

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
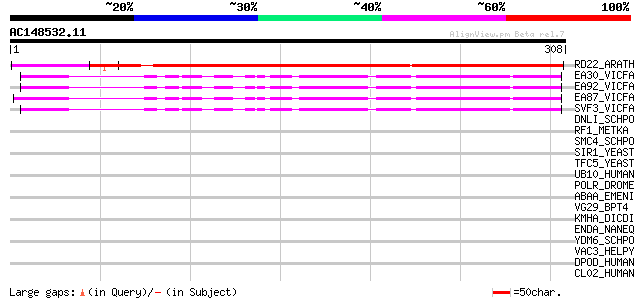
Query= AC148532.11 + phase: 0
(308 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor 265 8e-71
EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor 114 2e-25
EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor 110 3e-24
EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor 108 2e-23
SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1) 105 1e-22
DNLI_SCHPO (P12000) DNA ligase (EC 6.5.1.1) (Polydeoxyribonucleo... 35 0.18
RF1_METKA (Q8TXB5) Peptide chain release factor subunit 1 (Trans... 35 0.31
SMC4_SCHPO (P41004) Structural maintenance of chromosome 4 (Chro... 33 1.2
SIR1_YEAST (P21691) Regulatory protein SIR1 (Silent information ... 32 2.6
TFC5_YEAST (P46678) Transcription factor TFIIIB B" component (TF... 31 3.4
UB10_HUMAN (Q14694) Ubiquitin carboxyl-terminal hydrolase 10 (EC... 31 4.4
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 31 4.4
ABAA_EMENI (P20945) Regulatory protein abaA 31 4.4
VG29_BPT4 (P13337) Tail-tube assembly protein Gp29 (Tail length ... 30 5.8
KMHA_DICDI (P42527) Myosin heavy chain kinase A (EC 2.7.1.129) (... 30 5.8
ENDA_NANEQ (Q74MP4) tRNA-splicing endonuclease (EC 3.1.27.9) (tR... 30 5.8
YDM6_SCHPO (P87137) Hypothetical protein C57A7.06 in chromosome I 30 9.9
VAC3_HELPY (Q48253) Vacuolating cytotoxin precursor 30 9.9
DPOD_HUMAN (P28340) DNA polymerase delta catalytic subunit (EC 2... 30 9.9
CL02_HUMAN (Q8NHQ8) Protein C12orf2 (Carcinoma associated protei... 30 9.9
>RD22_ARATH (Q08298) Dehydration-responsive protein RD22 precursor
Length = 392
Score = 265 bits (678), Expect = 8e-71
Identities = 140/273 (51%), Positives = 180/273 (65%), Gaps = 17/273 (6%)
Query: 45 DVGVRKG-------YEGGGTYLNDEK-IIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLK 96
DVGV KG ++G Y+ + P +Y Y + QL D N LFFL+
Sbjct: 126 DVGVGKGGVTVHTRHKGRPIYVGVKPGANPFVYNYAA------KETQLHDDPNAALFFLE 179
Query: 97 KDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIV 156
KDL G ++N++F T FLPR A ++PF S K L +FS++ GS+EAE++
Sbjct: 180 KDLVRGKEMNVRFNAEDGYGGKTAFLPRGEAETVPFGSEKFSETLKRFSVEAGSEEAEMM 239
Query: 157 KRTISECEANGIKGEEKLCVTSLESMVDFTISKLGN-NVEAVSTEVDKNSNGLQQYVIAK 215
K+TI ECEA + GEEK C TSLESMVDF++SKLG +V AVSTEV K + +Q+Y IA
Sbjct: 240 KKTIEECEARKVSGEEKYCATSLESMVDFSVSKLGKYHVRAVSTEVAKKNAPMQKYKIAA 299
Query: 216 -GVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSE 274
GVKKL + +K++VCHK+ YP+AVFYCHK T VY+VPLEG +G K +AVCH +TS
Sbjct: 300 AGVKKLSD-DKSVVCHKQKYPFAVFYCHKAMMTTVYAVPLEGENGMRAKAVAVCHKNTSA 358
Query: 275 WNPKHLAFYVLKVQPGTVPICHILPQDHVVWVS 307
WNP HLAF VLKV+PGTVP+CH LP+ HVVW S
Sbjct: 359 WNPNHLAFKVLKVKPGTVPVCHFLPETHVVWFS 391
Score = 44.7 bits (104), Expect = 3e-04
Identities = 22/59 (37%), Positives = 30/59 (50%)
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLN 60
L P+ YW + LPN+P+P + NLL K GG++V KG G GT +N
Sbjct: 23 LTPERYWSTALPNTPIPNSLHNLLTFDFTDEKSTNVQVGKGGVNVNTHKGKTGSGTAVN 81
>EA30_VICFA (P21745) Embryonic abundant protein VF30.1 precursor
Length = 268
Score = 114 bits (286), Expect = 2e-25
Identities = 89/301 (29%), Positives = 136/301 (44%), Gaps = 72/301 (23%)
Query: 7 YWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEKIIP 66
YWQS+ PN+P+PK F++LL P+G +
Sbjct: 28 YWQSIWPNTPLPKTFSDLLIPSGKTNS--------------------------------- 54
Query: 67 LIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREV 126
+P+ ++ KQ TLFF + DLH G L N N + R
Sbjct: 55 --------LPIKSEEL----KQYSTLFF-EHDLHPGKNFIL------GNTNSVGSIIR-- 93
Query: 127 ANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFT 186
PF+ ++ + + + SI +KE + ++ C + E K CV+SL+SM+D
Sbjct: 94 ----PFTKSR-QGVTD--SIWLANKEKQSLE---DFCYSPTAIAEHKHCVSSLKSMIDQV 143
Query: 187 ISKLGNN-VEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD 245
IS G+ ++A+S+ N Q + + VKK+G+ ++CH+ N+ VF CH+
Sbjct: 144 ISHFGSTKIKAISS----NFAPYQDQYVVEDVKKVGDN--AVMCHRLNFEKVVFNCHQVR 197
Query: 246 STEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
T Y V L DG K + VCH DT NP+ L + L+V PGTVP+CH + W
Sbjct: 198 DTTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTPGTVPVCHFIGNKAAAW 256
Query: 306 V 306
V
Sbjct: 257 V 257
>EA92_VICFA (P21747) Embryonic abundant protein USP92 precursor
Length = 268
Score = 110 bits (276), Expect = 3e-24
Identities = 88/301 (29%), Positives = 134/301 (44%), Gaps = 72/301 (23%)
Query: 7 YWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEKIIP 66
YWQS+ PN+P+PK F++LL P+G +
Sbjct: 28 YWQSIWPNTPLPKTFSDLLIPSGKTNS--------------------------------- 54
Query: 67 LIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREV 126
+P+ ++ KQ TLFF + DLH L N N + R
Sbjct: 55 --------LPIKSEEL----KQYSTLFF-EHDLHPRKNFIL------GNTNSVGSIIR-- 93
Query: 127 ANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFT 186
PF+ ++ + + + SI +KE + + C + E K CV+SL+SM+D
Sbjct: 94 ----PFTKSR-QGVTD--SIWLANKEKQSFE---DFCYSPTAIAEHKHCVSSLKSMIDQV 143
Query: 187 ISKLGNN-VEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD 245
IS G+ ++A+S+ N Q + + VKK+G+ ++CH+ N+ VF CH+
Sbjct: 144 ISHFGSTKIKAISS----NFAPYQDQYVVEDVKKVGDN--AVMCHRLNFEKVVFNCHQVR 197
Query: 246 STEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
T Y V L DG K + VCH DT NP+ L + L+V PGTVP+CH + W
Sbjct: 198 ETTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTPGTVPVCHFIGNKAAAW 256
Query: 306 V 306
V
Sbjct: 257 V 257
>EA87_VICFA (P21746) Embryonic abundant protein USP87 precursor
Length = 268
Score = 108 bits (269), Expect = 2e-23
Identities = 87/305 (28%), Positives = 136/305 (44%), Gaps = 72/305 (23%)
Query: 3 PPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDE 62
P + YWQS+ PN+P+PK F+++L P+G +
Sbjct: 24 PREDYWQSIWPNTPLPKTFSDMLIPSGKTNS----------------------------- 54
Query: 63 KIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFL 122
+P+ ++ KQ TLFF + DLH L N N +
Sbjct: 55 ------------LPIKSEEL----KQYSTLFF-EHDLHPRKNFIL------GNTNSVGSI 91
Query: 123 PREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESM 182
R PF+ ++ + + + SI +KE + ++ C + E K CV+SL+SM
Sbjct: 92 IR------PFTKSR-QGVTD--SIWLANKEKQSLE---DFCYSPTAIAEHKHCVSSLKSM 139
Query: 183 VDFTISKLGNN-VEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYC 241
+D IS G+ ++A+S+ N Q + + VKK+G+ ++CH+ N+ VF C
Sbjct: 140 IDQVISHFGSTKIKAISS----NFAPYQDQYVVEDVKKVGDN--AVMCHRLNFEKVVFNC 193
Query: 242 HKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQD 301
H+ T Y V L DG K + VCH DT NP+ L + L+V GTVP+CH +
Sbjct: 194 HQVRDTTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTLGTVPVCHFIGNK 252
Query: 302 HVVWV 306
WV
Sbjct: 253 AAAWV 257
>SVF3_VICFA (P09059) Unknown seed protein 30.1 precursor (VF30.1)
Length = 268
Score = 105 bits (262), Expect = 1e-22
Identities = 86/301 (28%), Positives = 133/301 (43%), Gaps = 72/301 (23%)
Query: 7 YWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEKIIP 66
YWQS+ PN+P+PK F++L P+G +
Sbjct: 28 YWQSIWPNTPLPKTFSDLSIPSGKTNS--------------------------------- 54
Query: 67 LIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREV 126
+P+ ++ KQ TLFF + DLH L N N + R
Sbjct: 55 --------LPIKSEEL----KQYSTLFF-EHDLHPRKNFIL------GNTNSVGSIIR-- 93
Query: 127 ANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFT 186
PF+ ++ + + + SI +KE + ++ C + E K CV+SL+SM+D
Sbjct: 94 ----PFTKSR-QGVTD--SIWLANKEKQSLE---DFCYSPTAIAEHKHCVSSLKSMIDQV 143
Query: 187 ISKLGNN-VEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD 245
IS G+ ++A+S+ N Q + + VKK+G+ ++CH+ N+ VF CH+
Sbjct: 144 ISHFGSTKIKAISS----NFAPYQDQYVVEDVKKVGDN--AVMCHRLNFEKVVFNCHQVR 197
Query: 246 STEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
T Y V L DG K + VCH DT NP+ L + L+V GTVP+CH + W
Sbjct: 198 DTTAYVVSLVASDGTKTKALTVCHHDTRGMNPE-LLYEALEVTLGTVPVCHFIGNKAAAW 256
Query: 306 V 306
V
Sbjct: 257 V 257
>DNLI_SCHPO (P12000) DNA ligase (EC 6.5.1.1)
(Polydeoxyribonucleotide synthase [ATP])
Length = 768
Score = 35.4 bits (80), Expect = 0.18
Identities = 40/170 (23%), Positives = 73/170 (42%), Gaps = 19/170 (11%)
Query: 58 YLNDEKIIPLIYFYPIPIPLNESQIQ--LDDKQNVTLFFLKKDLHHGTKLNLQFKETTSN 115
YL+ K+ P + + + + ES I + + TL +K H L L +T+
Sbjct: 192 YLSINKLGP--DYSGLELGIGESIIMKAIGESTGQTLQQIKLSFHKVGDLGL-VAQTSRQ 248
Query: 116 NNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLC 175
N T F P A +IPF + ++ I + +++ ++KR +S CE E K
Sbjct: 249 NQPTMFKP--AALTIPFLFDSLKKIAQMSGNQSQNRKIGVIKRLLSSCEG----AEPKYL 302
Query: 176 VTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNK 225
+ +LE + + A T + +N QY K +KL ++++
Sbjct: 303 IRALEGKLRLQL--------AEKTILVALANATAQYHADKNGEKLSQQDR 344
>RF1_METKA (Q8TXB5) Peptide chain release factor subunit 1
(Translation termination factor aRF1)
Length = 409
Score = 34.7 bits (78), Expect = 0.31
Identities = 23/73 (31%), Positives = 40/73 (54%), Gaps = 9/73 (12%)
Query: 151 KEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTIS---KLGNNVEAVSTEVDKNSNG 207
KE + K+ + EC G E +L V ++ +VD+ + ++G+NVE +STE ++ G
Sbjct: 337 KEKDEAKKYVEECPECG---EAELNVEEIKDIVDYYVELAEQMGSNVEIISTETEE---G 390
Query: 208 LQQYVIAKGVKKL 220
Q Y +G+ L
Sbjct: 391 AQFYNAFRGLGAL 403
>SMC4_SCHPO (P41004) Structural maintenance of chromosome 4
(Chromosome segregation protein cut3)
Length = 1324
Score = 32.7 bits (73), Expect = 1.2
Identities = 34/145 (23%), Positives = 63/145 (43%), Gaps = 18/145 (12%)
Query: 83 QLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILN 142
+L+D +N L FLK + K N ++ K L + + NS+ K++ L
Sbjct: 335 KLEDSKNSVLSFLKDENELFMKQNQLYRTILYETRNKKTLVQNLLNSL---EGKLQAHLE 391
Query: 143 KF-----SIKEGSKEAEIVKRTISECEANGIKGEEKL------CVTSLESMVDFTIS--- 188
KF I E ++E + ++ ++ + N E+K +E + F ++
Sbjct: 392 KFEQTERDISEKNEEVKSLREKAAKVK-NDCTSEKKTRQSYEQQTVKIEEQLKFLLNKEK 450
Query: 189 KLGNNVEAVSTEVDKNSNGLQQYVI 213
KL ++EA+S E + N L + I
Sbjct: 451 KLKKSIEALSFEKSEAENSLSSHDI 475
>SIR1_YEAST (P21691) Regulatory protein SIR1 (Silent information
regulator 1)
Length = 678
Score = 31.6 bits (70), Expect = 2.6
Identities = 45/172 (26%), Positives = 69/172 (39%), Gaps = 26/172 (15%)
Query: 114 SNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTIS---------ECE 164
S NN +F+ RE P N + N+ K+ EAE+VKR IS +
Sbjct: 304 SKNNDYRFIVREK----PIVENTISNLDYSDIKKQQFTEAEVVKRKISADISQIENVHTQ 359
Query: 165 ANGIKGEEKLCVTSLESMVDFTISKLGNN---VEAVSTEVDKN--------SNGLQQYVI 213
N K + + V + S V ISK + + +S DKN + L++ I
Sbjct: 360 FNSQKEKNNIRVNKVSSEVLDQISKFPVSRVTLLLMSAGQDKNYIELVEELARRLEKICI 419
Query: 214 AKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTI 265
K + L E T + E A F S E Y + LE + +++ T+
Sbjct: 420 EKTTQSLEEIRDTFQANPE--MQASFDKEYYQSIEEYKITLELIKEDLLITL 469
>TFC5_YEAST (P46678) Transcription factor TFIIIB B" component
(TFIIIB90)
Length = 594
Score = 31.2 bits (69), Expect = 3.4
Identities = 37/153 (24%), Positives = 61/153 (39%), Gaps = 12/153 (7%)
Query: 74 PIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFS 133
P PL + Q + + L L KD F +++SN+NGT A +
Sbjct: 115 PAPLQTHRDQKVPRSS-RLASLSKDNESRPSFKPSFLDSSSNSNGT-------ARRLSTI 166
Query: 134 SNKMENILNKFSIKEGSKEAEIVKRTISECEANGIK---GEEKLCVTSLESMVDFTISKL 190
SNK+ + SI E + KR + + K G ++ + S S L
Sbjct: 167 SNKLPKKIRLGSITENDMNLKTFKRHRVLGKPSSAKKPAGAHRISIVSKISPPTAMTDSL 226
Query: 191 GNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEK 223
N + T + ++ + YVI+K VK + +K
Sbjct: 227 DRNEFSSETSTSREADENENYVISK-VKDIPKK 258
>UB10_HUMAN (Q14694) Ubiquitin carboxyl-terminal hydrolase 10 (EC
3.1.2.15) (Ubiquitin thiolesterase 10)
(Ubiquitin-specific processing protease 10)
(Deubiquitinating enzyme 10)
Length = 798
Score = 30.8 bits (68), Expect = 4.4
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 9/62 (14%)
Query: 213 IAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD------STEVYSVPLEG---VDGNMVK 263
I+K + G KNK CH+ +AV Y H +T+V+ + L G +D VK
Sbjct: 712 ISKELLSPGVKNKNFKCHRTYRLFAVVYHHGNSATGGHYTTDVFQIGLNGWLRIDDQTVK 771
Query: 264 TI 265
I
Sbjct: 772 VI 773
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 30.8 bits (68), Expect = 4.4
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 2/51 (3%)
Query: 12 LPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDE 62
+ N + KAF +L H A + +A A G +D V+ YEGGGT LN +
Sbjct: 478 IANLDVSKAFDSLSH-ASIYDTLRAYGAPKGFVDY-VQNTYEGGGTSLNGD 526
>ABAA_EMENI (P20945) Regulatory protein abaA
Length = 796
Score = 30.8 bits (68), Expect = 4.4
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 8/58 (13%)
Query: 12 LPNSPMPKAFTNLLHPA----GYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEKII 65
+P+ P+P A NL HP Y K+ R SNG G R+ G +YL +K +
Sbjct: 69 VPSHPLPTATANLYHPQLLSHRYQVKKLRRLQSNGSSYAGSRR----GRSYLKSQKYL 122
>VG29_BPT4 (P13337) Tail-tube assembly protein Gp29 (Tail length
regulator) (Folylpolyglutamate synthase) (EC 6.3.2.17)
Length = 590
Score = 30.4 bits (67), Expect = 5.8
Identities = 30/130 (23%), Positives = 64/130 (49%), Gaps = 21/130 (16%)
Query: 110 KETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEA--EIVKRTISECEANG 167
++ S+N T+ + +N++ + LN S K +A E++ +T+ E ++N
Sbjct: 12 RKVISDNKPTQEAAKSASNTL--------SGLNDISTKLDDAQAASELIAQTVEE-KSNE 62
Query: 168 IKG-----EEKLCVTSLESMVDFTISKLGNNV-----EAVSTEVDKNSNGLQQYVIAKGV 217
I G E + TS S + ++GNN+ E++ +++DK ++ L+Q + G+
Sbjct: 63 IIGAIDNVESAVSDTSAGSELIAETVEIGNNINKEIGESLGSKLDKLTSLLEQKIQTAGI 122
Query: 218 KKLGEKNKTI 227
++ G T+
Sbjct: 123 QQTGTSLATV 132
>KMHA_DICDI (P42527) Myosin heavy chain kinase A (EC 2.7.1.129)
(MHCK A)
Length = 1146
Score = 30.4 bits (67), Expect = 5.8
Identities = 28/109 (25%), Positives = 47/109 (42%), Gaps = 9/109 (8%)
Query: 107 LQFKETTSNN-----NGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTIS 161
L+ + T+S N NGT P ++ IP S + N +NK I E K+ E +
Sbjct: 323 LKLESTSSGNLMAGLNGTSGRPSSSSHFIPSSVSAAANNINKNEIMEEVKKVEEKLQKKI 382
Query: 162 ECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQ 210
E + K E ++ +E V S++ + + DK N ++Q
Sbjct: 383 REEIDNTKAE----LSKVERSVKDNRSEIEGLEKDCKNQFDKQDNKIKQ 427
>ENDA_NANEQ (Q74MP4) tRNA-splicing endonuclease (EC 3.1.27.9)
(tRNA-intron endonuclease)
Length = 154
Score = 30.4 bits (67), Expect = 5.8
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Query: 51 GYEGGGTYLNDEKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFK 110
GY G L D+K +PLI Y + + E ++ DDK+ FLKK L + + +++K
Sbjct: 3 GYLFGNRVLVDDKELPLIEAYYL-LDKGELEVYEDDKKLSKEEFLKKCLTYDERFLIRYK 61
>YDM6_SCHPO (P87137) Hypothetical protein C57A7.06 in chromosome I
Length = 929
Score = 29.6 bits (65), Expect = 9.9
Identities = 35/119 (29%), Positives = 54/119 (44%), Gaps = 17/119 (14%)
Query: 115 NNNGTK-FLPREVANSI-------PFSSNKMENILNKFSIKEGSKEAEIVKRTISECEAN 166
NN G + F P E A + PF S+ +NK +KEG K+AE K+ S EA
Sbjct: 635 NNTGRRSFKPSEEAAKLSLPSRKNPFVSDSAVLKVNKPEMKEGQKKAEARKKKESPLEAT 694
Query: 167 GIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNK 225
EE + + KL N S++ DK ++ L+ +A +++L E+ K
Sbjct: 695 ----EETNPWLQVPDQRTSSAKKLDKN----SSKADKKNHKLKMDKVA-SLQELVEEPK 744
>VAC3_HELPY (Q48253) Vacuolating cytotoxin precursor
Length = 1310
Score = 29.6 bits (65), Expect = 9.9
Identities = 47/196 (23%), Positives = 77/196 (38%), Gaps = 30/196 (15%)
Query: 85 DDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPR--------EVANSIPFSSNK 136
+D N T F D+H G +NL+ T+N NG +L + A+ + K
Sbjct: 468 EDTNNATATF-NNDIHLGKAVNLRVDAHTANFNGNIYLGKSTNLRVNGHTAHFKNIDATK 526
Query: 137 MENILNKFSIKEG--SKEAEIVKRTISECEANGIKG---EEKLCVTSLESMVDFTISKLG 191
+N LN ++ + + I K T + N IK +E + T ++S +TI G
Sbjct: 527 SDNGLNTSTLDFSGVTDKVNINKLTTAATNVN-IKNFDIKELVVTTRVQSFGQYTI--FG 583
Query: 192 NNVEAVSTEVDKNSNGLQQY------VIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD 245
N+ DK+ G+ + GV G K K ++ + P+ F
Sbjct: 584 ENIG------DKSRIGVVSLQTGYSPAYSGGVTFKGGK-KLVIDEIYHAPWNYFDARNVT 636
Query: 246 STEVYSVPLEGVDGNM 261
E+ L G GN+
Sbjct: 637 DVEINKRILFGAPGNI 652
>DPOD_HUMAN (P28340) DNA polymerase delta catalytic subunit (EC
2.7.7.7) (DNA polymerase delta subunit p125)
Length = 1107
Score = 29.6 bits (65), Expect = 9.9
Identities = 50/196 (25%), Positives = 84/196 (42%), Gaps = 20/196 (10%)
Query: 37 RDASNGGLDVGVRKGYEGGGTYLNDEKIIPLIYFYPIPI-PLNESQIQLDDKQNVTLFFL 95
R A + GL + V K EGG Y I PL +Y +PI L+ S + L +
Sbjct: 561 RQAMHEGLLMPVVKS-EGGEDYTGATVIEPLKGYYDVPIATLDFSSLYPSIMMAHNLCYT 619
Query: 96 KKDLHHGT--KLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILN--KFSIKEGSK 151
L GT KL L + G +F+ V + +EN+L+ K + E +K
Sbjct: 620 TL-LRPGTAQKLGLTEDQFIRTPTGDEFVKTSVRKGL--LPQILENLLSARKRAKAELAK 676
Query: 152 EAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQY 211
E + ++R + + G + S S+ FT +++G + E+ ++ G +
Sbjct: 677 ETDPLRRQV-------LDGRQLALKVSANSVYGFTGAQVG---KLPCLEISQSVTGFGRQ 726
Query: 212 VIAKGVKKLGEKNKTI 227
+I K K+L E T+
Sbjct: 727 MIEK-TKQLVESKYTV 741
>CL02_HUMAN (Q8NHQ8) Protein C12orf2 (Carcinoma associated protein
HOJ-1)
Length = 392
Score = 29.6 bits (65), Expect = 9.9
Identities = 21/115 (18%), Positives = 52/115 (44%), Gaps = 15/115 (13%)
Query: 124 REVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEAN---------------GI 168
+E+ I NK+++ L + E EAE ++R + E + N I
Sbjct: 245 QEIRQKITECENKLKDYLAQIRTMESGLEAEKLQREVQEAQVNEEEVKGKIGKVKGEIDI 304
Query: 169 KGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEK 223
+G++ L + + V+ ++ + ++ E+++ + L+Q + + +++ G K
Sbjct: 305 QGQQSLRLENGIKAVERSLGQATKRLQDKEQELEQLTKELRQVNLQQFIQQTGTK 359
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,088,008
Number of Sequences: 164201
Number of extensions: 1720195
Number of successful extensions: 3156
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 3135
Number of HSP's gapped (non-prelim): 27
length of query: 308
length of database: 59,974,054
effective HSP length: 110
effective length of query: 198
effective length of database: 41,911,944
effective search space: 8298564912
effective search space used: 8298564912
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC148532.11


