

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148526.1 + phase: 0
(598 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
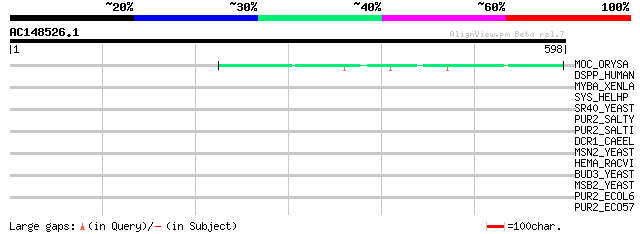
Score E
Sequences producing significant alignments: (bits) Value
MOC_ORYSA (Q84MM9) Protein MONOCULM 1 111 6e-24
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 37 0.15
MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB) (MYB-r... 37 0.19
SYS_HELHP (Q7VG34) Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--... 34 0.95
SR40_YEAST (P32583) Suppressor protein SRP40 33 1.6
PUR2_SALTY (P26977) Phosphoribosylamine--glycine ligase (EC 6.3.... 33 1.6
PUR2_SALTI (Q8Z334) Phosphoribosylamine--glycine ligase (EC 6.3.... 33 1.6
DCR1_CAEEL (P34529) Endoribonuclease dcr-1 (EC 3.1.26.-) 33 2.1
MSN2_YEAST (P33748) Zinc finger protein MSN2 (Multicopy suppress... 32 3.6
HEMA_RACVI (Q00716) Hemagglutinin precursor 32 3.6
BUD3_YEAST (P25558) Bud site selection protein BUD3 32 3.6
MSB2_YEAST (P32334) MSB2 protein (Multicopy suppressor of bud em... 32 4.7
PUR2_ECOL6 (Q8FB69) Phosphoribosylamine--glycine ligase (EC 6.3.... 31 8.1
PUR2_ECO57 (Q8X612) Phosphoribosylamine--glycine ligase (EC 6.3.... 31 8.1
>MOC_ORYSA (Q84MM9) Protein MONOCULM 1
Length = 441
Score = 111 bits (277), Expect = 6e-24
Identities = 96/404 (23%), Positives = 163/404 (39%), Gaps = 50/404 (12%)
Query: 226 KEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAI 285
++ L+ CA + GD A VL S G RL + L RV++
Sbjct: 51 RDLLLACADLLQRGDLPAARRAAEIVLAAAASPRGDAADRLAYHFARALALRVDAKAGHG 110
Query: 286 YKALK--CEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQ 343
+ + P S A + QI P+ +FA++++N I E + R+HI+D
Sbjct: 111 HVVVGGGAARPASSGAYLAFN---QIAPFLRFAHLTANQAILEAVDGARRVHILDLDAVH 167
Query: 344 GSQWMLLLHALKHKPG---GPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPF 400
G QW LL A+ + GPP +RVTG + R G +L A++ +PF
Sbjct: 168 GVQWPPLLQAIAERADPALGPPEVRVTGAGADRDTLLR------TGNRLRAFARSIHLPF 221
Query: 401 EFNSVKMY--------------------GCEVQLEDFEVQHDEVLVVNFPFALHHIPDES 440
F + + E DE L VN LH++
Sbjct: 222 HFTPLLLSCATTAPHHVAGTSTGAAAAASTAAAATGLEFHPDETLAVNCVMFLHNLAGH- 280
Query: 441 VSMENHRDRLLRLVKILSPKVVLFVEQESN------TNTSPFLPRFAETLNYYTAMFESI 494
+ L+ VK +SP VV E+E+ + R +++Y+A+FE++
Sbjct: 281 ----DELAAFLKWVKAMSPAVVTIAEREAGGGGGGGDHIDDLPRRVGVAMDHYSAVFEAL 336
Query: 495 DVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSP 554
+ +P ++R+ EQ + R+I + G + E +W AGF LS
Sbjct: 337 EATVPPGSRERLAVEQEVLGREIEAAVGPSGGRWWRGIE---RWGGAARAAGFAARPLSA 393
Query: 555 SVIDSVRTLLKDF--NKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
+ R LL+ ++ Y +++ A L W+++ + + SAW+
Sbjct: 394 FAVSQARLLLRLHYPSEGYLVQEARGACFLGWQTRPLLSVSAWQ 437
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 37.0 bits (84), Expect = 0.15
Identities = 44/191 (23%), Positives = 68/191 (35%), Gaps = 7/191 (3%)
Query: 37 TNSNNGGRRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPSSASGDPFNMRST 96
+NS++ ++ S E+SN N + S + + S SS SGD N +
Sbjct: 912 SNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDS 971
Query: 97 FSPQISRYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGS 156
S S N+++ S S + D S + S +D SNS+ S
Sbjct: 972 SDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSD-------SSNSSDS 1024
Query: 157 LHQGTSQDNEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYDWSQF 216
+ S D+ D S S+ S SSD DS S+ + + D S
Sbjct: 1025 SNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSGSSDSSDSSDSSDSSDSSDS 1084
Query: 217 EEIIPKLDLKE 227
+ D E
Sbjct: 1085 SDSSDSSDSSE 1095
Score = 32.0 bits (71), Expect = 4.7
Identities = 39/174 (22%), Positives = 69/174 (39%), Gaps = 8/174 (4%)
Query: 26 DSNKNRFKTLQTNSNNGGRRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPSS 85
DS+ ++ + ++ S++ +++ S S+ S+ + SS ++ + DS S
Sbjct: 618 DSSDSKSDSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSDS 677
Query: 86 ASGDPFNMRSTFSPQISRYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEV 145
+S + S S SSD++ + S+D DS + S ++
Sbjct: 678 SSSSDSSSSSDSSNSSDSSDSSDSSNSSESSDSSDSSDSDSSD--------SSDSSNSNS 729
Query: 146 FGSCLSNSNGSLHQGTSQDNEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSN 199
S SNS+ S S D+ D S S+ S SSD DS S+
Sbjct: 730 SDSDSSNSSDSSDSSDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSS 783
Score = 31.6 bits (70), Expect = 6.2
Identities = 44/175 (25%), Positives = 74/175 (42%), Gaps = 12/175 (6%)
Query: 26 DSNKNRFKTLQTNSNNGGRRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFD-SPS 84
DS+ + + ++S+N G ++ S ++++ +S+ +SS ++ + D S S
Sbjct: 946 DSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSS--------DSSNSSDSSDSSDSSDS 997
Query: 85 SASGDPFNMRSTFSPQISRYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIE 144
S S D N S+ S S S N+++ S S + D S + S +D
Sbjct: 998 SDSSDSSN--SSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSD-S 1054
Query: 145 VFGSCLSNSNGSLHQGTSQDNEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSN 199
S S+S+GS S D+ D S S+ S SSD DS S+
Sbjct: 1055 SDSSDSSDSSGSSDSSDSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSSDSS 1109
>MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB)
(MYB-related protein 2) (XMYB2)
Length = 728
Score = 36.6 bits (83), Expect = 0.19
Identities = 46/182 (25%), Positives = 78/182 (42%), Gaps = 40/182 (21%)
Query: 47 RTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPSS------ASGDPFNMRSTF--- 97
RT+ S++ ++W S +++ +E D++ D PS A+ + ++ F
Sbjct: 170 RTDNSIK-NHWNSTMKRK---VEQEGYLQDLMNCDRPSKLQAKSCAAPNHLQAQNQFYIP 225
Query: 98 -SPQISRYCSSDNTTNVSPLG--------VYSSADDDSFEFKMKEIELSLVGTDIEV--- 145
QI RY S + N + + ADD E ++KE+EL L+ + EV
Sbjct: 226 VQTQIPRYSSLSHDDNCIEIQNSFSFIQQPFVDADDPEKEKRIKELELLLMSAENEVRRK 285
Query: 146 ---FGSCLSNSNGSLHQGTSQDNE-----------FHIDEMISEQGFKESEISLLGPSSD 191
GS L+ S S H G S N + +DE+ + G + E +L P ++
Sbjct: 286 RVPSGSSLTWSE-SYHMGESMSNTMSHLEEQTHDFYSLDEIPNTSGQQSVEDKILAPEAN 344
Query: 192 IV 193
IV
Sbjct: 345 IV 346
>SYS_HELHP (Q7VG34) Seryl-tRNA synthetase (EC 6.1.1.11)
(Serine--tRNA ligase) (SerRS)
Length = 422
Score = 34.3 bits (77), Expect = 0.95
Identities = 27/100 (27%), Positives = 46/100 (46%), Gaps = 5/100 (5%)
Query: 369 IDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVN 428
+++ Q+F + KL K+ + KV + N VKM E L++ E H E L
Sbjct: 45 LEELQAFQNKTSKLFGEYKREQKDITALKVALDENKVKMSTLESTLKEIE-SHLETLAYG 103
Query: 429 FPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQE 468
P ++PD++ + + + K+LSP FV +E
Sbjct: 104 IP----NLPDDATPEGEDENDNVEIKKVLSPPHFDFVPKE 139
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 33.5 bits (75), Expect = 1.6
Identities = 45/193 (23%), Positives = 79/193 (40%), Gaps = 24/193 (12%)
Query: 37 TNSNNGGRRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPSSASGDPFNMRST 96
++S++ ++ S +S+ ES+ + SS ++D + DS SS+S + S+
Sbjct: 30 SSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSSDSSD--SSDSESSSSSSSSSSSSS 87
Query: 97 FSPQISRYCSSDNTTNVSPLGVYSSADDDSFEFKMKE--------------IELSLVGTD 142
S SD++++ S SS+D+ S E + ++ E T+
Sbjct: 88 SSSDSESSSESDSSSSGSSSSSSSSSDESSSESESEDETKKRARESDNEDAKETKKAKTE 147
Query: 143 IEVFGSCLSNSNGSLHQGTSQD-NEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLN 201
E S S+S+GS S+ +E D S +SE SD QS+ +
Sbjct: 148 PESSSSSESSSSGSSSSSESESGSESDSDSSSSSSSSSDSE-------SDSESDSQSSSS 200
Query: 202 GSLHQGTSQYDWS 214
S +S D S
Sbjct: 201 SSSSDSSSDSDSS 213
>PUR2_SALTY (P26977) Phosphoribosylamine--glycine ligase (EC
6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase)
(Phosphoribosylglycinamide synthetase)
Length = 429
Score = 33.5 bits (75), Expect = 1.6
Identities = 27/112 (24%), Positives = 51/112 (45%), Gaps = 3/112 (2%)
Query: 103 RYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTS 162
R + D N +G YS A + E + +E ++ ++ + + G L+ G
Sbjct: 217 RVGNGDTGPNTGGMGAYSPAPVVTDEVHQRTME-RIIWPTVKGMAAEGNTYTGFLYAGLM 275
Query: 163 QDNEFH--IDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYD 212
D + + + E G E++ +L SD+VD C + +G L + TS++D
Sbjct: 276 IDKQGNPKVIEFNCRFGDPETQPIMLRMKSDLVDLCLAACDGKLDEKTSEWD 327
>PUR2_SALTI (Q8Z334) Phosphoribosylamine--glycine ligase (EC
6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase)
(Phosphoribosylglycinamide synthetase)
Length = 429
Score = 33.5 bits (75), Expect = 1.6
Identities = 27/112 (24%), Positives = 51/112 (45%), Gaps = 3/112 (2%)
Query: 103 RYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTS 162
R + D N +G YS A + E + +E ++ ++ + + G L+ G
Sbjct: 217 RVGNGDTGPNTGGMGAYSPAPVVTDEVHQRTME-RIIWPTVKGMAAEGNTYTGFLYAGLM 275
Query: 163 QDNEFH--IDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYD 212
D + + + E G E++ +L SD+VD C + +G L + TS++D
Sbjct: 276 IDKQGNPKVIEFNCRFGDPETQPIMLRMKSDLVDLCLAACDGKLDEKTSEWD 327
>DCR1_CAEEL (P34529) Endoribonuclease dcr-1 (EC 3.1.26.-)
Length = 1845
Score = 33.1 bits (74), Expect = 2.1
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 13 IQLYYQQQSLRQIDSNKNRFKTLQTNSNNGGRRQRTNVSLETSN 56
+++Y Q QSL +D R LQ N RR RT + TSN
Sbjct: 861 LEIYDQNQSLLDVDFTSTRLNLLQPRIQNQPRRSRTVSNSSTSN 904
>MSN2_YEAST (P33748) Zinc finger protein MSN2 (Multicopy suppressor
of SNF1 protein 2)
Length = 704
Score = 32.3 bits (72), Expect = 3.6
Identities = 44/180 (24%), Positives = 70/180 (38%), Gaps = 40/180 (22%)
Query: 26 DSNKNRFKTLQTNSNNGGRRQRTNVSLETSNWESNWEKQFCTLESSQVANDVI-----AF 80
DSNKNR +T N+ + N S +N Q+ TL S ND++
Sbjct: 96 DSNKNR--DARTIENDSEIKSTNNASGSGAN-------QYTTLTSPYPMNDILYNMNNPL 146
Query: 81 DSPSSAS-------GDPFNMRSTFSPQISRYCSSDNTTNVSPL----------------G 117
SPS +S P N S +S S+ N T +SP G
Sbjct: 147 QSPSPSSVPQNPTINPPINTASN-ETNLSPQTSNGNETLISPRAQQHTSIKDNRLSLPNG 205
Query: 118 VYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSL--HQGTSQDNEFHIDEMISE 175
S+ D+ + E + + +D + + +SNSN + + +S N +ID M+ +
Sbjct: 206 ANSNLFIDTNPNNLNEKLRNQLNSDTNSYSNSISNSNSNSTGNLNSSYFNSLNIDSMLDD 265
>HEMA_RACVI (Q00716) Hemagglutinin precursor
Length = 310
Score = 32.3 bits (72), Expect = 3.6
Identities = 37/155 (23%), Positives = 65/155 (41%), Gaps = 18/155 (11%)
Query: 59 SNWEKQFCTLESSQVANDVIAFDSPSS---ASGDPFNMRSTFSPQISRYCSSDNTTNVSP 115
S+W K+ ++ NDV+ FD ++ + P++ +T I S+D T +
Sbjct: 48 SSWYKKPDSIILLAAKNDVVYFDDYTADKVSYDSPYDTLATIIT-IKSLTSADAGTYICA 106
Query: 116 LGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTSQDNEFHIDEM--- 172
+ S+ D D +++ I+L + ++ + LS S T QD H +E
Sbjct: 107 FFITSTNDTDKIDYEEYFIDLVVNPANVSTIDAILSGST------TQQDIISHTEEQHDS 160
Query: 173 -----ISEQGFKESEISLLGPSSDIVDSCQSNLNG 202
SE + SE S SS I ++ +S G
Sbjct: 161 DTTICTSESTTQISETSESTTSSQISETSESTSYG 195
>BUD3_YEAST (P25558) Bud site selection protein BUD3
Length = 1636
Score = 32.3 bits (72), Expect = 3.6
Identities = 33/142 (23%), Positives = 59/142 (41%), Gaps = 11/142 (7%)
Query: 64 QFCTLESSQVANDVIAFDS--PSSASGDPFNMRSTFSPQISRYCSSDNTTNVSPLGVYSS 121
+ C +S ++ + F + PS + N+ F +R+CS+D N+ L V +
Sbjct: 633 ELCAYDSEKIKSKFALFLNIPPSKEILEVNNLHLAF---FARFCSNDGRDNIVILDVLTK 689
Query: 122 ADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTSQDNEFHIDEMISEQGFKES 181
DD E I +++ C S+ N S+ + NE I + E +E
Sbjct: 690 HDDKHIEVTSDNIVFTIINQLAIEIPICFSSLNSSMAKDLLCVNENLIKNL--EHQLEE- 746
Query: 182 EISLLGPSSDIVDSCQSNLNGS 203
+ PS+D + S L+G+
Sbjct: 747 ---VKHPSTDEHRAVNSKLSGA 765
>MSB2_YEAST (P32334) MSB2 protein (Multicopy suppressor of bud
emergence 2)
Length = 1306
Score = 32.0 bits (71), Expect = 4.7
Identities = 27/102 (26%), Positives = 46/102 (44%), Gaps = 5/102 (4%)
Query: 111 TNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLH---QGTSQDNEF 167
TN++ L + DDS + + E+ +IE + ++NS+ + + GT++
Sbjct: 1048 TNITVLQIVP-LQDDSLNYLVSVAEVYFPTAEIEELSNLITNSSSAFYTDGMGTAKSMAA 1106
Query: 168 HIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTS 209
+D I G S G SSD S SN +GS G++
Sbjct: 1107 MVDSSIPLTGLLHDSNSNSGGSSDGSSSSNSN-SGSSGSGSN 1147
>PUR2_ECOL6 (Q8FB69) Phosphoribosylamine--glycine ligase (EC
6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase)
(Phosphoribosylglycinamide synthetase)
Length = 429
Score = 31.2 bits (69), Expect = 8.1
Identities = 26/112 (23%), Positives = 49/112 (43%), Gaps = 3/112 (2%)
Query: 103 RYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTS 162
R D N +G YS A + E + +E ++ ++ + + G L+ G
Sbjct: 217 RVGDKDTGPNTGGMGAYSPAPVVTDEVHQRTME-RIIWPTVKGMAAEGNTYTGFLYAGLM 275
Query: 163 QDNEFH--IDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYD 212
D + + + E G E++ +L SD+V+ C + G L + TS++D
Sbjct: 276 IDKQGNPKVIEFNCRFGDPETQPIMLRMKSDLVELCLAACEGKLDEKTSEWD 327
>PUR2_ECO57 (Q8X612) Phosphoribosylamine--glycine ligase (EC
6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase)
(Phosphoribosylglycinamide synthetase)
Length = 429
Score = 31.2 bits (69), Expect = 8.1
Identities = 26/112 (23%), Positives = 49/112 (43%), Gaps = 3/112 (2%)
Query: 103 RYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTS 162
R D N +G YS A + + + +E ++ ++ S + G L+ G
Sbjct: 217 RVGDKDTGPNTGGMGAYSPAPVVTDDVHQRTME-RIIWPTVKGMASEGNTYTGFLYAGLM 275
Query: 163 QDNEFH--IDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYD 212
D + + + E G E++ +L SD+V+ C + G L + TS++D
Sbjct: 276 IDKQGNPKVIEFNCRFGDPETQPIMLRMKSDLVELCLAACEGKLDEKTSEWD 327
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,872,468
Number of Sequences: 164201
Number of extensions: 3030307
Number of successful extensions: 7094
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 7060
Number of HSP's gapped (non-prelim): 28
length of query: 598
length of database: 59,974,054
effective HSP length: 116
effective length of query: 482
effective length of database: 40,926,738
effective search space: 19726687716
effective search space used: 19726687716
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC148526.1


