

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.6 + phase: 0
(125 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
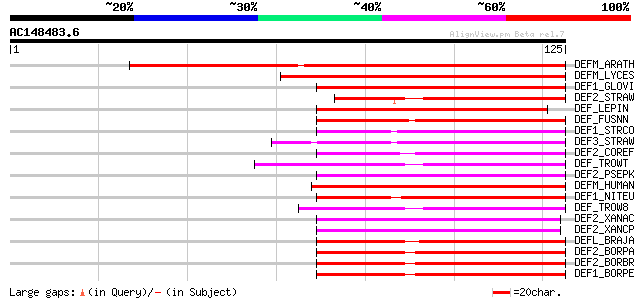
Score E
Sequences producing significant alignments: (bits) Value
DEFM_ARATH (Q9FV53) Peptide deformylase, mitochondrial precursor... 90 1e-18
DEFM_LYCES (Q9FUZ0) Peptide deformylase, mitochondrial precursor... 84 6e-17
DEF1_GLOVI (Q7NJV3) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1) ... 57 6e-09
DEF2_STRAW (Q826Q0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 50 9e-07
DEF_LEPIN (Q93LE9) Peptide deformylase (EC 3.5.1.88) (PDF) (Poly... 50 1e-06
DEF_FUSNN (Q8REF0) Peptide deformylase (EC 3.5.1.88) (PDF) (Poly... 50 1e-06
DEF1_STRCO (Q9RD27) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1) ... 48 3e-06
DEF3_STRAW (Q825U9) Peptide deformylase 3 (EC 3.5.1.88) (PDF 3) ... 47 8e-06
DEF2_COREF (Q8FMD0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 47 1e-05
DEF_TROWT (Q83GH8) Peptide deformylase (EC 3.5.1.88) (PDF) (Poly... 45 2e-05
DEF2_PSEPK (Q88EA7) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 45 2e-05
DEFM_HUMAN (Q9HBH1) Peptide deformylase, mitochondrial precursor... 45 3e-05
DEF1_NITEU (Q82TW4) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1) ... 45 3e-05
DEF_TROW8 (Q83HQ3) Peptide deformylase (EC 3.5.1.88) (PDF) (Poly... 45 4e-05
DEF2_XANAC (Q8PG20) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 44 5e-05
DEF2_XANCP (Q8P4F9) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 44 6e-05
DEFL_BRAJA (Q89N37) Peptide deformylase-like (Polypeptide deform... 44 8e-05
DEF2_BORPA (Q7W4K0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 43 1e-04
DEF2_BORBR (Q7WG25) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2) ... 43 1e-04
DEF1_BORPE (Q7W0Q0) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1) ... 43 1e-04
>DEFM_ARATH (Q9FV53) Peptide deformylase, mitochondrial precursor
(EC 3.5.1.88) (PDF) (Polypeptide deformylase)
Length = 259
Score = 89.7 bits (221), Expect = 1e-18
Identities = 53/98 (54%), Positives = 64/98 (65%), Gaps = 1/98 (1%)
Query: 28 SSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDH 87
SS N K+ T S ST + +K V+ LP IV +GDPV+HE AREVD
Sbjct: 32 SSHLLNRKLYNLPTSSSSSLSTKAGWLLGLGEKKKKVD-LPEIVASGDPVLHEKAREVDP 90
Query: 88 SEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
EI S++IQ IIDDMI VMR APGVG+AAPQIG+PLR+
Sbjct: 91 GEIGSERIQKIIDDMIKVMRLAPGVGLAAPQIGVPLRI 128
>DEFM_LYCES (Q9FUZ0) Peptide deformylase, mitochondrial precursor
(EC 3.5.1.88) (PDF) (Polypeptide deformylase)
Length = 277
Score = 84.0 bits (206), Expect = 6e-17
Identities = 38/64 (59%), Positives = 52/64 (80%)
Query: 62 KTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGI 121
K +P IVKAGDPV+HEP++++ EI S++IQ II++M+ VMR APGVG+AAPQIGI
Sbjct: 83 KKKQAMPDIVKAGDPVLHEPSQDIPLEEIGSERIQKIIEEMVKVMRNAPGVGLAAPQIGI 142
Query: 122 PLRV 125
PL++
Sbjct: 143 PLKI 146
>DEF1_GLOVI (Q7NJV3) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1)
(Polypeptide deformylase 1)
Length = 227
Score = 57.4 bits (137), Expect = 6e-09
Identities = 27/56 (48%), Positives = 41/56 (73%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IVK GDPV+ A+ ++ EI+S+ IQ +I M MR+APGVG+AAPQ+G+ +++
Sbjct: 48 IVKTGDPVLRLTAKPLNSDEIQSEAIQQLIAAMAERMREAPGVGLAAPQVGVSVQL 103
>DEF2_STRAW (Q826Q0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 186
Score = 50.1 bits (118), Expect = 9e-07
Identities = 26/53 (49%), Positives = 34/53 (64%), Gaps = 5/53 (9%)
Query: 74 GDPVIHEPAREV-DHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
GDPV+H P EV DH ++ +++DM M A GVG+AA QIG+PLRV
Sbjct: 20 GDPVLHAPCEEVTDHGP----ELARLVEDMFATMYAANGVGLAANQIGVPLRV 68
>DEF_LEPIN (Q93LE9) Peptide deformylase (EC 3.5.1.88) (PDF)
(Polypeptide deformylase)
Length = 178
Score = 49.7 bits (117), Expect = 1e-06
Identities = 23/52 (44%), Positives = 35/52 (67%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGI 121
I++ GDP++ + + V EI++ + + +I DM MR A GVG+AAPQIGI
Sbjct: 6 ILRMGDPILRKISEPVTEDEIQTKEFKKLIRDMFDTMRHAEGVGLAAPQIGI 57
>DEF_FUSNN (Q8REF0) Peptide deformylase (EC 3.5.1.88) (PDF)
(Polypeptide deformylase)
Length = 174
Score = 49.7 bits (117), Expect = 1e-06
Identities = 25/56 (44%), Positives = 39/56 (69%), Gaps = 1/56 (1%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
I K G+ V+ + A+EV+ SEI +D+ + +DDM+ M + GVG+AAPQIG+ R+
Sbjct: 5 IKKYGEDVLKQIAKEVELSEI-NDEFRQFLDDMVETMYETDGVGLAAPQIGVSKRI 59
>DEF1_STRCO (Q9RD27) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1)
(Polypeptide deformylase 1)
Length = 218
Score = 48.1 bits (113), Expect = 3e-06
Identities = 24/56 (42%), Positives = 34/56 (59%), Gaps = 1/56 (1%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IV AGDPV+ A D ++ + ++ + L M APGVG+AAPQ+G+ LRV
Sbjct: 26 IVAAGDPVLRRAAEPYD-GQVAPALFERFVEALRLTMHAAPGVGLAAPQVGVGLRV 80
>DEF3_STRAW (Q825U9) Peptide deformylase 3 (EC 3.5.1.88) (PDF 3)
(Polypeptide deformylase 3)
Length = 224
Score = 47.0 bits (110), Expect = 8e-06
Identities = 27/66 (40%), Positives = 38/66 (56%), Gaps = 2/66 (3%)
Query: 60 LRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQI 119
L T LP IV AGDPV+ A D ++ + ++ + L M APGVG+AAPQ+
Sbjct: 26 LLATGGPLP-IVAAGDPVLRRGAEPYD-GQLGPGLLARFVEALRLTMHAAPGVGLAAPQV 83
Query: 120 GIPLRV 125
G+ LR+
Sbjct: 84 GVGLRI 89
>DEF2_COREF (Q8FMD0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 193
Score = 46.6 bits (109), Expect = 1e-05
Identities = 26/56 (46%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IV GDPV+H P REV ++Q +I DM M A GVG+AA QIG+ R+
Sbjct: 6 IVIHGDPVLHNPTREVTEP---ISELQELIADMYETMEVANGVGLAANQIGVSKRI 58
>DEF_TROWT (Q83GH8) Peptide deformylase (EC 3.5.1.88) (PDF)
(Polypeptide deformylase)
Length = 228
Score = 45.4 bits (106), Expect = 2e-05
Identities = 28/70 (40%), Positives = 38/70 (54%), Gaps = 4/70 (5%)
Query: 56 ETALLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVA 115
E A+ + + K+ I G V+H PA+ V IQ I+ DM M APGVG+A
Sbjct: 25 EGAVPKISGGKILPIYITGHAVLHAPAKPVTDFS----GIQEIVRDMFATMFAAPGVGLA 80
Query: 116 APQIGIPLRV 125
PQIG+ LR+
Sbjct: 81 GPQIGLGLRI 90
>DEF2_PSEPK (Q88EA7) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 178
Score = 45.4 bits (106), Expect = 2e-05
Identities = 24/56 (42%), Positives = 34/56 (59%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
I+K GD + A V + S ++Q +IDDM MR GVG+AAPQ+GI L++
Sbjct: 5 ILKMGDERLLRIAPPVPEHMLGSAELQQLIDDMFETMRHVGGVGLAAPQVGIDLQL 60
>DEFM_HUMAN (Q9HBH1) Peptide deformylase, mitochondrial precursor
(EC 3.5.1.88) (PDF) (Polypeptide deformylase)
Length = 243
Score = 45.1 bits (105), Expect = 3e-05
Identities = 18/57 (31%), Positives = 37/57 (64%)
Query: 69 YIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
++ + GDPV+ A V+ +++ ++Q + ++ VMR+ VG++APQ+G+P +V
Sbjct: 66 HVCQVGDPVLRGVAAPVERAQLGGPELQRLTQRLVQVMRRRRCVGLSAPQLGVPRQV 122
>DEF1_NITEU (Q82TW4) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1)
(Polypeptide deformylase 1)
Length = 176
Score = 45.1 bits (105), Expect = 3e-05
Identities = 22/56 (39%), Positives = 36/56 (64%), Gaps = 2/56 (3%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
++K GDP + +PAR VD + + +++ ++ DM M G G+AAPQIG+ L+V
Sbjct: 5 VLKMGDPCLLQPARRVD--QFGTPELEALLQDMQDTMAALNGAGLAAPQIGVSLQV 58
>DEF_TROW8 (Q83HQ3) Peptide deformylase (EC 3.5.1.88) (PDF)
(Polypeptide deformylase)
Length = 201
Score = 44.7 bits (104), Expect = 4e-05
Identities = 26/60 (43%), Positives = 33/60 (54%), Gaps = 4/60 (6%)
Query: 66 KLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
K+ I G V+H PA+ V IQ I+ DM M APGVG+A PQIG+ LR+
Sbjct: 8 KILPIYITGHAVLHAPAKPVTDFS----GIQEIVRDMFATMFAAPGVGLAGPQIGLGLRI 63
>DEF2_XANAC (Q8PG20) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 170
Score = 44.3 bits (103), Expect = 5e-05
Identities = 21/55 (38%), Positives = 33/55 (59%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLR 124
I++ DP + A VD +E+ S Q ++DDM M +APG+G+AA Q+ + R
Sbjct: 6 ILEFPDPRLRTKAVPVDAAEVTSQAFQTLLDDMFHTMYEAPGIGLAASQVDVHKR 60
>DEF2_XANCP (Q8P4F9) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 170
Score = 43.9 bits (102), Expect = 6e-05
Identities = 21/55 (38%), Positives = 33/55 (59%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLR 124
I++ DP + A VD +E+ S Q ++DDM M +APG+G+AA Q+ + R
Sbjct: 6 ILEFPDPRLRTKAVPVDAAEVVSPAFQTLLDDMFQTMYEAPGIGLAASQVDVHKR 60
>DEFL_BRAJA (Q89N37) Peptide deformylase-like (Polypeptide
deformylase-like)
Length = 170
Score = 43.5 bits (101), Expect = 8e-05
Identities = 22/56 (39%), Positives = 35/56 (62%), Gaps = 3/56 (5%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
+V+ D + PAR V + D+++ + D++ MR APG+G+ AP IG+PLRV
Sbjct: 11 LVRYPDRRLAIPARPVTAFD---DELRELAADLLDTMRAAPGIGITAPHIGVPLRV 63
>DEF2_BORPA (Q7W4K0) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 176
Score = 43.1 bits (100), Expect = 1e-04
Identities = 23/56 (41%), Positives = 36/56 (64%), Gaps = 2/56 (3%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
I+K GDP + A V+ + + +++ +IDDM M A GVG+AAPQIG+ L++
Sbjct: 5 ILKMGDPRLLRVAAPVERYD--TPELRALIDDMFETMAHAQGVGLAAPQIGVDLQL 58
>DEF2_BORBR (Q7WG25) Peptide deformylase 2 (EC 3.5.1.88) (PDF 2)
(Polypeptide deformylase 2)
Length = 176
Score = 43.1 bits (100), Expect = 1e-04
Identities = 23/56 (41%), Positives = 36/56 (64%), Gaps = 2/56 (3%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
I+K GDP + A V+ + + +++ +IDDM M A GVG+AAPQIG+ L++
Sbjct: 5 ILKMGDPRLLRVAAPVERYD--TPELRALIDDMFETMAHAQGVGLAAPQIGVDLQL 58
>DEF1_BORPE (Q7W0Q0) Peptide deformylase 1 (EC 3.5.1.88) (PDF 1)
(Polypeptide deformylase 1)
Length = 176
Score = 43.1 bits (100), Expect = 1e-04
Identities = 23/56 (41%), Positives = 36/56 (64%), Gaps = 2/56 (3%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
I+K GDP + A V+ + + +++ +IDDM M A GVG+AAPQIG+ L++
Sbjct: 5 ILKMGDPRLLRVAAPVERYD--TPELRALIDDMFETMAHAQGVGLAAPQIGVDLQL 58
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,480,905
Number of Sequences: 164201
Number of extensions: 471395
Number of successful extensions: 1874
Number of sequences better than 10.0: 186
Number of HSP's better than 10.0 without gapping: 133
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 1715
Number of HSP's gapped (non-prelim): 190
length of query: 125
length of database: 59,974,054
effective HSP length: 101
effective length of query: 24
effective length of database: 43,389,753
effective search space: 1041354072
effective search space used: 1041354072
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148483.6


