

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148526.6 + phase: 0 /pseudo
(420 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
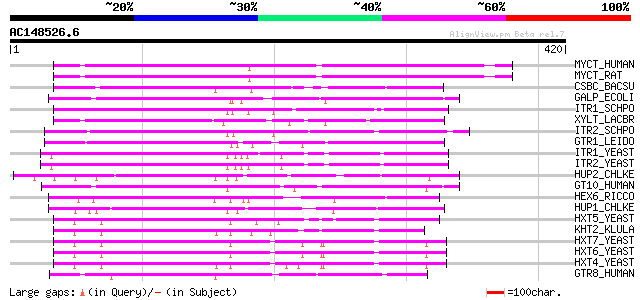
Score E
Sequences producing significant alignments: (bits) Value
MYCT_HUMAN (Q96QE2) Proton myo-inositol cotransporter (H(+)-myo-... 201 2e-51
MYCT_RAT (Q921A2) Proton myo-inositol cotransporter (H(+)-myo-in... 199 1e-50
CSBC_BACSU (P46333) Probable metabolite transport protein csbC 168 2e-41
GALP_ECOLI (P37021) Galactose-proton symporter (Galactose transp... 159 1e-38
ITR1_SCHPO (Q10286) Myo-inositol transporter 1 158 3e-38
XYLT_LACBR (O52733) D-xylose-proton symporter (D-xylose transpor... 151 3e-36
ITR2_SCHPO (P87110) Myo-inositol transporter 2 150 6e-36
GTR1_LEIDO (Q01440) Membrane transporter D1 149 1e-35
ITR1_YEAST (P30605) Myo-inositol transporter 1 147 5e-35
ITR2_YEAST (P30606) Myo-inositol transporter 2 142 2e-33
HUP2_CHLKE (Q39524) H(+)/hexose cotransporter 2 (Galactose-H+ sy... 118 2e-26
GT10_HUMAN (O95528) Solute carrier family 2, facilitated glucose... 110 7e-24
HEX6_RICCO (Q07423) Hexose carrier protein HEX6 109 2e-23
HUP1_CHLKE (P15686) H(+)/hexose cotransporter 1 108 3e-23
HXT5_YEAST (P38695) Probable glucose transporter HXT5 107 4e-23
KHT2_KLULA (P53387) Hexose transporter 2 107 7e-23
HXT7_YEAST (P39004) High-affinity hexose transporter HXT6 105 3e-22
HXT6_YEAST (P39003) High-affinity hexose transporter HXT6 105 3e-22
HXT4_YEAST (P32467) Low-affinity glucose transporter HXT4 (Low-a... 104 5e-22
GTR8_HUMAN (Q9NY64) Solute carrier family 2, facilitated glucose... 103 8e-22
>MYCT_HUMAN (Q96QE2) Proton myo-inositol cotransporter
(H(+)-myo-inositol cotransporter) (Hmit)
Length = 629
Score = 201 bits (512), Expect = 2e-51
Identities = 130/379 (34%), Positives = 199/379 (52%), Gaps = 47/379 (12%)
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGR 93
GG LFGYDTGV+SGA L ++ Q+ QE +VS A + A GG +N GR
Sbjct: 72 GGFLFGYDTGVVSGAMLLLK---RQLSLDALWQELLVSSTVGAAAVSALAGGALNGVFGR 128
Query: 94 KKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGA 153
+ IL+A +F AG+ V+AAA ++ GRL+VGLG+G ASMT P+YI+E SP +RG
Sbjct: 129 RAAILLASALFTAGSAVLAAANNKETLLAGRLVVGLGIGIASMTVPVYIAEVSPPNLRGR 188
Query: 154 LVCRLHQWFTHNRWSISLLPHQFGIYQ-------------------------------VY 182
LV + T ++ S++ F Q +
Sbjct: 189 LVTINTLFITGGQFFASVVDGAFSYLQKDGWRYMLGLAXVPAVIQFFGFLFLPESPRWLI 248
Query: 183 RQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVR 242
++ + ++A++ILS++ ++EE ++ +IE E+ E G G + + L S R
Sbjct: 249 QKGQTQKARRILSQMRGNQTIDEEYDSIKNNIEEEEKEVGSAGPVICRML----SYPPTR 304
Query: 243 RGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMV 302
R L G +Q+ QQ+ GINTIMYYS TI+Q +G+ + A L+ VT+ N + T+V +
Sbjct: 305 RALIVGCGLQMFQQLSGINTIMYYSATILQMSGVEDDRLAIWLASVTAFTNFIFTLVGVW 364
Query: 303 LIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTTAP 362
L+++ GRRKL SL G V+L+ L++ F +A +P ++ + G N+TC Y+
Sbjct: 365 LVEKVGRRKLTFGSLAGTTVALIILALGFVLSAQVSPRITFKPIAPSGQNATCTRYS--- 421
Query: 363 NHLSWNCMQC-LHEDCAFC 380
C +C L DC FC
Sbjct: 422 -----YCNECMLDPDCGFC 435
>MYCT_RAT (Q921A2) Proton myo-inositol cotransporter
(H(+)-myo-inositol cotransporter) (Hmit)
Length = 618
Score = 199 bits (505), Expect = 1e-50
Identities = 129/379 (34%), Positives = 198/379 (52%), Gaps = 47/379 (12%)
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGR 93
GG LFGYDTGV+SGA L +R Q+ QE +VS A A + A GG +N +GR
Sbjct: 61 GGFLFGYDTGVVSGAMLLLR---RQMRLGAMWQELLVSGAVGAAAVAALAGGALNGALGR 117
Query: 94 KKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGA 153
+ IL+A + G+ V+AAA ++ GRL+VGLG+G ASMT P+YI+E SP +RG
Sbjct: 118 RSAILLASALCTVGSAVLAAAANKETLLAGRLVVGLGIGIASMTVPVYIAEVSPPNLRGR 177
Query: 154 LVCRLHQWFTHNRWSISLLPHQFGIYQ-------------------------------VY 182
LV + T ++ S++ F Q +
Sbjct: 178 LVTINTLFITGGQFFASVVDGAFSYLQKDGWRYMLGLAAIPAVIQFLGFLFLPESPRWLI 237
Query: 183 RQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVR 242
++ + ++A++ILS++ ++EE ++ SIE E+ E G + + L S R
Sbjct: 238 QKGQTQKARRILSQMRGNQTIDEEYDSIRNSIEEEEKEASAAGPIICRML----SYPPTR 293
Query: 243 RGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMV 302
R L G +Q+ QQ+ GINTIMYYS TI+Q +G+ + A L+ +T+ N + T+V +
Sbjct: 294 RALAVGCGLQMFQQLSGINTIMYYSATILQMSGVEDDRLAIWLASITAFTNFIFTLVGVW 353
Query: 303 LIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTTAP 362
L+++ GRRKL SL G V+L L++ F +A +P ++ + + G N+TC Y+
Sbjct: 354 LVEKVGRRKLTFGSLAGTTVALTILALGFLLSAQVSPRVTFRPTAPSGQNATCTEYS--- 410
Query: 363 NHLSWNCMQC-LHEDCAFC 380
C +C L DC FC
Sbjct: 411 -----YCNECMLDPDCGFC 424
>CSBC_BACSU (P46333) Probable metabolite transport protein csbC
Length = 461
Score = 168 bits (426), Expect = 2e-41
Identities = 111/324 (34%), Positives = 172/324 (52%), Gaps = 44/324 (13%)
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGR 93
GGLL+GYDTGVISGA L+I ++ L +VSM GAI G+A G +D+ GR
Sbjct: 17 GGLLYGYDTGVISGALLFINNDIPLTTLTEGL---VVSMLLLGAIFGSALSGTCSDRWGR 73
Query: 94 KKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGA 153
+K + + ++F+ GAL A + ++I R+++GL VG ++ P+Y+SE +P KIRG
Sbjct: 74 RKVVFVLSIIFIIGALACAFSQTIGMLIASRVILGLAVGGSTALVPVYLSEMAPTKIRGT 133
Query: 154 L-----------------VCRLHQWFTHNRWSISLLPHQFGIYQV------------YRQ 184
L V L F RW + L + + ++
Sbjct: 134 LGTMNNLMIVTGILLAYIVNYLFTPFEAWRWMVGLAAVPAVLLLIGIAFMPESPRWLVKR 193
Query: 185 SKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRG 244
EEEA++I++ + P ++E E+ M + EAEK E L LK W +R
Sbjct: 194 GSEEEARRIMNITHDPKDIEMELAEMKQG-EAEKKETTL------GVLKAKW----IRPM 242
Query: 245 LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLI 304
L G+ + + QQ VGINT++YY+PTI AG+ ++++A ++ LN + I +M+LI
Sbjct: 243 LLIGVGLAIFQQAVGINTVIYYAPTIFTKAGLGTSASALG-TMGIGILNVIMCITAMILI 301
Query: 305 DRFGRRKLMLISLIGIFVSLVTLS 328
DR GR+KL++ +GI +SL LS
Sbjct: 302 DRVGRKKLLIWGSVGITLSLAALS 325
>GALP_ECOLI (P37021) Galactose-proton symporter (Galactose
transporter)
Length = 464
Score = 159 bits (403), Expect = 1e-38
Identities = 109/337 (32%), Positives = 174/337 (51%), Gaps = 37/337 (10%)
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
LA GLLFG D GVI+GA +I DEF+ QE +VS GA +GA G+++
Sbjct: 21 LAALAGLLFGLDIGVIAGALPFIADEFQITSHT---QEWVVSSMMFGAAVGAVGSGWLSF 77
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
K+GRKK++++ ++FVAG+L AAAP V+I+ R+L+GL VG AS TAPLY+SE +P K
Sbjct: 78 KLGRKKSLMIGAILFVAGSLFSAAAPNVEVLILSRVLLGLAVGVASYTAPLYLSEIAPEK 137
Query: 150 IRGALVCRLHQWFTHN-----------------RW--SISLLPH---QFGIYQVYRQSKE 187
IRG+++ T RW + ++P G++ + +
Sbjct: 138 IRGSMISMYQLMITIGILGAYLSDTAFSYTGAWRWMLGVIIIPAILLLIGVFFLPDSPRW 197
Query: 188 EEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWS----NDVVRR 243
AK+ R + E + + ++ K E I SL K G W+ N RR
Sbjct: 198 FAAKR------RFVDAERVLLRLRDTSAEAKRELDEIRESLQVKQSG-WALFKENSNFRR 250
Query: 244 GLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVL 303
++ G+ +QV+QQ G+N IMYY+P I + AG + + +++ N + T +++ L
Sbjct: 251 AVFLGVLLQVMQQFTGMNVIMYYAPKIFELAGYTNTTEQMWGTVIVGLTNVLATFIAIGL 310
Query: 304 IDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPS 340
+DR+GR+ + + + + + L T H+PS
Sbjct: 311 VDRWGRKPTLTLGFLVMAAGMGVLG-TMMHIGIHSPS 346
>ITR1_SCHPO (Q10286) Myo-inositol transporter 1
Length = 575
Score = 158 bits (399), Expect = 3e-38
Identities = 106/332 (31%), Positives = 169/332 (49%), Gaps = 38/332 (11%)
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGR 93
GGLLFGYDTGVISGA + I + +E I S S GA++G G + D GR
Sbjct: 97 GGLLFGYDTGVISGALVVIGTSLGGHELTNGGKEFITSATSLGALLGGIIAGALADFFGR 156
Query: 94 KKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGA 153
K I +A ++ + G++V A W +I+GR ++G GVG AS+ PLY+SE +P+KIRG
Sbjct: 157 KPVIAIASIIIIVGSIVQVTAHHLWHMIVGRFVIGWGVGIASLIIPLYLSEIAPSKIRGR 216
Query: 154 LVCRLHQWFT----------------HNRW----SISLLPHQFGIY----------QVYR 183
LV T HN W ++++P F ++ + +
Sbjct: 217 LVIIYVLLITAGQVIAYGIDTAFEHVHNGWRWMVGLAMVPAAFQLFILIWLPESPRLLVK 276
Query: 184 QSKEEEAKQILSKIY---RPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDV 240
+ + +EA L++IY P E++ ++ + E + G + + K + N
Sbjct: 277 KERSQEAYNTLARIYPTAHPYEIKTKLYLIQEGV--RDPFSGSRWQKIVKTFKELYFNPS 334
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVS 300
R L +Q +QQ+ G N++MY+S TI + G +N T A L+ + N V TIV+
Sbjct: 335 NFRALILACGLQAMQQLSGFNSLMYFSSTIFEVVGF-NNPT--ATGLIIAATNFVFTIVA 391
Query: 301 MVLIDRFGRRKLMLISLIGIFVSLVTLSVTFN 332
+ID FGRR L+L+++ G+ +L+ +V F+
Sbjct: 392 FGVIDFFGRRILLLLTVWGMIAALIVCAVAFH 423
>XYLT_LACBR (O52733) D-xylose-proton symporter (D-xylose
transporter)
Length = 457
Score = 151 bits (382), Expect = 3e-36
Identities = 100/324 (30%), Positives = 171/324 (51%), Gaps = 40/324 (12%)
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGR 93
GGLLFGYDTGVISGA L+I+ +Q++ W Q +VS GAI+GAA G +D+ GR
Sbjct: 16 GGLLFGYDTGVISGAILFIQ---KQMNLGSWQQGWVVSAVLLGAILGAAIIGPSSDRFGR 72
Query: 94 KKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGA 153
+K +L++ ++F GAL A +P W +II R+++G+ VGAAS P Y++E +P+ RG
Sbjct: 73 RKLLLLSAIIFFVGALGSAFSPEFWTLIISRIILGMAVGAASALIPTYLAELAPSDKRGT 132
Query: 154 LVCRLHQ-------------------WFTHNRWSISLLPHQFGIYQVYRQSKEEEAKQIL 194
V L Q ++T RW + + + E + ++
Sbjct: 133 -VSSLFQLMVMTGILLAYITNYSFSGFYTGWRWMLGFAAIPAALLFLGGLILPESPRFLV 191
Query: 195 SKIYRPGEVEEEMKAM-----HESIEAEKAEDGLIGHSLAQKLKGAWS---NDVVRRGLY 246
+ G ++E + H+ + K + + A+ + G WS +VR L
Sbjct: 192 ----KSGHLDEARHVLDTMNKHDQVAVNKEINDI--QESAKIVSGGWSELFGKMVRPSLI 245
Query: 247 AGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGL-NAVGTIVSMVLID 305
GI + + QQ++G NT++YY+PTI F + +A L+ + G+ N + T +++ ++D
Sbjct: 246 IGIGLAIFQQVMGCNTVLYYAPTI--FTDVGFGVSAALLAHIGIGIFNVIVTAIAVAIMD 303
Query: 306 RFGRRKLMLISLIGIFVSLVTLSV 329
+ R+K++ I +G+ +SL +S+
Sbjct: 304 KIDRKKIVNIGAVGMGISLFVMSI 327
>ITR2_SCHPO (P87110) Myo-inositol transporter 2
Length = 557
Score = 150 bits (379), Expect = 6e-36
Identities = 108/355 (30%), Positives = 173/355 (48%), Gaps = 42/355 (11%)
Query: 27 LMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGY 86
L +A GLLFGYDTGVISGA + + V +E I S S A+I A G+
Sbjct: 84 LSAVAGISGLLFGYDTGVISGALAVLGSDLGHV-LSSGQKELITSATSFAALISATTSGW 142
Query: 87 MNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEAS 146
+ D +GRK+ +L AD +FV G+++MAA+ ++++GR +VG G+G S+ P+YI+E +
Sbjct: 143 LADWVGRKRLLLCADAIFVIGSVIMAASRNVAMMVVGRFIVGYGIGLTSLIVPMYITELA 202
Query: 147 PAKIRGALVCRLHQWFT----------------HNRWS--------------ISLLPHQF 176
PA++RG LV + T H W ISL
Sbjct: 203 PARLRGRLVIIYVVFITGGQLIAYSLNAAFEHVHQGWRIMFGIGAAPALGQLISLFWTPE 262
Query: 177 GIYQVYRQSKEEEAKQILSKIY---RPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLK 233
+ R + E+ +ILS+I+ +P E+ ++ + E ++ + E H LK
Sbjct: 263 SPRYLLRHNHVEKVYKILSRIHPEAKPAEIAYKVSLIQEGVKVDFPEGNKFQH-FFHSLK 321
Query: 234 GAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLN 293
++ RR L+ G +Q QQ G N I Y+S I Q G + ++S+V N
Sbjct: 322 VLFTVPSNRRSLFIGCFLQWFQQFSGTNAIQYFSAIIFQSVGF---KNSISVSIVVGATN 378
Query: 294 AVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLS 348
V TIV+ + IDR GRR+++L + + L ++ + H P+ + Q++ S
Sbjct: 379 FVFTIVAFMFIDRIGRRRILLCTSAVMIAGLALCAIAY----HFLPADTTQNTNS 429
>GTR1_LEIDO (Q01440) Membrane transporter D1
Length = 547
Score = 149 bits (377), Expect = 1e-35
Identities = 104/330 (31%), Positives = 172/330 (51%), Gaps = 37/330 (11%)
Query: 27 LMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGY 86
+M A GG LFGYDTGVI+ A ++D F + W IV++A AGA +GA G+
Sbjct: 5 VMLCAALGGFLFGYDTGVINAALFQMKDHFG-FSEHSWQYALIVAIAIAGAFVGAFISGF 63
Query: 87 MNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEAS 146
++ GR+ I +AD +FV G+++M AAP V+++ R++VGL +G +S T P+Y++E +
Sbjct: 64 ISAAFGRRPCIAVADALFVIGSVLMGAAPNVEVVLVSRVIVGLAIGISSATIPVYLAEVT 123
Query: 147 PAKIRGALVCRLHQWFTHNR-------------------WSISL----LPHQFGIYQVY- 182
K RGA + + + T + W +++ LP + Q +
Sbjct: 124 SPKHRGATIVLNNLFLTGGQFVAAGFTAIMVVFTSKNIGWRVAIGIGALP---AVVQAFC 180
Query: 183 -RQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAE--DGLIGHSLAQKLKGAWSND 239
E + +LSK + + KA+ + E + E +G S+ + + D
Sbjct: 181 LLFFLPESPRWLLSKGH-----ADRAKAVADKFEVDLCEFQEGDELPSVRIDYRPLMARD 235
Query: 240 VVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIV 299
+R + +Q++QQ GINTIMYYS I+ AG LS+ + +NA+ T V
Sbjct: 236 -MRFRVVLSSGLQIIQQFSGINTIMYYSSVILYDAGFRDAIMPVVLSIPLAFMNALFTAV 294
Query: 300 SMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
++ +DRFGRR+++LIS+ G V LV +++
Sbjct: 295 AIFTVDRFGRRRMLLISVFGCLVLLVVIAI 324
>ITR1_YEAST (P30605) Myo-inositol transporter 1
Length = 584
Score = 147 bits (371), Expect = 5e-35
Identities = 109/349 (31%), Positives = 176/349 (50%), Gaps = 49/349 (14%)
Query: 24 SPYLMRL---ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIG 80
SP+++ L A G +FGYDTG IS A + I + + +E + + S GA+I
Sbjct: 83 SPFIITLTFVASISGFMFGYDTGYISSALISIGTDLDHKVLTYGEKEIVTAATSLGALIT 142
Query: 81 AAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPL 140
+ F G D GRK+ ++ ++++FV GA++ +A W + +GRL++G GVG S+ APL
Sbjct: 143 SIFAGTAADIFGRKRCLMGSNLMFVIGAILQVSAHTFWQMAVGRLIMGFGVGIGSLIAPL 202
Query: 141 YISEASPAKIRGALVCRLHQWFT----------------HNRWSI----SLLPH--QFGI 178
+ISE +P IRG L W T +N W I SL+P QF
Sbjct: 203 FISEIAPKMIRGRLTVINSLWLTGGQLVAYGCGAGLNYVNNGWRILVGLSLIPTAVQFTC 262
Query: 179 ---------YQVYRQSKEEEAKQILSKIYRPGEVE------EEMKAMHESIEAEKAEDGL 223
Y V + A ++L + Y E EE+ +++SI + + +
Sbjct: 263 LCFLPDTPRYYVMKGDLAR-ATEVLKRSYTDTSEEIIERKVEELVTLNQSIPGKNVPEKV 321
Query: 224 IGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAF 283
+ ++L SN R L G +Q +QQ G N++MY+S TI + G ++S
Sbjct: 322 --WNTIKELHTVPSN---LRALIIGCGLQAIQQFTGWNSLMYFSGTIFETVGFKNSS--- 373
Query: 284 ALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFN 332
A+S++ SG N + T+V+ ID+ GRR ++LI L G+ ++LV S+ F+
Sbjct: 374 AVSIIVSGTNFIFTLVAFFSIDKIGRRTILLIGLPGMTMALVVCSIAFH 422
>ITR2_YEAST (P30606) Myo-inositol transporter 2
Length = 612
Score = 142 bits (358), Expect = 2e-33
Identities = 106/348 (30%), Positives = 172/348 (48%), Gaps = 47/348 (13%)
Query: 24 SPYLMRL---ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIG 80
SP+++ L A G +FGYDTG IS A + I + + +E I + S GA+I
Sbjct: 109 SPFIITLTFVASISGFMFGYDTGYISSALISINRDLDNKVLTYGEKELITAATSLGALIT 168
Query: 81 AAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPL 140
+ G D GR+ ++ ++++F+ GA++ A W + GRL++G GVG S+ +PL
Sbjct: 169 SVGAGTAADVFGRRPCLMFSNLMFLIGAILQITAHKFWQMAAGRLIMGFGVGIGSLISPL 228
Query: 141 YISEASPAKIRGALVCRLHQWFT----------------HNRWSI----SLLPH--QFGI 178
+ISE +P IRG L W T N W I SL+P QF
Sbjct: 229 FISEIAPKMIRGRLTVINSLWLTGGQLIAYGCGAGLNHVKNGWRILVGLSLIPTVLQFSF 288
Query: 179 Y--------QVYRQSKEEEAKQILSKIYRPGEVE------EEMKAMHESIEAEKAEDGLI 224
+ + + AK +L + Y E E EE+ ++++SI +
Sbjct: 289 FCFLPDTPRYYVMKGDLKRAKMVLKRSYVNTEDEIIDQKVEELSSLNQSIPGKNPITKF- 347
Query: 225 GHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFA 284
++ ++L SN R L G +Q +QQ G N++MY+S TI + G ++S A
Sbjct: 348 -WNMVKELHTVPSN---FRALIIGCGLQAIQQFTGWNSLMYFSGTIFETVGFKNSS---A 400
Query: 285 LSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFN 332
+S++ SG N V T+++ ID+ GRR ++LI L G+ V+LV ++ F+
Sbjct: 401 VSIIVSGTNFVFTLIAFFCIDKIGRRYILLIGLPGMTVALVICAIAFH 448
>HUP2_CHLKE (Q39524) H(+)/hexose cotransporter 2 (Galactose-H+
symporter)
Length = 540
Score = 118 bits (296), Expect = 2e-26
Identities = 115/398 (28%), Positives = 177/398 (43%), Gaps = 72/398 (18%)
Query: 4 GGHPLADKTEFTEC---WRRTAESPYLMRLAL---SGGLLFGYDTGVISGA--------- 48
GG P+A T + R + Y+ +AL SGGLLFGYD GV G
Sbjct: 3 GGGPVASTTTNRASQYGYARGGLNWYIFIVALTAGSGGLLFGYDIGVTGGVTSMPEFLQK 62
Query: 49 ---SLYIRDEFEQVDKKPW-------LQETIVSMASAGAIIGAAFGGYMNDKMGRKKTIL 98
S+Y R + K P+ LQ S AG + + F G + + GRK T+L
Sbjct: 63 FFPSIYDRTQQPSDSKDPYCTYDDQKLQLFTSSFFLAGMFV-SFFAGSVVRRWGRKPTML 121
Query: 99 MADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGAL---- 154
+A V+F+AGA + A A +++IGR+L+G GVG + PLY+SE +P K RG L
Sbjct: 122 IASVLFLAGAGLNAGAQDLAMLVIGRVLLGFGVGGGNNAVPLYLSECAPPKYRGGLNMMF 181
Query: 155 ---------VCRLHQWFT---HNRWSIS----------------LLPHQFGIYQVYRQSK 186
V +L + T +N W +S LLP + +
Sbjct: 182 QLAVTIGIIVAQLVNYGTQTMNNGWRLSLGLAGVPAIILLIGSLLLPETPN--SLIERGH 239
Query: 187 EEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRGLY 246
+ +L+++ R V+ E + + AE++ + S A +S ++ L
Sbjct: 240 RRRGRAVLARLRRTEAVDTEFEDI--CAAAEESTRYTLRQSWAALFSRQYSPMLIVTSLI 297
Query: 247 AGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDR 306
A ++QQ+ GIN IM+Y P + G A ++ A +++ +N T VS+ +D+
Sbjct: 298 A-----MLQQLTGINAIMFYVPVLFSSFGTARHA-ALLNTVIIGAVNVAATFVSIFSVDK 351
Query: 307 FGRRKLMLIS----LIGIFVSLVTLSVTFNQAAHHAPS 340
FGRR L L IG V+ L V N+ + PS
Sbjct: 352 FGRRGLFLEGGIQMFIGQVVTAAVLGVELNKYGTNLPS 389
>GT10_HUMAN (O95528) Solute carrier family 2, facilitated glucose
transporter, member 10 (Glucose transporter type 10)
Length = 541
Score = 110 bits (275), Expect = 7e-24
Identities = 92/329 (27%), Positives = 156/329 (46%), Gaps = 22/329 (6%)
Query: 25 PYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFG 84
P ++L GGL FGY+ VISGA L ++ +F + QE +V GA++ + G
Sbjct: 9 PLCASVSLLGGLTFGYELAVISGALLPLQLDFGLSCLE---QEFLVGSLLLGALLASLVG 65
Query: 85 GYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISE 144
G++ D GRK+ IL +++V +AG+L + A + +++GR +VG + +SM +Y+SE
Sbjct: 66 GFLIDCYGRKQAILGSNLVLLAGSLTLGLAGSLAWLVLGRAVVGFAISLSSMACCIYVSE 125
Query: 145 ASPAKIRGALVCRLHQWFT-----HNRWSISLLPHQFGIYQVYRQSKEEEAKQILSKIYR 199
+ RG LV T + +L +G ++ + Q LS ++
Sbjct: 126 LVGPRQRGVLVSLYEAGITVGILLSYALNYALAGTPWGWRHMFGWATAPAVLQSLSLLFL 185
Query: 200 PGEVEEEMKAMHESI------EAEKAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQV 253
P +E A H+ + EA K G +S + + D +R G+ + +
Sbjct: 186 PAGTDE--TATHKDLIPLQGGEAPKLGPGRPRYSFLDLFR---ARDNMRGRTTVGLGLVL 240
Query: 254 VQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLM 313
QQ+ G ++ Y+ TI G S+A S+ + T+ +M L+DR GRR L+
Sbjct: 241 FQQLTGQPNVLCYASTIFSSVGFHGGSSAVLASVGLGAVKVAATLTAMGLVDRAGRRALL 300
Query: 314 L--ISLIGIFVSLVTLSVTFNQAAHHAPS 340
L +L+ + VS + L V+F PS
Sbjct: 301 LAGCALMALSVSGIGL-VSFAVPMDSGPS 328
>HEX6_RICCO (Q07423) Hexose carrier protein HEX6
Length = 510
Score = 109 bits (272), Expect = 2e-23
Identities = 90/350 (25%), Positives = 156/350 (43%), Gaps = 68/350 (19%)
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKK--------------PWLQETIVSM 72
+A GG++FGYD GV G + +++ F V +K L + S
Sbjct: 28 MAAMGGVIFGYDIGVSGGVTSMDPFLKKFFPDVYRKMKEDTEISNYCKFDSQLLTSFTSS 87
Query: 73 ASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
++ + F + GRK +IL+ VF+A A + AA +++I GR+L+G+GVG
Sbjct: 88 LYVAGLVASFFASSVTRAFGRKPSILLGGXVFLAXAALGGAAVNVYMLIFGRVLLGVGVG 147
Query: 133 AASMTAPLYISEASPAKIRGA-------------LVCRLHQWFTHN-----RWSISLLPH 174
A+ PLY+SE +P + RGA L L + T W ISL
Sbjct: 148 FANQAVPLYLSEMAPPRYRGAINNGFQFSVGIGALSANLINYGTEKIEGGWGWRISLAMA 207
Query: 175 Q-------FGIY--------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKA 219
FG + R + E AK +L ++ +V+ E+
Sbjct: 208 AVPAAILTFGALFLPETPNSLIQRSNDHERAKLMLQRVRGTTDVQAEL------------ 255
Query: 220 EDGLIGHSLAQKLKGAWSNDVVRRG----LYAGITVQVVQQIVGINTIMYYSPTIVQFAG 275
D LI S+ + +++RR L + + QQ+ GIN I +Y+P + + G
Sbjct: 256 -DDLIKASIISRTIQHPFKNIMRRKYRPQLVMAVAIPFFQQVTGINVIAFYAPILFRTIG 314
Query: 276 IASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLV 325
+ +++ + S+VT + + T +SM+++D+ GRR L + + +FV+ +
Sbjct: 315 LEESASLLS-SIVTGLVGSASTFISMLIVDKLGRRALFIFGGVQMFVAQI 363
>HUP1_CHLKE (P15686) H(+)/hexose cotransporter 1
Length = 534
Score = 108 bits (270), Expect = 3e-23
Identities = 94/353 (26%), Positives = 152/353 (42%), Gaps = 67/353 (18%)
Query: 30 LALSGGLLFGYDTGVISGA---SLYIRDEFEQV--------DKKPW-------LQETIVS 71
+A GGLL GYD GV G + + F V + P+ LQ + S
Sbjct: 33 MAACGGLLLGYDNGVTGGVVSLEAFEKKFFPDVWAKKQEVHEDSPYCTYDNAKLQLFVSS 92
Query: 72 MASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGV 131
+ AG ++ F ++ GRK T+ + FVAG LV A A ++I+GR+L+G GV
Sbjct: 93 LFLAG-LVSCLFASWITRNWGRKVTMGIGGAFFVAGGLVNAFAQDMAMLIVGRVLLGFGV 151
Query: 132 GAASMTAPLYISEASPAKIRGALVCRLHQWFT----------------HNRWSISL---- 171
G S P Y+SE +P RG L + T N W +SL
Sbjct: 152 GLGSQVVPQYLSEVAPFSHRGMLNIGYQLFVTIGILIAGLVNYAVRDWENGWRLSLGPAA 211
Query: 172 ------------LPHQFGIYQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKA 219
LP + + K E+ +++L K+ EV+ E + ++E +
Sbjct: 212 APGAILFLGSLVLPESPNF--LVEKGKTEKGREVLQKLCGTSEVDAEFADIVAAVEIARP 269
Query: 220 EDGLIGHSLAQKLKGAWSNDVVRR---GLYAGITVQVVQQIVGINTIMYYSPTIVQFAGI 276
++ +W++ RR L +Q QQ GIN I++Y P + G
Sbjct: 270 IT----------MRQSWASLFTRRYMPQLLTSFVIQFFQQFTGINAIIFYVPVLFSSLGS 319
Query: 277 ASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
A NS A ++V +N T+++++ D+FGRR L++ I ++++T V
Sbjct: 320 A-NSAALLNTVVVGAVNVGSTLIAVMFSDKFGRRFLLIEGGIQCCLAMLTTGV 371
>HXT5_YEAST (P38695) Probable glucose transporter HXT5
Length = 592
Score = 107 bits (268), Expect = 4e-23
Identities = 96/337 (28%), Positives = 149/337 (43%), Gaps = 54/337 (16%)
Query: 34 GGLLFGYDTGVISG----ASLYIRDEFEQVDKKPWLQET----IVSMASAGAIIGAAFGG 85
GG +FG+DTG ISG R + + +L + +VS+ + G IG
Sbjct: 96 GGFVFGWDTGTISGFVRQTDFIRRFGSTRANGTTYLSDVRTGLMVSIFNIGCAIGGIVLS 155
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYISE 144
+ D GRK ++ V++ G ++ A+ W IGR++ GLGVG ++ AP+ ISE
Sbjct: 156 KLGDMYGRKIGLMTVVVIYSIGIIIQIASIDKWYQYFIGRIISGLGVGGITVLAPMLISE 215
Query: 145 ASPAKIRGALVCRLHQWFTHN------------------RWSISL-LPHQFGIYQVYRQS 185
SP ++RG LV T +W + L L + I+ + +
Sbjct: 216 VSPKQLRGTLVSCYQLMITFGIFLGYCTNFGTKNYSNSVQWRVPLGLCFAWSIFMIVGMT 275
Query: 186 -------------KEEEAKQILSKIYRPGE----VEEEMKAMHESIEAEKAEDGLIGHSL 228
K EEAK+ L++ + E V EM+ SIEAE+ S
Sbjct: 276 FVPESPRYLVEVGKIEEAKRSLARANKTTEDSPLVTLEMENYQSSIEAERLAGSA---SW 332
Query: 229 AQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLV 288
+ + G + RR L G+ +Q +QQ+ G N YY TI Q G+ +F ++V
Sbjct: 333 GELVTG--KPQMFRRTL-MGMMIQSLQQLTGDNYFFYYGTTIFQAVGL---EDSFETAIV 386
Query: 289 TSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLV 325
+N V T S+ +DRFGRR +L +G+ V
Sbjct: 387 LGVVNFVSTFFSLYTVDRFGRRNCLLWGCVGMICCYV 423
>KHT2_KLULA (P53387) Hexose transporter 2
Length = 566
Score = 107 bits (266), Expect = 7e-23
Identities = 91/326 (27%), Positives = 151/326 (45%), Gaps = 54/326 (16%)
Query: 34 GGLLFGYDTGVISG----ASLYIRDEFEQVDKKPWLQET----IVSMASAGAIIGAAFGG 85
GG +FG+DTG ISG R E+ D +L IVS+ + G IG
Sbjct: 72 GGFVFGWDTGTISGFVNQTDFIRRFGQEKADGSHYLSNVRTGLIVSIFNIGCAIGGIILS 131
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYISE 144
+ D GR+ +++ +++V G ++ A+ W IGR++ GLGVG S+ +P+ ISE
Sbjct: 132 KLGDMYGRRIGLMIVVLIYVVGIIIQIASIDKWYQYFIGRIISGLGVGGISVLSPMLISE 191
Query: 145 ASPAKIRGALV---------------CRLHQWFTHN---RWSISL-LPHQFGIYQV---- 181
+P IRG LV C + T++ +W + L L + I+ +
Sbjct: 192 TAPKHIRGTLVSFYQLMITFGIFLGYCTNYGTKTYSNSVQWRVPLGLCFAWAIFMITGML 251
Query: 182 ---------YRQSKEEEAKQILSK----IYRPGEVEEEMKAMHESIEAEKAEDGLIGHSL 228
+ + +EAK+ ++K Y V+ E+ + +EAE+ L G +
Sbjct: 252 FVPESPRFLVEKDRIDEAKRSIAKSNKVSYEDPAVQAEVDLICAGVEAER----LAGSAS 307
Query: 229 AQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLV 288
++L + V + L G+ +Q QQ+ G N YY TI G+ +F S+V
Sbjct: 308 IKELFS--TKTKVFQRLIMGMLIQSFQQLTGNNYFFYYGTTIFNSVGM---DDSFETSIV 362
Query: 289 TSGLNAVGTIVSMVLIDRFGRRKLML 314
+N T V++ ++D+FGRRK +L
Sbjct: 363 LGIVNFASTFVAIYVVDKFGRRKCLL 388
>HXT7_YEAST (P39004) High-affinity hexose transporter HXT6
Length = 570
Score = 105 bits (261), Expect = 3e-22
Identities = 90/342 (26%), Positives = 156/342 (45%), Gaps = 51/342 (14%)
Query: 34 GGLLFGYDTGVISG----ASLYIRDEFEQVDKKPWLQET----IVSMASAGAIIGAAFGG 85
GG +FG+DTG ISG R + D +L + IVS+ + G IG
Sbjct: 75 GGFVFGWDTGTISGFINQTDFIRRFGMKHKDGTNYLSKVRTGLIVSIFNIGCAIGGIILS 134
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYISE 144
+ D GRK +++ V+++ G ++ A+ W IGR++ GLGVG ++ +P+ ISE
Sbjct: 135 KLGDMYGRKVGLIVVVVIYIIGIIIQIASINKWYQYFIGRIISGLGVGGIAVLSPMLISE 194
Query: 145 ASPAKIRGALVCRLHQWFTHN------------------RWSISL-LPHQFGIYQVYRQS 185
SP +RG LV T +W + L L + ++ + +
Sbjct: 195 VSPKHLRGTLVSCYQLMITAGIFLGYCTNFGTKNYSNSVQWRVPLGLCFAWALFMIGGMT 254
Query: 186 KEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAE-------DGLIGHSLAQKLKG--AW 236
E+ + L+++ G++EE +++ S + + + ++ A+KL G +W
Sbjct: 255 FVPESPRYLAEV---GKIEEAKRSIAVSNKVAVDDPSVLAEVEAVLAGVEAEKLAGNASW 311
Query: 237 -----SNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSG 291
S V + L G +Q +QQ+ G N YY TI + G+ S +F S+V
Sbjct: 312 GELFSSKTKVLQRLIMGAMIQSLQQLTGDNYFFYYGTTIFKAVGL---SDSFETSIVLGI 368
Query: 292 LNAVGTIVSMVLIDRFGRRKLML---ISLIGIFVSLVTLSVT 330
+N T V + +++R+GRR +L S+ V ++ VT
Sbjct: 369 VNFASTFVGIYVVERYGRRTCLLWGAASMTACMVVYASVGVT 410
>HXT6_YEAST (P39003) High-affinity hexose transporter HXT6
Length = 570
Score = 105 bits (261), Expect = 3e-22
Identities = 90/342 (26%), Positives = 156/342 (45%), Gaps = 51/342 (14%)
Query: 34 GGLLFGYDTGVISG----ASLYIRDEFEQVDKKPWLQET----IVSMASAGAIIGAAFGG 85
GG +FG+DTG ISG R + D +L + IVS+ + G IG
Sbjct: 75 GGFVFGWDTGTISGFINQTDFIRRFGMKHKDGTNYLSKVRTGLIVSIFNIGCAIGGIILS 134
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYISE 144
+ D GRK +++ V+++ G ++ A+ W IGR++ GLGVG ++ +P+ ISE
Sbjct: 135 KLGDMYGRKVGLIVVVVIYIIGIIIQIASINKWYQYFIGRIISGLGVGGIAVLSPMLISE 194
Query: 145 ASPAKIRGALVCRLHQWFTHN------------------RWSISL-LPHQFGIYQVYRQS 185
SP +RG LV T +W + L L + ++ + +
Sbjct: 195 VSPKHLRGTLVSCYQLMITAGIFLGYCTNFGTKNYSNSVQWRVPLGLCFAWALFMIGGMT 254
Query: 186 KEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAE-------DGLIGHSLAQKLKG--AW 236
E+ + L+++ G++EE +++ S + + + ++ A+KL G +W
Sbjct: 255 FVPESPRYLAEV---GKIEEAKRSIAVSNKVAVDDPSVLAEVEAVLAGVEAEKLAGNASW 311
Query: 237 -----SNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSG 291
S V + L G +Q +QQ+ G N YY TI + G+ S +F S+V
Sbjct: 312 GELFSSKTKVLQRLIMGAMIQSLQQLTGDNYFFYYGTTIFKAVGL---SDSFETSIVLGI 368
Query: 292 LNAVGTIVSMVLIDRFGRRKLML---ISLIGIFVSLVTLSVT 330
+N T V + +++R+GRR +L S+ V ++ VT
Sbjct: 369 VNFASTFVGIYVVERYGRRTCLLWGAASMTACMVVYASVGVT 410
>HXT4_YEAST (P32467) Low-affinity glucose transporter HXT4
(Low-affinity glucose transporter LGT1)
Length = 576
Score = 104 bits (259), Expect = 5e-22
Identities = 93/342 (27%), Positives = 160/342 (46%), Gaps = 51/342 (14%)
Query: 34 GGLLFGYDTGVISG----ASLYIRDEFEQVDKKPWLQET----IVSMASAGAIIGAAFGG 85
GG +FG+DTG ISG R + D +L + IVS+ + G IG
Sbjct: 81 GGFVFGWDTGTISGFVAQTDFIRRFGMKHHDGTYYLSKVRTGLIVSIFNIGCAIGGIILA 140
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYISE 144
+ D GRK +++ V+++ G ++ A+ W IGR++ GLGVG ++ +P+ ISE
Sbjct: 141 KLGDMYGRKMGLIVVVVIYIIGIIIQIASINKWYQYFIGRIISGLGVGGIAVLSPMLISE 200
Query: 145 ASPAKIRGALV---------------CRLHQWFTHN---RWSISL-LPHQFGIYQVYRQS 185
SP IRG LV C + T+ +W + L L + ++ + +
Sbjct: 201 VSPKHIRGTLVSCYQLMITLGIFLGYCTNYGTKTYTNSVQWRVPLGLGFAWALFMIGGMT 260
Query: 186 KEEEAKQILSKIYRPGEVEEEMK--AMHESIEAE----KAEDGLIGHSL-AQKLKG--AW 236
E+ + L ++ G++EE + A+ + A+ AE ++ ++ A+KL G +W
Sbjct: 261 FVPESPRYLVEV---GKIEEAKRSIALSNKVSADDPAVMAEVEVVQATVEAEKLAGNASW 317
Query: 237 -----SNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSG 291
+ V + L G +Q +QQ+ G N YY T+ G+ +F S+V
Sbjct: 318 GEIFSTKTKVFQRLIMGAMIQSLQQLTGDNYFFYYGTTVFTAVGL---EDSFETSIVLGI 374
Query: 292 LNAVGTIVSMVLIDRFGRRKLML---ISLIGIFVSLVTLSVT 330
+N T V + L++R+GRR+ +L S+ V ++ VT
Sbjct: 375 VNFASTFVGIFLVERYGRRRCLLWGAASMTACMVVFASVGVT 416
>GTR8_HUMAN (Q9NY64) Solute carrier family 2, facilitated glucose
transporter, member 8 (Glucose transporter type 8)
(Glucose transporter type X1)
Length = 477
Score = 103 bits (257), Expect = 8e-22
Identities = 89/304 (29%), Positives = 141/304 (46%), Gaps = 29/304 (9%)
Query: 31 ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASA----GAIIGAAFGGY 86
A G L FG+ G S A ++ P L + S A GA G GG+
Sbjct: 33 AALGPLSFGFALGYSSPAIPSLQ---RAAPPAPRLDDAAASWFGAVVTLGAAAGGVLGGW 89
Query: 87 MNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEAS 146
+ D+ GRK ++L+ V FVAG V+ AA W+++ GRLL GL G AS+ AP+YISE +
Sbjct: 90 LVDRAGRKLSLLLCSVPFVAGFAVITAAQDVWMLLGGRLLTGLACGVASLVAPVYISEIA 149
Query: 147 PAKIRGAL-------------VCRLHQWFTHNRWSISLLPHQFGIYQVYRQSKEEEAKQI 193
+RG L + L W RW L + + E + +
Sbjct: 150 YPAVRGLLGSCVQLMVVVGILLAYLAGWVLEWRWLAVLGCVPPSLMLLLMCFMPETPRFL 209
Query: 194 LSKIYRPGEVEEEMKAMHESIEAEKA-EDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQ 252
L++ R +E M A+ +E+ ED IG + L + + G+++
Sbjct: 210 LTQHRR----QEAMAALRFLWGSEQGWEDPPIGAEQSFHL-ALLRQPGIYKPFIIGVSLM 264
Query: 253 VVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKL 312
QQ+ G+N +M+Y+ TI + A +S A S+V + + T V+ +++DR GRR L
Sbjct: 265 AFQQLSGVNAVMFYAETIFEEAKFKDSSLA---SVVVGVIQVLFTAVAALIMDRAGRRLL 321
Query: 313 MLIS 316
+++S
Sbjct: 322 LVLS 325
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,493,100
Number of Sequences: 164201
Number of extensions: 1816720
Number of successful extensions: 6271
Number of sequences better than 10.0: 257
Number of HSP's better than 10.0 without gapping: 200
Number of HSP's successfully gapped in prelim test: 57
Number of HSP's that attempted gapping in prelim test: 5610
Number of HSP's gapped (non-prelim): 480
length of query: 420
length of database: 59,974,054
effective HSP length: 113
effective length of query: 307
effective length of database: 41,419,341
effective search space: 12715737687
effective search space used: 12715737687
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148526.6


