

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.13 + phase: 0
(303 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
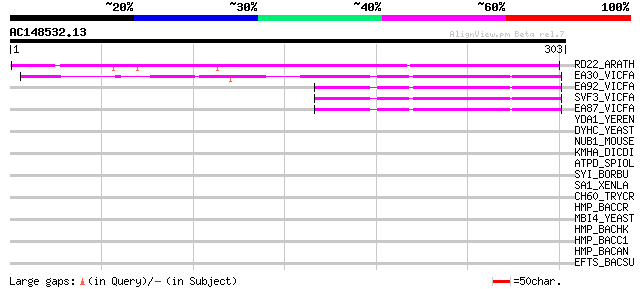
Score E
Sequences producing significant alignments: (bits) Value
RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor 308 1e-83
EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor 107 5e-23
EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor 103 4e-22
SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1) 100 6e-21
EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor 100 6e-21
YDA1_YEREN (P31489) Adhesin yadA precursor 35 0.18
DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC) 33 0.87
NUB1_MOUSE (P54729) NEDD8 ultimate buster-1 (BS4 protein) 31 3.3
KMHA_DICDI (P42527) Myosin heavy chain kinase A (EC 2.7.1.129) (... 31 3.3
ATPD_SPIOL (P11402) ATP synthase delta chain, chloroplast precur... 31 3.3
SYI_BORBU (O51773) Isoleucyl-tRNA synthetase (EC 6.1.1.5) (Isole... 31 4.3
SA1_XENLA (Q9DGN1) Cohesin subunit SA-1 (XSA-1) (Stromal antigen... 30 5.6
CH60_TRYCR (Q95046) Chaperonin HSP60, mitochondrial precursor (P... 30 5.6
HMP_BACCR (Q81FW4) Flavohemoprotein (Hemoglobin-like protein) (F... 30 7.4
MBI4_YEAST (P03879) Intron-encoded RNA maturase bI4 precursor (R... 30 9.6
HMP_BACHK (Q6HLA6) Flavohemoprotein (Hemoglobin-like protein) (F... 30 9.6
HMP_BACC1 (Q73B49) Flavohemoprotein (Hemoglobin-like protein) (F... 30 9.6
HMP_BACAN (Q81T23) Flavohemoprotein (Hemoglobin-like protein) (F... 30 9.6
EFTS_BACSU (P80700) Elongation factor Ts (EF-Ts) 30 9.6
>RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor
Length = 392
Score = 308 bits (788), Expect = 1e-83
Identities = 171/370 (46%), Positives = 220/370 (59%), Gaps = 74/370 (20%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYY------ 55
L P+ YW + LPN+P+P ++ NLL +++EK T V VGKGGV+V KG
Sbjct: 23 LTPERYWSTALPNTPIPNSLHNLL--TFDFTDEKSTNVQVGKGGVNVNTHKGKTGSGTAV 80
Query: 56 ---------------EGGGTDVNVGVGR-------------------------------- 68
GGGT V+VG G+
Sbjct: 81 NVGKGGVRVDTGKGKPGGGTHVSVGSGKGHGGGVAVHTGKPGKRTDVGVGKGGVTVHTRH 140
Query: 69 -------------SPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTS--N 113
+PF+YNYAA ETQLHD PN ALFFLEKDL G ++N++F+
Sbjct: 141 KGRPIYVGVKPGANPFVYNYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGG 200
Query: 114 AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVT 173
FLPR A ++PF S K + + +++ GS+ +++K TI ECE + + GEEK C T
Sbjct: 201 KTAFLPRGEAETVPFGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCAT 260
Query: 174 SLESMVDFTTSKLGK-NVEAVSTEV-NKESNLQQYTIAS-GVKKLGEKNKAVVCHKENYP 230
SLESMVDF+ SKLGK +V AVSTEV K + +Q+Y IA+ GVKKL + +K+VVCHK+ YP
Sbjct: 261 SLESMVDFSVSKLGKYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSD-DKSVVCHKQKYP 319
Query: 231 YAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVC 290
+AVFYCHK T Y+VPLEG +G R KA+AVCH +TS WNP HLAF+VLKV+PGTVPVC
Sbjct: 320 FAVFYCHKAMMTTVYAVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVC 379
Query: 291 HLLPEDHVVW 300
H LPE HVVW
Sbjct: 380 HFLPETHVVW 389
>EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor
Length = 268
Score = 107 bits (266), Expect = 5e-23
Identities = 83/300 (27%), Positives = 125/300 (41%), Gaps = 75/300 (25%)
Query: 7 YWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTDVNVGV 66
YW+S+ PN+P+PK ++LL P+ G T+
Sbjct: 28 YWQSIWPNTPLPKTFSDLLIPS-----------------------------GKTN----- 53
Query: 67 GRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLP----RQV 122
+ + + + F E DLH G N T S + P RQ
Sbjct: 54 ----------SLPIKSEELKQYSTLFFEHDLHPGK--NFILGNTNSVGSIIRPFTKSRQG 101
Query: 123 ANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFT 182
+ +NK + + C E K CV+SL+SM+D
Sbjct: 102 VTDSIWLANKEKQSLEDF------------------CYSPTAIAEHKHCVSSLKSMIDQV 143
Query: 183 TSKLGKN-VEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT 241
S G ++A+S+ + Q + VKK+G+ AV+CH+ N+ VF CH+
Sbjct: 144 ISHFGSTKIKAISSNF---APYQDQYVVEDVKKVGDN--AVMCHRLNFEKVVFNCHQVRD 198
Query: 242 TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
T AY V L +DG++ KA+ VCH DT NP+ L ++ L+V PGTVPVCH + W+
Sbjct: 199 TTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTPGTVPVCHFIGNKAAAWV 257
>EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor
Length = 268
Score = 103 bits (258), Expect = 4e-22
Identities = 55/136 (40%), Positives = 80/136 (58%), Gaps = 7/136 (5%)
Query: 167 EEKVCVTSLESMVDFTTSKLGKN-VEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCH 225
E K CV+SL+SM+D S G ++A+S+ + Q + VKK+G+ AV+CH
Sbjct: 128 EHKHCVSSLKSMIDQVISHFGSTKIKAISSNF---APYQDQYVVEDVKKVGDN--AVMCH 182
Query: 226 KENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPG 285
+ N+ VF CH+ T AY V L +DG++ KA+ VCH DT NP+ L ++ L+V PG
Sbjct: 183 RLNFEKVVFNCHQVRETTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTPG 241
Query: 286 TVPVCHLLPEDHVVWI 301
TVPVCH + W+
Sbjct: 242 TVPVCHFIGNKAAAWV 257
Score = 34.7 bits (78), Expect = 0.30
Identities = 12/23 (52%), Positives = 19/23 (82%)
Query: 7 YWKSMLPNSPMPKAITNLLHPAG 29
YW+S+ PN+P+PK ++LL P+G
Sbjct: 28 YWQSIWPNTPLPKTFSDLLIPSG 50
>SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1)
Length = 268
Score = 100 bits (248), Expect = 6e-21
Identities = 54/136 (39%), Positives = 79/136 (57%), Gaps = 7/136 (5%)
Query: 167 EEKVCVTSLESMVDFTTSKLGKN-VEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCH 225
E K CV+SL+SM+D S G ++A+S+ + Q + VKK+G+ AV+CH
Sbjct: 128 EHKHCVSSLKSMIDQVISHFGSTKIKAISSNF---APYQDQYVVEDVKKVGDN--AVMCH 182
Query: 226 KENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPG 285
+ N+ VF CH+ T AY V L +DG++ KA+ VCH DT NP+ L ++ L+V G
Sbjct: 183 RLNFEKVVFNCHQVRDTTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTLG 241
Query: 286 TVPVCHLLPEDHVVWI 301
TVPVCH + W+
Sbjct: 242 TVPVCHFIGNKAAAWV 257
Score = 32.3 bits (72), Expect = 1.5
Identities = 11/23 (47%), Positives = 18/23 (77%)
Query: 7 YWKSMLPNSPMPKAITNLLHPAG 29
YW+S+ PN+P+PK ++L P+G
Sbjct: 28 YWQSIWPNTPLPKTFSDLSIPSG 50
>EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor
Length = 268
Score = 100 bits (248), Expect = 6e-21
Identities = 54/136 (39%), Positives = 79/136 (57%), Gaps = 7/136 (5%)
Query: 167 EEKVCVTSLESMVDFTTSKLGKN-VEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCH 225
E K CV+SL+SM+D S G ++A+S+ + Q + VKK+G+ AV+CH
Sbjct: 128 EHKHCVSSLKSMIDQVISHFGSTKIKAISSNF---APYQDQYVVEDVKKVGDN--AVMCH 182
Query: 226 KENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPG 285
+ N+ VF CH+ T AY V L +DG++ KA+ VCH DT NP+ L ++ L+V G
Sbjct: 183 RLNFEKVVFNCHQVRDTTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTLG 241
Query: 286 TVPVCHLLPEDHVVWI 301
TVPVCH + W+
Sbjct: 242 TVPVCHFIGNKAAAWV 257
Score = 35.0 bits (79), Expect = 0.23
Identities = 12/27 (44%), Positives = 21/27 (77%)
Query: 3 PPQLYWKSMLPNSPMPKAITNLLHPAG 29
P + YW+S+ PN+P+PK +++L P+G
Sbjct: 24 PREDYWQSIWPNTPLPKTFSDMLIPSG 50
>YDA1_YEREN (P31489) Adhesin yadA precursor
Length = 455
Score = 35.4 bits (80), Expect = 0.18
Identities = 45/215 (20%), Positives = 78/215 (35%), Gaps = 29/215 (13%)
Query: 16 PMPKAITN--LLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTDVNVGVGRSPFI- 72
P+ KA+ + + + A +++ G + D GV G+ +V +G S +
Sbjct: 105 PLSKALGDSAVTYGAASTAQKDGVAIGARASTSDTGVAVGFNSKADAKNSVAIGHSSHVA 164
Query: 73 ----YNYAASETQLHDKPN---VALFFLEKDLHH---GTK---------LNLQFSKTTSN 113
Y+ A + D+ N + L + L H GTK L + KT N
Sbjct: 165 ANHGYSIAIGDRSKTDRENSVSIGHESLNRQLTHLAAGTKDTDAVNVAQLKKEIEKTQEN 224
Query: 114 AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQ-------GIKG 166
+AN+ ++ NK ++ N SK + ++N E Q
Sbjct: 225 TNKRSAELLANANAYADNKSSSVLGIANNYTDSKSAETLENARKEAFAQSKDVLNMAKAH 284
Query: 167 EEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKES 201
V T+LE+ + S +E NK+S
Sbjct: 285 SNSVARTTLETAEEHANSVARTTLETAEEHANKKS 319
>DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC)
Length = 4092
Score = 33.1 bits (74), Expect = 0.87
Identities = 34/162 (20%), Positives = 69/162 (41%), Gaps = 24/162 (14%)
Query: 53 GYYEGGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTS 112
GY D+ + + FI+N +T LH KP + + E+ L + F+ T
Sbjct: 3145 GYQFSNWRDIQQFIRKDDFIHNIVHYDTTLHMKPQIRKYMEEEFLS-----DPNFTYETI 3199
Query: 113 NAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCV 172
N A+ +++ ++N K + V ++ + E + +K + +
Sbjct: 3200 NRAS----------KACGPLYQWVNAQINFSKVLENVDPLRQEMKRIEFESLKTKANLLA 3249
Query: 173 T-----SLESMVDFTTSK---LGKNVEAVSTEV-NKESNLQQ 205
LE+ ++ + K L ++VEA+ TE+ N ++NL +
Sbjct: 3250 AEEMTQDLEASIEVSKRKYSLLIRDVEAIKTEMSNVQANLDR 3291
>NUB1_MOUSE (P54729) NEDD8 ultimate buster-1 (BS4 protein)
Length = 614
Score = 31.2 bits (69), Expect = 3.3
Identities = 28/111 (25%), Positives = 45/111 (40%), Gaps = 14/111 (12%)
Query: 121 QVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVD 180
Q+A + F N ++ +INK ++ G EEQG+ K V L+ +
Sbjct: 116 QIAETFGFQENYIKIVINKKQLQLG-----------KSLEEQGVTHNVKAMVLELKQSEE 164
Query: 181 FTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPY 231
L E + KE +Q+ G++ L E+ + VV E PY
Sbjct: 165 DVRKNLQLEEEEQNEAELKERRIQR--TKRGLEILAERAEMVV-DPETMPY 212
>KMHA_DICDI (P42527) Myosin heavy chain kinase A (EC 2.7.1.129)
(MHCK A)
Length = 1146
Score = 31.2 bits (69), Expect = 3.3
Identities = 37/173 (21%), Positives = 71/173 (40%), Gaps = 18/173 (10%)
Query: 50 VRKGYYE---GGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQ 106
+R+ Y+ G GTD N +G + ++ + + K N A+ L +GTK L+
Sbjct: 591 LREAYHTVSLGVGTDENYPLGTTTKLFPPIEMISPI-SKNNEAM----TQLKNGTKFVLK 645
Query: 107 FSKTTSNAATFLPRQVANSIPFSSNKMEYII----NKLNIKKGSKGVQIVKNTISECEEQ 162
K + +Q + + F KM+ + NK N KK K ++ + + + E ++
Sbjct: 646 LYKKEAE------QQASRELYFEDVKMQMVCRDWGNKFNQKKPPKKIEFLMSWVVELIDR 699
Query: 163 GIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKL 215
+ + S+E ++ K N AV T + +T K++
Sbjct: 700 SPSSNGQPILCSIEPLLVGEFKKNNSNYGAVLTNRSTPQAFSHFTYELSNKQM 752
>ATPD_SPIOL (P11402) ATP synthase delta chain, chloroplast precursor
(EC 3.6.3.14)
Length = 257
Score = 31.2 bits (69), Expect = 3.3
Identities = 21/81 (25%), Positives = 40/81 (48%), Gaps = 2/81 (2%)
Query: 120 RQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEE--QGIKGEEKVCVTSLES 177
R V + I +S + N +NI S+ + +VK ++E E+ I G E VTS+
Sbjct: 126 RSVLDEIITTSGLQPHTANFINILIDSERINLVKEILNEFEDVFNKITGTEVAVVTSVVK 185
Query: 178 MVDFTTSKLGKNVEAVSTEVN 198
+ + +++ K V+ ++ N
Sbjct: 186 LENDHLAQIAKGVQKITGAKN 206
>SYI_BORBU (O51773) Isoleucyl-tRNA synthetase (EC 6.1.1.5)
(Isoleucine--tRNA ligase) (IleRS)
Length = 1042
Score = 30.8 bits (68), Expect = 4.3
Identities = 35/134 (26%), Positives = 64/134 (47%), Gaps = 11/134 (8%)
Query: 82 LHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLN 141
L+D P F+ K + K+NL T+ + + +P S+ YI+ K N
Sbjct: 783 LNDYPKANENFINKTIEE--KINLARKITSMARSLRSLHNIKIRMPISTI---YIVTK-N 836
Query: 142 IKKGSKGVQIVKNTISE--CEEQGIKGEEKVCVTSLESMVDFTT--SKLGKNVEAVSTEV 197
+ + +++ + + E +E IK E+ +T ++ +F KLGK+++AVSTE+
Sbjct: 837 QNEQNMLMEMQEIILDEINAKEMKIKANEEELIT-YKAKANFKELGKKLGKDMKAVSTEI 895
Query: 198 NKESNLQQYTIASG 211
+K N I +G
Sbjct: 896 SKLKNEDIIKIING 909
>SA1_XENLA (Q9DGN1) Cohesin subunit SA-1 (XSA-1) (Stromal antigen 1
homolog) (SCC3 homolog 1)
Length = 1265
Score = 30.4 bits (67), Expect = 5.6
Identities = 17/86 (19%), Positives = 39/86 (44%), Gaps = 1/86 (1%)
Query: 151 IVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNL-QQYTIA 209
I+K T+S+ + K + SL+ + + + G N++ S V+ L +++ +
Sbjct: 910 IIKETLSKTRQMDKIQCAKTLILSLQQLFNELVQEQGPNLDRTSAHVSGIKELARRFALT 969
Query: 210 SGVKKLGEKNKAVVCHKENYPYAVFY 235
G+ ++ + HK+ +A Y
Sbjct: 970 FGLDQIKTREAVATLHKDGIEFAFKY 995
>CH60_TRYCR (Q95046) Chaperonin HSP60, mitochondrial precursor
(Protein Cpn60) (groEL protein) (Heat shock protein 60)
Length = 562
Score = 30.4 bits (67), Expect = 5.6
Identities = 26/99 (26%), Positives = 47/99 (47%), Gaps = 7/99 (7%)
Query: 198 NKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT-TKAYSVPLEGADGS- 255
NK + +Q I +G + +GE+ + EN+ A+ K T TK +V L G S
Sbjct: 291 NKTAMMQDIAIFAGARLVGEEGSGLELDAENFDPAILGTVKKATITKDDTVLLNGGGESS 350
Query: 256 ----RVKAI-AVCHTDTSEWNPKHLAFQVLKVQPGTVPV 289
RV+ + + +TS++N + L ++ K+ G +
Sbjct: 351 MVKERVELLRGLIDGETSDYNREKLQERLAKLSGGVAVI 389
>HMP_BACCR (Q81FW4) Flavohemoprotein (Hemoglobin-like protein)
(Flavohemoglobin) (Nitric oxide dioxygenase) (EC
1.14.12.17) (NO oxygenase) (NOD)
Length = 402
Score = 30.0 bits (66), Expect = 7.4
Identities = 23/95 (24%), Positives = 49/95 (51%), Gaps = 12/95 (12%)
Query: 146 SKGVQIVKNTISECEEQGIKGEEKVCV------TSLESMVDFTTSKLGKNVEAVSTEVNK 199
+K ++IVK+T+ +E+G++ + L ++ + T K G+ +A++ V
Sbjct: 4 AKTIEIVKSTVPLLQEKGVEITTRFYQILFSEHPELLNIFNHTNQKKGRQQQALANAVYA 63
Query: 200 ES----NLQQYTIASGVKKLGEKNKAVVCHKENYP 230
+ NL+ I VK++G K++++ E+YP
Sbjct: 64 AATYIDNLE--AIIPVVKQIGHKHRSLGIKAEHYP 96
>MBI4_YEAST (P03879) Intron-encoded RNA maturase bI4 precursor (RNA
maturase SCbI4) [Contains: Truncated, nonfunctional
cytochrome b; RNA maturase bI4 (EC 3.1.-.-)]
Length = 638
Score = 29.6 bits (65), Expect = 9.6
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 1/59 (1%)
Query: 86 PNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKK 144
PN ++ ++D T +N + TSN L + + + NK Y++N N+KK
Sbjct: 327 PNRVKYYYKEDNQQVTNMNSSNTHLTSNKKNLLV-DTSETTRTTKNKFNYLLNIFNMKK 384
>HMP_BACHK (Q6HLA6) Flavohemoprotein (Hemoglobin-like protein)
(Flavohemoglobin) (Nitric oxide dioxygenase) (EC
1.14.12.17) (NO oxygenase) (NOD)
Length = 402
Score = 29.6 bits (65), Expect = 9.6
Identities = 23/94 (24%), Positives = 48/94 (50%), Gaps = 12/94 (12%)
Query: 147 KGVQIVKNTISECEEQGIKGEEKVCV------TSLESMVDFTTSKLGKNVEAVSTEVNKE 200
K ++IVK+T+ +E+G++ + L ++ + T K G+ +A++ V
Sbjct: 5 KTIEIVKSTVPLLQEKGVEITTRFYEILFSEHPELLNIFNHTNQKKGRQQQALANAVYAA 64
Query: 201 S----NLQQYTIASGVKKLGEKNKAVVCHKENYP 230
+ NL+ I VK++G K++++ E+YP
Sbjct: 65 ATYIDNLE--AIIPVVKQIGHKHRSLGIKAEHYP 96
>HMP_BACC1 (Q73B49) Flavohemoprotein (Hemoglobin-like protein)
(Flavohemoglobin) (Nitric oxide dioxygenase) (EC
1.14.12.17) (NO oxygenase) (NOD)
Length = 402
Score = 29.6 bits (65), Expect = 9.6
Identities = 23/94 (24%), Positives = 48/94 (50%), Gaps = 12/94 (12%)
Query: 147 KGVQIVKNTISECEEQGIKGEEKVCV------TSLESMVDFTTSKLGKNVEAVSTEVNKE 200
K ++IVK+T+ +E+G++ + L ++ + T K G+ +A++ V
Sbjct: 5 KTIEIVKSTVPLLQEKGVEITTRFYEILFSEHPELLNIFNHTNQKKGRQQQALANAVYAA 64
Query: 201 S----NLQQYTIASGVKKLGEKNKAVVCHKENYP 230
+ NL+ I VK++G K++++ E+YP
Sbjct: 65 ATYIDNLE--VIIPVVKQIGHKHRSLGIKAEHYP 96
>HMP_BACAN (Q81T23) Flavohemoprotein (Hemoglobin-like protein)
(Flavohemoglobin) (Nitric oxide dioxygenase) (EC
1.14.12.17) (NO oxygenase) (NOD)
Length = 402
Score = 29.6 bits (65), Expect = 9.6
Identities = 23/94 (24%), Positives = 48/94 (50%), Gaps = 12/94 (12%)
Query: 147 KGVQIVKNTISECEEQGIKGEEKVCV------TSLESMVDFTTSKLGKNVEAVSTEVNKE 200
K ++IVK+T+ +E+G++ + L ++ + T K G+ +A++ V
Sbjct: 5 KTIEIVKSTVPLLQEKGVEITTRFYEILFSEHPELLNIFNHTNQKKGRQQQALANAVYAA 64
Query: 201 S----NLQQYTIASGVKKLGEKNKAVVCHKENYP 230
+ NL+ I VK++G K++++ E+YP
Sbjct: 65 ATYIDNLE--AIIPVVKQIGHKHRSLGIKAEHYP 96
>EFTS_BACSU (P80700) Elongation factor Ts (EF-Ts)
Length = 292
Score = 29.6 bits (65), Expect = 9.6
Identities = 35/146 (23%), Positives = 60/146 (40%), Gaps = 29/146 (19%)
Query: 145 GSKGVQIVKNTISE--CEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVE-AVSTEVNKES 201
G+KGV + N+ ++ + +G K L ++ D + +VE A+ ++ S
Sbjct: 66 GNKGVILEVNSETDFVAKNEGFK-------ELLNTLADHLLANTPADVEEAMGQKMENGS 118
Query: 202 NLQQYTIASGVKKLGEK---NKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVK 258
+++Y I S V K+GEK + V K++ Y H G R+
Sbjct: 119 TVEEY-ITSAVAKIGEKITLRRFTVLTKDDSSAFGAYLHM---------------GGRIG 162
Query: 259 AIAVCHTDTSEWNPKHLAFQVLKVQP 284
+ V + T E K +A V V P
Sbjct: 163 VLTVLNGTTDEETAKDIAMHVAAVNP 188
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,961,678
Number of Sequences: 164201
Number of extensions: 1646992
Number of successful extensions: 3163
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 3134
Number of HSP's gapped (non-prelim): 26
length of query: 303
length of database: 59,974,054
effective HSP length: 110
effective length of query: 193
effective length of database: 41,911,944
effective search space: 8089005192
effective search space used: 8089005192
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC148532.13


