----KTGQYRDHLVSNLVGKCYVIHFTRYQRGNPDM 502
>BXEN_CLOBO (P46082) Botulinum neurotoxin type E, nontoxic component
Length = 1162
Score = 32.3 bits (72), Expect = 0.59
Identities = 23/93 (24%), Positives = 43/93 (45%), Gaps = 5/93 (5%)
Query: 4 SPPAPSP-ESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDN--VYVKNDFAEP 60
S P P E+ T +++ Q K + + + + G ++ N +Y K ++AE
Sbjct: 102 STAIPFPYENNTEDYR--QTNYLSSKNNEHYYTANLVIFGPGSNIIKNNVIYYKKEYAES 159
Query: 61 RIGKIIKIWETPTLERKIKVQWFFRPIEVSKCL 93
+G +++IW P L K + +E+ KCL
Sbjct: 160 GMGTMLEIWFQPFLTHKYDEFYVDPALELIKCL 192
>BXEN_CLOBU (Q06366) Botulinum neurotoxin type E, nontoxic component
Length = 1162
Score = 32.0 bits (71), Expect = 0.77
Identities = 23/93 (24%), Positives = 43/93 (45%), Gaps = 5/93 (5%)
Query: 4 SPPAPSP-ESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDN--VYVKNDFAEP 60
S P P E+ T +++ Q K + + + + G ++ N +Y K ++AE
Sbjct: 102 STAIPFPYENNTEDYR--QTNYLSSKNNEHYYTANLVIFGPGSNIIKNNVIYYKKEYAEN 159
Query: 61 RIGKIIKIWETPTLERKIKVQWFFRPIEVSKCL 93
+G +++IW P L K + +E+ KCL
Sbjct: 160 GMGTMLEIWFQPFLTHKYDEFYVDPALELIKCL 192
>YANC_SCHPO (Q10077) Hypothetical protein C3H1.12c in chromosome I
Length = 1131
Score = 31.6 bits (70), Expect = 1.0
Identities = 29/126 (23%), Positives = 57/126 (45%), Gaps = 19/126 (15%)
Query: 49 DNVYVKNDF-AEP-RIGKIIKIWET-----PTLERKIKVQWFFRPIEVSKCLTWIKIYFN 101
D V V + F EP +I +II ++ L +++ W+FRP ++ + LT ++ F
Sbjct: 109 DFVLVNSPFPGEPFQIARIISFEKSRPCVSTNLYDSVRLNWYFRPRDIQRHLTDTRLLFA 168
Query: 102 ELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQR 161
+ + I+ + S+ KC + S+ L + + + ++ F R FD
Sbjct: 169 SMH---------SDIYNIGSVQEKCT---VKHRSQIENLDEYKSQAKSYYFDRLFDQNIN 216
Query: 162 KIVEEV 167
K+ + V
Sbjct: 217 KVFDVV 222
>DNM1_PARLI (Q27746) DNA (cytosine-5)-methyltransferase PliMCI (EC
2.1.1.37) (Dnmt1) (DNA methyltransferase PliMCI) (DNA
MTase PliMCI) (MCMT) (M.PliMCI)
Length = 1612
Score = 31.6 bits (70), Expect = 1.0
Identities = 34/126 (26%), Positives = 57/126 (44%), Gaps = 13/126 (10%)
Query: 52 YVKN---DFAEP-RIGKIIKIWETPTLER-KIKVQWFFRPIEVSKCLTWI-KIYFNELFF 105
YVK + EP RIGKII I+ T + +++V +RP + K T + N L++
Sbjct: 974 YVKGSNLECPEPFRIGKIISIYTTKSNSTVRLRVNKMYRPEDTHKGRTAAYQADLNVLYW 1033
Query: 106 ACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVE 165
+ + + LE + GKC++VC + N + ++R +RK E
Sbjct: 1034 SEEEA-----VTELEVVQGKCSVVC--AEDLNVSTDEYSAGGPHKFYFREAYDSERKCFE 1086
Query: 166 EVDDKS 171
+ KS
Sbjct: 1087 DPPSKS 1092
>YKT0_CAEEL (P34321) Hypothetical protein C07A9.10 in chromosome III
Length = 254
Score = 30.4 bits (67), Expect = 2.3
Identities = 15/58 (25%), Positives = 27/58 (45%), Gaps = 3/58 (5%)
Query: 92 CLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAA 149
C T +K+ +L C + +HP K ++VC ++ +NP + DK + A
Sbjct: 175 CTTALKVLMKDLAAECKRNTEVKVLHPTRF---KSHLVCSAQGCKNPTIPDKPYFKVA 229
>MEL_SACPS (Q03647) Alpha-galactosidase precursor (EC 3.2.1.22)
(Melibiase) (Alpha-D-galactoside galactohydrolase)
(MELx)
Length = 471
Score = 29.6 bits (65), Expect = 3.8
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 3/35 (8%)
Query: 26 GGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEP 60
G ++ D +F+ S GVDY +DN Y K F P
Sbjct: 129 GHEEEDAEFFAS---NGVDYLKYDNCYNKGQFGAP 160
>ORC1_HUMAN (Q13415) Origin recognition complex subunit 1
(Replication control protein 1)
Length = 861
Score = 29.3 bits (64), Expect = 5.0
Identities = 13/41 (31%), Positives = 24/41 (57%), Gaps = 3/41 (7%)
Query: 55 NDFAEPRIGKIIKIWET---PTLERKIKVQWFFRPIEVSKC 92
+D P + K+++++E P +++ +VQWF R EV C
Sbjct: 58 DDDENPYVAKLLELFEDDSDPPPKKRARVQWFVRFCEVPAC 98
>T3MO_BPP1 (P08763) Type III restriction-modification system EcoPI
enzyme mod (EC 2.1.1.72) (EcoPI methyltransferase)
(M.EcoPI)
Length = 646
Score = 28.9 bits (63), Expect = 6.6
Identities = 45/182 (24%), Positives = 74/182 (39%), Gaps = 36/182 (19%)
Query: 9 SPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKI--- 65
SP E L + WG+ KK + +FY L +D N+Y K P +G++
Sbjct: 333 SPTGEELSWSWGK------KKINDEFY---NLIVIDIKDGKNIYKKQ---RPALGELPTK 380
Query: 66 --IKIWETP---------TLERKIKVQWFFRPIEVSKCLTWIKIYFNE----LFFACGDG 110
IW P L+ + + F P V +KI + L F G G
Sbjct: 381 KPKSIWYKPEYSTSTATTELKNLLGAKLFEGPKPVPLITDLVKIGTKKDSLVLDFFAGSG 440
Query: 111 DGLATIHPL-ESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDD 169
+ L E +G N +CI KD ++ +K + + + F++ +++I +EV
Sbjct: 441 TTAEAVAYLNEKDSGCRNFICIQKD----EVINKTKNAYSLGYRSIFEITKKRI-QEVFK 495
Query: 170 KS 171
KS
Sbjct: 496 KS 497
>SC21_RICPR (Q9ZEB4) SCO2-like protein RP031
Length = 191
Score = 28.9 bits (63), Expect = 6.6
Identities = 19/58 (32%), Positives = 30/58 (50%), Gaps = 6/58 (10%)
Query: 11 ESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKI 68
+S ++F +++GK ++Q Y S G YSL+DN +K R+ IIKI
Sbjct: 46 DSLKIKFTLIEQQGKKFDSTNLQGYLSLIYFGTTYSLYDNQALK------RVEDIIKI 97
>SM4F_RAT (Q9Z143) Semaphorin 4F precursor (Semaphorin W) (Sema W)
Length = 776
Score = 28.5 bits (62), Expect = 8.6
Identities = 14/25 (56%), Positives = 15/25 (60%), Gaps = 1/25 (4%)
Query: 2 SNSPPAPSPESETLEFKWGQKRGKG 26
S PP+PSPE E L G KRG G
Sbjct: 719 SQDPPSPSPEDERLPLALG-KRGSG 742
>SM4F_MOUSE (Q9Z123) Semaphorin 4F precursor (Semaphorin W) (Sema W)
Length = 777
Score = 28.5 bits (62), Expect = 8.6
Identities = 14/25 (56%), Positives = 15/25 (60%), Gaps = 1/25 (4%)
Query: 2 SNSPPAPSPESETLEFKWGQKRGKG 26
S PP+PSPE E L G KRG G
Sbjct: 720 SQDPPSPSPEDERLPLALG-KRGSG 743
>MEL_SACMI (Q11129) Alpha-galactosidase precursor (EC 3.2.1.22)
(Melibiase) (Alpha-D-galactoside galactohydrolase)
(MELj)
Length = 471
Score = 28.5 bits (62), Expect = 8.6
Identities = 14/35 (40%), Positives = 18/35 (51%), Gaps = 3/35 (8%)
Query: 26 GGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEP 60
G ++ D +F F GVDY +DN Y K F P
Sbjct: 129 GHEQEDAEF---FARNGVDYLKYDNCYNKGKFGTP 160
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,502,588
Number of Sequences: 164201
Number of extensions: 984678
Number of successful extensions: 1620
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1613
Number of HSP's gapped (non-prelim): 15
length of query: 178
length of database: 59,974,054
effective HSP length: 103
effective length of query: 75
effective length of database: 43,061,351
effective search space: 3229601325
effective search space used: 3229601325
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148532.15


