

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
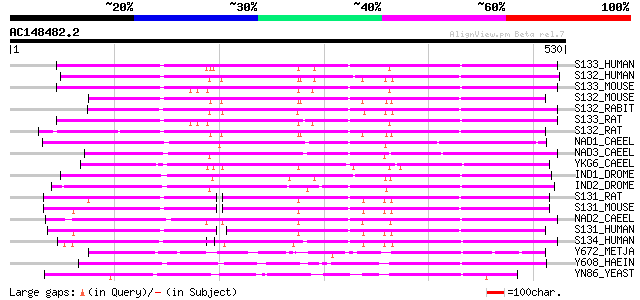
Score E
Sequences producing significant alignments: (bits) Value
S133_HUMAN (Q8WWT9) Solute carrier family 13, member 3 (Sodium-d... 296 9e-80
S132_HUMAN (Q13183) Solute carrier family 13, member 2 (Renal so... 290 6e-78
S133_MOUSE (Q91Y63) Solute carrier family 13, member 3 (Sodium-d... 286 9e-77
S132_MOUSE (Q9ES88) Solute carrier family 13, member 2 (Renal so... 286 1e-76
S132_RABIT (Q28615) Solute carrier family 13, member 2 (Renal so... 285 2e-76
S133_RAT (Q9Z0Z5) Solute carrier family 13, member 3 (Sodium-dep... 284 4e-76
S132_RAT (P70545) Solute carrier family 13, member 2 (Intestinal... 258 2e-68
NAD1_CAEEL (Q93655) Sodium-dependent low-affinity dicarboxylate ... 255 2e-67
NAD3_CAEEL (Q21339) Sodium-dependent high-affinity dicarboxylate... 252 1e-66
YKG6_CAEEL (P46556) Hypothetical protein B0285.6 in chromosome III 224 4e-58
IND1_DROME (Q9VVT2) I'm not dead yet protein (INDY transporter p... 220 6e-57
IND2_DROME (Q9VDQ0) I'm not dead yet protein 2 213 8e-55
S131_RAT (Q07782) Solute carrier family 13, member 1 (Renal sodi... 209 2e-53
S131_MOUSE (Q9JHI4) Solute carrier family 13, member 1 (Renal so... 209 2e-53
NAD2_CAEEL (P32739) Sodium-dependent high-affinity dicarboxylate... 203 8e-52
S131_HUMAN (Q9BZW2) Solute carrier family 13, member 1 (Renal so... 197 6e-50
S134_HUMAN (Q9UKG4) Solute carrier family 13, member 4 (Na(+)/su... 189 2e-47
Y672_METJA (Q58086) Hypothetical protein MJ0672 154 7e-37
Y608_HAEIN (Q57486) Hypothetical protein HI0608 152 2e-36
YN86_YEAST (P27514) Hypothetical 99.5 kDa protein in URK1-SMM1 i... 127 7e-29
>S133_HUMAN (Q8WWT9) Solute carrier family 13, member 3
(Sodium-dependent high-affinity dicarboxylate
transporter 2) (Na(+)/dicarboxylate cotransporter 3)
(NaDC-3) (hNaDC3)
Length = 602
Score = 296 bits (758), Expect = 9e-80
Identities = 183/558 (32%), Positives = 285/558 (50%), Gaps = 84/558 (15%)
Query: 45 LLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADT 104
L + + L V P + L VI + +W T A+PL VT++ P+ LFP GI ++
Sbjct: 21 LFTPLALLPVVFALPPKEGRCLFVILLMAVYWCTEALPLSVTALLPIVLFPFMGILPSNK 80
Query: 105 VAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFF 164
V Y D L L I+A A+E +N+HRR+AL + ++ + P+ L+LG+ TT F
Sbjct: 81 VCPQYFLDTNFLFLSGLIMASAIEEWNLHRRIALKILMLV---GVQPARLILGMMVTTSF 137
Query: 165 VSMWLHNVATAVMMMPVATGIL-----------------------------HRLP----- 190
+SMWL N A+ MM+P+A IL H +P
Sbjct: 138 LSMWLSNTASTAMMLPIANAILKSLFGQKEVRKDPSQESEENTAAVRRNGLHTVPTEMQF 197
Query: 191 -------------------PAE--EQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNL 229
PA+ ++ E + ++++ Y+ IGG +TLTGT NL
Sbjct: 198 LASTEAKDHPGETEVPLDLPADSRKEDEYRRNIWKGFLISIPYSASIGGTATLTGTAPNL 257
Query: 230 IIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV-------RKGSSRAL 282
I++G KS P+ ++F SWF F FP+ +L LL W + +Y RK S
Sbjct: 258 ILLGQLKSFFPQCDVVNFGSWFIFAFPLMLLFLLAGWLWISFLYGGLSFRGWRKNKSEIR 317
Query: 283 SDYLDRA--LLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-GLV 339
++ DRA +++ + + LGP+ FAE+ V +F + +L TR +PGW LF+ G +
Sbjct: 318 TNAEDRARAVIREEYQNLGPIKFAEQAVFILFCMFAILLFTRD-PKFIPGWASLFNPGFL 376
Query: 340 GDGSISVLAAVLLFIIPNMKQN-------------GEKLMDWNECKK-LPWNLILLLGAG 385
D V +LF P+ + + E L+ W + ++ +PWN+ILLLG G
Sbjct: 377 SDAVTGVAIVTILFFFPSQRPSLKWWFDFKAPNTETEPLLTWKKAQETVPWNIILLLGGG 436
Query: 386 FALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLL 445
FA+A G + SGL+ + L L++VP V ++++ + TEF SN AT + +P+L
Sbjct: 437 FAMAKGCEESGLSVWIGGQLHPLENVPPALAVLLITVVIAFFTEF-ASNTATIIIFLPVL 495
Query: 446 YHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAG 505
+AI + VHPL LMIPG + FAF LP STP N + FA+GH+ +KDM++ G+ + + G
Sbjct: 496 AELAIRLRVHPLYLMIPGTVGCSFAFMLPVSTPPNSIAFASGHLLVKDMVRTGLLMNLMG 555
Query: 506 VSVLALLMPTLGKLLLSL 523
V +L+L M T + + L
Sbjct: 556 VLLLSLAMNTWAQTIFQL 573
>S132_HUMAN (Q13183) Solute carrier family 13, member 2 (Renal
sodium/dicarboxylate cotransporter) (Na(+)/dicarboxylate
cotransporter 1) (NaDC-1)
Length = 592
Score = 290 bits (742), Expect = 6e-78
Identities = 189/551 (34%), Positives = 292/551 (52%), Gaps = 79/551 (14%)
Query: 49 IICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHS 108
I+ L + + P+ I + +W T A+PL VT++ PL LFP+ GI A VA
Sbjct: 22 ILLLPLPILVPSKEAYCAYAIILMALFWCTEALPLAVTALFPLILFPMMGIVDASEVAVE 81
Query: 109 YMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMW 168
Y+ D L G ++A+AVE +N+H+R+AL V L+ + P+ L+LG T F+SMW
Sbjct: 82 YLKDSNLLFFGGLLVAIAVEHWNLHKRIALRVLLIV---GVRPAPLILGFMLVTAFLSMW 138
Query: 169 LHNVATAVMMMPVATGILHRL-----------------------PPAEEQPELMN----- 200
+ N AT+ MM+P+A +L +L P +E +L N
Sbjct: 139 ISNTATSAMMVPIAHAVLDQLHSSQASSNVEEGSNNPTFELQEPSPQKEVTKLDNGQALP 198
Query: 201 -----------------KFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPE-A 242
++ + L V Y+ IGGI+TLTGT NL++ G SL P+
Sbjct: 199 VTSASSEGRAHLSQKHLHLTQCMSLCVCYSASIGGIATLTGTAPNLVLQGQINSLFPQNG 258
Query: 243 KPISFNSWFFFGFPVAVLLLLCFWCILCLIYV----RK--GSSRALSDYLDRA--LLKRD 294
++F SWF F FP V+LLL W L ++++ RK G + + A +++ +
Sbjct: 259 NVVNFASWFSFAFPTMVILLLLAWLWLQILFLGFNFRKNFGIGEKMQEQQQAAYCVIQTE 318
Query: 295 LEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLF------HGLVGDGSISVLA 348
LGPM+FAEK + +F +L++LW TR L GWG L +V DG++++
Sbjct: 319 HRLLGPMTFAEKAISILFVILVLLWFTREPGFFL-GWGNLAFPNAKGESMVSDGTVAIFI 377
Query: 349 AVLLFIIPN----MKQNGEK---------LMDWNEC-KKLPWNLILLLGAGFALADGVQS 394
+++FIIP+ + Q+ E L+DW +K+PWN++LLLG G+ALA G +
Sbjct: 378 GIIMFIIPSKFPGLTQDPENPGKLKAPLGLLDWKTVNQKMPWNIVLLLGGGYALAKGSER 437
Query: 395 SGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHV 454
SGL++ + L L+ VP AI +SLL + TE TSN AT T+ +P+L +A + +
Sbjct: 438 SGLSEWLGNKLTPLQSVPAPAIAIILSLLVATFTE-CTSNVATTTIFLPILASMAQAICL 496
Query: 455 HPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMP 514
HPL +M+P +AT AF LP +TP N + F+ G +++ DM + G L + GV ++AL +
Sbjct: 497 HPLYVMLPCTLATSLAFMLPVATPPNAIVFSFGDLKVLDMARAGFLLNIIGVLIIALAIN 556
Query: 515 TLGKLLLSLFS 525
+ G L SL S
Sbjct: 557 SWGIPLFSLHS 567
>S133_MOUSE (Q91Y63) Solute carrier family 13, member 3
(Sodium-dependent high-affinity dicarboxylate
transporter 2) (Na(+)/dicarboxylate cotransporter 3)
(NaDC-3) (mNaDC3)
Length = 600
Score = 286 bits (732), Expect = 9e-77
Identities = 184/557 (33%), Positives = 282/557 (50%), Gaps = 83/557 (14%)
Query: 45 LLSIIICLLVKLDA-PTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASAD 103
LL + + LL L A P + L VI + +W T A+PL VT++ P+ LFP GI +
Sbjct: 20 LLLVPLALLPILFALPPKEGRCLYVILLMAVYWCTEALPLSVTALLPIILFPFMGILPSS 79
Query: 104 TVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTF 163
V Y D L L I+A A+E +N+HRR+AL V ++ + P+ L+LG+ TT
Sbjct: 80 KVCPQYFLDTNFLFLSGLIMASAIEEWNLHRRIALKVLMLV---GVQPARLILGMMVTTS 136
Query: 164 FVSMWLHN---------VATAVMM------------------------------MPVATG 184
F+SMWL N +A+A++ +P
Sbjct: 137 FLSMWLSNTASTAMMLPIASAILKSLFGQREARKDLPREGDESTAAVQGNGLRTVPTEMQ 196
Query: 185 ILH--------------RLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLI 230
L LP ++ E + ++++ Y+ IGG +TLTGT NLI
Sbjct: 197 FLASSEGGHTEDAEAPMELPDDSKEEEHRRNIWKGFLISIPYSASIGGTATLTGTAPNLI 256
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV-------RKGSSRALS 283
++G KS P+ ++F SWF F FP+ +L LL W + +Y RK S+ +
Sbjct: 257 LLGQLKSFFPQCDVVNFGSWFIFAFPLMLLFLLVGWLWISFLYGGMSWRSWRKKKSKIRA 316
Query: 284 DYLD--RALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-GLVG 340
D D +A+++ + + LGP+ FAE+ V +F +L +R +PGW LF G V
Sbjct: 317 DAEDQAKAVIQEEFQNLGPIKFAEQAVFILFCTFAILLFSRD-PKFIPGWASLFAPGFVS 375
Query: 341 DGSISVLAAVLLFIIPNMKQN-------------GEKLMDWNECKK-LPWNLILLLGAGF 386
D V +LF P+ K + E L+ W + ++ +PWN+ILLLG GF
Sbjct: 376 DAVTGVAIVTILFFFPSQKPSLKWWFDFKAPNSETEPLLSWKKAQETVPWNIILLLGGGF 435
Query: 387 ALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLY 446
A+A G + SGL+ + L L+ VP L V ++++ + TEF SN AT + +P+L
Sbjct: 436 AMAKGCEESGLSAWIGGQLHPLEHVPPLLAVLLITVVIAFFTEF-ASNTATIIIFLPVLA 494
Query: 447 HIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGV 506
+AI +HVHPL LMIPG + +AF LP STP N + F+TGH+ +KDM++ G+ + + GV
Sbjct: 495 ELAIRLHVHPLYLMIPGTVGCSYAFMLPVSTPPNSIAFSTGHLLVKDMVRTGLLMNLMGV 554
Query: 507 SVLALLMPTLGKLLLSL 523
+L+L M T + + L
Sbjct: 555 LLLSLAMNTWAQTIFQL 571
>S132_MOUSE (Q9ES88) Solute carrier family 13, member 2 (Renal
sodium/dicarboxylate cotransporter) (Na(+)/dicarboxylate
cotransporter 1) (NaDC-1)
Length = 586
Score = 286 bits (731), Expect = 1e-76
Identities = 181/505 (35%), Positives = 268/505 (52%), Gaps = 74/505 (14%)
Query: 76 WITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRR 135
W T A+PL VT++ P+ LFPL GI A V Y D L +G ++A+AVE +N+H+R
Sbjct: 49 WCTEALPLAVTALFPIILFPLMGIMEASKVCLEYFKDTNILFVGGLMVAIAVEHWNLHKR 108
Query: 136 LALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLP----- 190
+AL V L+ + P++LLLG T F+SMW+ N AT MM+P+ +L +L
Sbjct: 109 IALGVLLII---GVRPALLLLGFMLVTAFLSMWISNTATTAMMLPIGYAVLEQLQGSQKD 165
Query: 191 ----------------PAEEQPELMN-------------------KFSRAVILTVVYATP 215
P +E+ +L N +FS+ + L + Y+
Sbjct: 166 VEEGNSNPSFELQEASPQKEETKLDNGQAVSVSSEPRAQKTKEHHRFSQGLSLCICYSAS 225
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPE-AKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV 274
IGGI+TLTGT NL++ G S+ PE + ++F SWF F FP V+LLL W L ++++
Sbjct: 226 IGGIATLTGTTPNLVLQGQVNSIFPENSNVVNFASWFGFAFPTMVILLLLAWLWLQVLFL 285
Query: 275 ----RK----GSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISD 326
RK G ++K LGPMSFAEK V +F LL+VLW TR
Sbjct: 286 GVNFRKNFGFGEGEEERKQAAFQVIKTQHRLLGPMSFAEKAVTFLFVLLVVLWFTRE-PG 344
Query: 327 DLPGWGVLF------HGLVGDGSISVLAAVLLFIIPNM--------KQNGE-----KLMD 367
PGWG +V DG++++ ++++FIIP+ K+ G+ ++
Sbjct: 345 FFPGWGDTAFANKKGQSMVSDGTVAIFISLIMFIIPSKIPGLTEDPKKPGKLKAPPAILT 404
Query: 368 WNECK-KLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSI 426
W K+PWN+++LLG GFALA G + SGL+ + L L+ VP A V +SLL +I
Sbjct: 405 WKTVNDKMPWNILILLGGGFALAKGSEESGLSKWLGDKLTPLQHVPPSATVLILSLLVAI 464
Query: 427 ITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFAT 486
TE TSN AT TL +P+L +A + +HPL +M+P +A AF LP +TP N + F+
Sbjct: 465 FTE-CTSNVATTTLFLPILASMAQAICLHPLYVMLPCTLAASLAFMLPVATPPNAIVFSF 523
Query: 487 GHIEIKDMLKVGVPLKVAGVSVLAL 511
G +++ DM + G L + GV + L
Sbjct: 524 GGLKVSDMARAGFLLNIIGVLTITL 548
>S132_RABIT (Q28615) Solute carrier family 13, member 2 (Renal
sodium/dicarboxylate cotransporter) (Na(+)/dicarboxylate
cotransporter 1) (NaDC-1)
Length = 593
Score = 285 bits (730), Expect = 2e-76
Identities = 182/526 (34%), Positives = 274/526 (51%), Gaps = 82/526 (15%)
Query: 75 WWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHR 134
+W T+A+PL VT++ PL LFP+ GI A V Y+ D L +G +LA+AVE +N+H+
Sbjct: 48 FWCTDALPLAVTALLPLCLFPMMGIMEASEVGLEYLKDTNVLFIGGLLLAIAVEHWNLHK 107
Query: 135 RLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRL----- 189
R+AL V L+ + P++L+LG T F+SMW+ N A+ MM+P+A +L L
Sbjct: 108 RIALRVLLL---TGVRPALLILGFMVVTAFLSMWISNTASTAMMVPIAHAVLQELNNTQS 164
Query: 190 ----------------PPAEEQPELMNK-------------------------FSRAVIL 208
P +E ++ K FS+ + L
Sbjct: 165 NVEEGSDNPTFELQEPSPQKETSKVDEKDNGQAQPLPAVPLESGEHMTQEQLRFSQGMSL 224
Query: 209 TVVYATPIGGISTLTGTGVNLIIIGMWKSLAPE-AKPISFNSWFFFGFPVAVLLLLCFWC 267
V Y+ IGGI+TLTGT NL++ G SL P+ ++F SWF F FP+ V+LLL W
Sbjct: 225 CVCYSASIGGIATLTGTTPNLVLQGQMTSLFPQNPNVVNFASWFGFAFPIMVILLLLSWL 284
Query: 268 ILCLIY----------VRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIV 317
L +++ +R+ +++ LGPMSFAEK V +F +L++
Sbjct: 285 WLQILFLGINFRKNFGIREQEHEQQRKQAAYRVIQTQYRLLGPMSFAEKAVFILFVILVL 344
Query: 318 LWMTRRISDDLPGWGVLFHG------LVGDGSISVLAAVLLFIIPN----MKQNGEK--- 364
LW TR GWG L +V DGS S+L V LF++P+ + Q+ +
Sbjct: 345 LWFTRE-PGFFHGWGNLVFSDASGRVMVSDGSASILIGVFLFMVPSKIPGLTQDPDNPGR 403
Query: 365 ------LMDWNEC-KKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIV 417
L++W KK+PWN++LLLG G+ALA G + SGL+ + L L+ VP A V
Sbjct: 404 LKAPPALLNWKLVNKKMPWNIVLLLGGGYALAKGSEESGLSQWLGNKLMPLQHVPPPATV 463
Query: 418 PAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTST 477
+ LL + TE TSN AT TLL+P+L +A + +HPL +M+P +A+ AF LP +T
Sbjct: 464 FIICLLVATFTE-CTSNAATTTLLLPILASMAQAICLHPLYVMLPCTLASSLAFMLPVAT 522
Query: 478 PSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLLLSL 523
P N + F+ G + + DM + G+ L + GV V+ L + + G + L
Sbjct: 523 PPNAIVFSFGGLRVSDMARAGIMLNIIGVLVIMLAINSWGVPMFQL 568
>S133_RAT (Q9Z0Z5) Solute carrier family 13, member 3
(Sodium-dependent high-affinity dicarboxylate
transporter 2) (Na(+)/dicarboxylate cotransporter 3)
(NaDC-3) (rNaDC3)
Length = 600
Score = 284 bits (727), Expect = 4e-76
Identities = 185/558 (33%), Positives = 284/558 (50%), Gaps = 85/558 (15%)
Query: 45 LLSIIICLLVKLDA-PTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASAD 103
LL + + LL L A P + L VI + +W T A+PL VT++ P+ LFP GI +
Sbjct: 20 LLLVPLALLPILFALPPKEGRCLYVILLMAVYWCTEALPLSVTALLPIILFPFMGILPSS 79
Query: 104 TVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTF 163
V Y D L L I+A A+E N+HRR+AL V ++ + P+ L+LG+ TT
Sbjct: 80 KVCPQYFLDTNFLFLSGLIMASAIEERNLHRRIALKVLMLV---GVQPARLILGMMVTTS 136
Query: 164 FVSMWLHN---------VATAVMM------------------------------MPVATG 184
F+SMWL N +A+A++ +P
Sbjct: 137 FLSMWLSNTASTAMMLPIASAILKSLFGQRDTRKDLPREGEDSTAAVRGNGLRTVPTEMQ 196
Query: 185 ILH--------------RLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLI 230
L LP ++ E + ++++ Y+ IGG +TLTGT NLI
Sbjct: 197 FLASSEGGHAEDVEAPLELPDDSKEEEHRRNIWKGFLISIPYSASIGGTATLTGTAPNLI 256
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV-------RKGSSRALS 283
++G KS P+ ++F SWF F FP+ +L LL W + +Y RK +S+ L
Sbjct: 257 LLGQLKSFFPQCDVVNFGSWFIFAFPLMLLFLLVGWLWISFLYGGMSWRGWRKKNSK-LQ 315
Query: 284 DYLD---RALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-GLV 339
D + +A+++ + + LGP+ FAE+ V +F L +L +R +PGW LF G V
Sbjct: 316 DVAEDKAKAVIQEEFQNLGPIKFAEQAVFILFCLFAILLFSRD-PKFIPGWASLFAPGFV 374
Query: 340 GDGSISVLAAVLLFIIPNMKQN-------------GEKLMDWNECKK-LPWNLILLLGAG 385
D V +LF P+ K + E L+ W + ++ +PWN+ILLLG G
Sbjct: 375 SDAVTGVAIVTILFFFPSQKPSLKWWFDFKAPNSETEPLLSWKKAQETVPWNIILLLGGG 434
Query: 386 FALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLL 445
FA+A G + SGL+ + L L+ VP L V ++++ + TEF SN AT + +P+L
Sbjct: 435 FAMAKGCEESGLSAWIGGQLHPLEHVPPLLAVLLITVVIAFFTEF-ASNTATIIIFLPVL 493
Query: 446 YHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAG 505
+AI +HVHPL LMIPG ++ +AF LP STP N + F+TGH+ +KDM++ G+ + + G
Sbjct: 494 AELAIRLHVHPLYLMIPGTVSCSYAFMLPVSTPPNSIAFSTGHLLVKDMVRTGLLMNLMG 553
Query: 506 VSVLALLMPTLGKLLLSL 523
V +L+L M T + + L
Sbjct: 554 VLLLSLAMNTWAQAIFQL 571
>S132_RAT (P70545) Solute carrier family 13, member 2 (Intestinal
sodium/dicarboxylate cotransporter) (Na(+)/dicarboxylate
cotransporter 1) (NaDC-1)
Length = 587
Score = 258 bits (660), Expect = 2e-68
Identities = 185/554 (33%), Positives = 275/554 (49%), Gaps = 78/554 (14%)
Query: 28 TIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTS 87
T + L FY+I+ L I L + L T I + W T A+PL VT
Sbjct: 3 TCWPALWAYRFYLIV--LCLPIFLLPLPLIVQTKEAYCAYSIILMALLWCTEALPLAVTV 60
Query: 88 MCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGD 147
+ P+ LF L GI A Y D L +G ++A+AVE +N+H+R+AL V L+
Sbjct: 61 LFPIVLFLLMGIMDASE-GLEYFKDTNILFVGGLMVAIAVEHWNLHKRIALQVLLII--- 116
Query: 148 PLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLP----------------- 190
+ ++LLLG T F+SMW+ N AT MM+P+ +L +L
Sbjct: 117 GVRAALLLLGFMLVTAFLSMWISNTATTAMMVPIGHAVLEQLQASKKDVEGGNNNPTFEL 176
Query: 191 ----PAEEQPELMN-------------------KFSRAVILTVVYATPIGGISTLTGTGV 227
P +E +L N +FS+ + L + Y+ IGGI LTGT
Sbjct: 177 QEECPQKEVTKLDNGQPVSAPSEPRTQKTQEHHRFSQGLSLCICYSASIGGIDNLTGTTP 236
Query: 228 NLIIIGMWKSLAPE-AKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV----RK----GS 278
NL++ G SL P+ A ++F SWF F FP ++LLL W L ++++ RK G
Sbjct: 237 NLVLQGQVNSLFPQNANVVNFASWFGFAFPTMIILLLLAWLWLQVLFLGVNFRKNFGFGE 296
Query: 279 SRALSDYLDRALLKRDLEALGPMSFAEKMVLCV-FGLLIVLWMTRRISDDLPGWG-VLF- 335
++K LGPMSFAEK V F LL+VLW TR PGWG +F
Sbjct: 297 GEEERKQAAFQVIKTQYRLLGPMSFAEKTCSTVLFVLLVVLWFTRE-PGFFPGWGDTVFA 355
Query: 336 ----HGLVGDGSISVLAAVLLFIIPNM--------KQNGE-----KLMDWNECK-KLPWN 377
+ DG++++ ++++FIIP+ K+ G+ ++ W K+PWN
Sbjct: 356 NEKGQSMPSDGTVAIFISLVMFIIPSKIPGLMEDPKKPGKLKAPPAILTWKTVNDKMPWN 415
Query: 378 LILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDAT 437
+++LLG GFALA G + SGL++ + L L+ +P A + LL +I TE TSN AT
Sbjct: 416 IVILLGGGFALAKGSEQSGLSEWLGDKLTPLQHIPPSATAVILCLLIAIFTE-CTSNVAT 474
Query: 438 ATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKV 497
TL +P+L +A + +HPL +M+P +A AF LP +TP N + F+ G +++ DM +
Sbjct: 475 TTLFLPILASMAQAICLHPLYVMLPCTLAASLAFMLPVATPPNAIVFSFGGLKVSDMARA 534
Query: 498 GVPLKVAGVSVLAL 511
G L + GV L
Sbjct: 535 GFLLNIIGVLATTL 548
>NAD1_CAEEL (Q93655) Sodium-dependent low-affinity dicarboxylate
transporter 1 (Na(+)/dicarboxylate cotransporter 1)
(NaDC-1) (ceNaDC1)
Length = 582
Score = 255 bits (651), Expect = 2e-67
Identities = 157/536 (29%), Positives = 272/536 (50%), Gaps = 64/536 (11%)
Query: 32 ILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPL 91
+L + ++I G LL L+ D+ K L +A + ++W+ A+PL +T+ P+
Sbjct: 11 LLIYKQSFVIWGALLIFSPLLMFVGDSHGLQAKCLYCVAVMGSYWVFEALPLAITAFIPM 70
Query: 92 FLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNP 151
LFPLFGI ++ VA +Y+ D L +G ++ALAVE+ +H R+AL V +P
Sbjct: 71 ILFPLFGIMRSEEVARAYLPDTCFLFMGGLMVALAVEKCELHARVALFVLKTVGSEP--- 127
Query: 152 SMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELM------------ 199
+ ++ G T F+SMW+ N AT +M+P+ ++ L +L+
Sbjct: 128 ARVMAGFMGVTGFLSMWISNTATTALMVPILQSVITELVSNHRMEDLVALCEAHHNSSRK 187
Query: 200 ------------------------------NKFSRAVILTVVYATPIGGISTLTGTGVNL 229
K ++ ++L+V ++ IGG +T+TGT NL
Sbjct: 188 HSVGMRRLSLPNENNEIKREEMDTAMSPREQKMAKGLMLSVCFSANIGGAATITGTASNL 247
Query: 230 IIIGMWKSLAPEAKP-ISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDR 288
+++G L P A ++F SW F FP+ L+ WC+L L+Y+R ++ +
Sbjct: 248 VLVGQLNELFPGADTGVNFLSWLIFAFPMVFCCLIYCWCVLYLLYLRDAPKGSI---IVT 304
Query: 289 ALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHG-LVGDGSISVL 347
L++ L SFAE V+ F LL+VLW+ R +PGWG +F V D + ++
Sbjct: 305 RKLQQKYNELHAFSFAEMAVIFCFALLLVLWILRE-PQVVPGWGEMFKDEFVSDATSAMF 363
Query: 348 AAVLLFIIPN----------MKQNGEKLMDWNECK-KLPWNLILLLGAGFALADGVQSSG 396
+LLF +P ++ L+DW + + PW+++ LLG GFALA GV+ SG
Sbjct: 364 IVILLFTLPEKLPSSRGSSEQRKASSGLLDWATVQDRFPWSVLFLLGGGFALAAGVKESG 423
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L+ + + +L DV I+ + ++ S+ + SN A++ +P++ +A ++ + P
Sbjct: 424 LSHDIGAIMRYL-DVFNHNIIMLICIIISVTLTNVCSNTVIASIFIPIVAELARSLEIDP 482
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALL 512
L M+P I+ FAF LP +TP N + F++G++++ DM G+ + + G VL++L
Sbjct: 483 LNFMLPVTISASFAFLLPVATPPNAIVFSSGYLKVFDMFVSGLCVTL-GCVVLSML 537
>NAD3_CAEEL (Q21339) Sodium-dependent high-affinity dicarboxylate
transporter 3 (Na(+)/dicarboxylate cotransporter 3)
(NaDC-3) (ceNaDC3)
Length = 566
Score = 252 bits (644), Expect = 1e-66
Identities = 154/476 (32%), Positives = 260/476 (54%), Gaps = 33/476 (6%)
Query: 72 VFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYN 131
+ +W++ +PL VT+M P+ LFPL G+ A+T A YM+D L +G I+A AVE+ +
Sbjct: 84 IAVYWMSEVMPLAVTAMLPVVLFPLVGVLDANTTAKEYMNDTNFLFIGGLIMAAAVEKCD 143
Query: 132 VHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRL-- 189
+H R+AL+V + P ++LG T +S ++ N AT MM+P+ ++ +L
Sbjct: 144 LHERVALSVLRCVGSE---PKWIMLGFMTVTALLSSFISNTATTAMMVPIGQSVVQQLIS 200
Query: 190 -----PPAEEQPEL-MNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAK 243
P E+ L K + ++L++ +A IGG T TGT NL+++G +L P+
Sbjct: 201 SFQHHPTNGERGRLGCKKMATGLVLSICFAANIGGTGTATGTPSNLVMLGQLSALFPKVD 260
Query: 244 -PISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMS 302
+++ +W FF +P+ +L L W L ++R + D +LK L M+
Sbjct: 261 GSLNYVTWIFFAYPLMLLCLFVAWMTLVSFFLRDAPEK---DEAVTEMLKTRYNELPRMT 317
Query: 303 FAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLF-HGLVGDGSISVLAAVLLFIIP----- 356
+AEK V F +L+ LW+ R +PG+GV F G D + +++ A LLF++P
Sbjct: 318 YAEKSVFVCFCILLSLWVFRN-PGVVPGFGVFFKKGAYTDATSAMIVAFLLFVLPSERPD 376
Query: 357 --------NMKQNGEKLMDWNECKK-LPWNLILLLGAGFALADGVQSSGLADVMSKALEF 407
++K+ G LMDW ++ PW+++LLLG GFALA GV+ SGL+ ++ +L
Sbjct: 377 LATYIKKEDLKKRG-CLMDWKTMQETFPWSVVLLLGGGFALAAGVKESGLSLLIGNSLSS 435
Query: 408 LKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIAT 467
++ +P L I+ +++L +++ I SN TA++ VP++ +A HP LM+P +A+
Sbjct: 436 IEHLP-LWILQLLTMLIAMVITNICSNTVTASIFVPIVATLAQRAGHHPFTLMLPTTLAS 494
Query: 468 EFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLLLSL 523
FAF P TP N + F +G +++ DM VG + + + + L M ++ L L L
Sbjct: 495 SFAFIFPVGTPPNAIVFGSGMVKVSDMAFVGGIISLELLVLTVLYMNSIAYLTLPL 550
>YKG6_CAEEL (P46556) Hypothetical protein B0285.6 in chromosome III
Length = 577
Score = 224 bits (571), Expect = 4e-58
Identities = 156/511 (30%), Positives = 252/511 (48%), Gaps = 71/511 (13%)
Query: 68 VIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAV 127
VI + +W+ VPL VTS P+ P GI S VA Y D + S +L+LAV
Sbjct: 45 VILTMSCYWVAEVVPLAVTSFIPMIALPFLGIVSIKEVAPKYFADTNIVFFNSLMLSLAV 104
Query: 128 ERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILH 187
E +H+R+AL + L + G P L+ G T F+S+W+ + A +M P+A +L
Sbjct: 105 EECQLHKRIALKM-LTYVGT--RPHWLMAGFMIITSFISLWISDTACCALMAPIAYALLE 161
Query: 188 -----RLPPAE-------------EQPELMNK--------------FSRAVILTVVYATP 215
++ P E E PE K + ++L V +A+
Sbjct: 162 EIMIPKMRPEEKENEIEVMKIFDKEDPEEKEKKKLDTSRLSVRDRGICKCMMLLVAHASL 221
Query: 216 IGGISTLTGTGVNLIII-GMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYV 274
IGG T+ TG NLI + K+ E IS+ SW F P + + W I+ L ++
Sbjct: 222 IGGTGTINSTGPNLIFRDNIEKNFPNEDHGISYLSWMAFAIPPMIFYMFSSWFIVQLQFL 281
Query: 275 ----------RKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRI 324
+ + + + + + + LGPM++AEK L +F L ++ W++
Sbjct: 282 GPRHLMGMFREPTETEKQEEEVAKRAVWKSYDQLGPMTWAEKSTLVIFVLAVLSWVS--- 338
Query: 325 SDD--LPGWGVLFH-GLVGDGSISVLAAVLLFIIPNMKQN--------------GEKLMD 367
SD +PGW LF G V D ++A LLFI P K + E L+D
Sbjct: 339 SDPKVIPGWSDLFRKGYVTDSCSGLVAVFLLFIWPKKKPDFRIFRKDKSRPSVRQEPLID 398
Query: 368 WNEC--KKLPWNLILLLGAGFALADGVQSSGLADVMSKALEF-LKDVPYLAIVPAVSLLC 424
W +C ++ PW++ILLLGAGFA++D V+ SGL+ +++ +L + +P+ + +S++
Sbjct: 399 W-DCVRRRFPWSIILLLGAGFAISDAVRVSGLSSLIACSLNSTISKMPFFVMQIILSIVV 457
Query: 425 SIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGF 484
++TEF T N ATA++ +P+ + +A + HPL IP I F+F LP +TP+N + +
Sbjct: 458 VVMTEFST-NSATASIFIPISFKMAEAVGAHPLYFSIPTAIGPSFSFMLPMATPANAIVY 516
Query: 485 ATGHIEIKDMLKVGVPLKVAGVSVLALLMPT 515
T I + DM+ GV L + +++ A+ M T
Sbjct: 517 ETKTIRMIDMVSCGVFLNIFCIAITAINMNT 547
>IND1_DROME (Q9VVT2) I'm not dead yet protein (INDY transporter
protein) (drIndy)
Length = 572
Score = 220 bits (561), Expect = 6e-57
Identities = 152/519 (29%), Positives = 255/519 (48%), Gaps = 56/519 (10%)
Query: 49 IICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHS 108
++CL V L + + ++ + +W+T A+PL VTSM P+ FP+ GI S+D
Sbjct: 33 LLCLPVMLLNEGAEFRCMYLLLVMAIFWVTEALPLYVTSMIPIVAFPIMGIMSSDQTCRL 92
Query: 109 YMDDVITLVLGSFILALAVERYNVHRRLALNV-TLVFCGDPLNPSMLLLGLCATTFFVSM 167
Y D + + +G ++ALAVE N+H+RLAL V +V C +P L GL T F+SM
Sbjct: 93 YFKDTLVMFMGGIMVALAVEYCNLHKRLALRVIQIVGC----SPRRLHFGLIMVTMFLSM 148
Query: 168 WLHNVATAVMMMPVATGILHRLPPA---------------------EEQPELMNKFSRAV 206
W+ N A MM P+ +L L E++P K +
Sbjct: 149 WISNAACTAMMCPIIQAVLEELQAQGVCKINHEPQYQIVGGNKKNNEDEPPYPTKITLCY 208
Query: 207 ILTVVYATPIGGISTLTGTGVNLIIIGMWKS-LAPEAKPISFNSWFFFGFP-VAVLLLLC 264
L + YA+ +GG T+ GT NL G++++ + + F ++ F+ P + V LL
Sbjct: 209 YLGIAYASSLGGCGTIIGTATNLTFKGIYEARFKNSTEQMDFPTFMFYSVPSMLVYTLLT 268
Query: 265 F----WCILCLIYVRKGSSRALSDYLDRA-----LLKRDLEALGPMSFAEKMVLCVFGLL 315
F W + L + ++ + + A ++ + + LGPMS E V+ +F +
Sbjct: 269 FVFLQWHFMGLWRPKSKEAQEVQRGREGADVAKKVIDQRYKDLGPMSIHEIQVMILFIFM 328
Query: 316 IVLWMTRRISDDLPGWGVLFHGL-VGDGSISVLAAVLLFIIPN--------MKQNG---- 362
+V++ TR+ L GW L + + + ++ V+ F++P ++ G
Sbjct: 329 VVMYFTRKPGIFL-GWADLLNSKDIRNSMPTIFVVVMCFMLPANYAFLRYCTRRGGPVPT 387
Query: 363 ---EKLMDWNECK-KLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVP 418
L+ W + K+PW L+ LLG GFALA+G + SG+A ++ AL LK +P ++
Sbjct: 388 GPTPSLITWKFIQTKVPWGLVFLLGGGFALAEGSKQSGMAKLIGNALIGLKVLPNSVLLL 447
Query: 419 AVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTP 478
V L+ +T F +SN A A +++P+L +++ + +HPL L++P G+A AF LP STP
Sbjct: 448 VVILVAVFLTAF-SSNVAIANIIIPVLAEMSLAIEIHPLYLILPAGLACSMAFHLPVSTP 506
Query: 479 SNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
N + +I KDM G+ + + L + T G
Sbjct: 507 PNALVAGYANIRTKDMAIAGIGPTIITIITLFVFCQTWG 545
>IND2_DROME (Q9VDQ0) I'm not dead yet protein 2
Length = 562
Score = 213 bits (543), Expect = 8e-55
Identities = 147/525 (28%), Positives = 253/525 (48%), Gaps = 53/525 (10%)
Query: 41 ILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIA 100
I+ PL+++ I L+ K L +I + WIT +P+ VT++ PL PL G+
Sbjct: 26 IIIPLITLPI-LIYGFQTDMAEFKCLWLIVTMALLWITETLPIYVTALFPLVFCPLLGLV 84
Query: 101 SADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCA 160
+A V Y D I + LG I+AL +E N+H R+AL V + G +P L +GL +
Sbjct: 85 NASIVCKQYFTDTIVVFLGGLIVALGIEYSNLHTRIALRVIRIVGG---SPRRLFVGLMS 141
Query: 161 TTFFVSMWLHNVATAVMMMPVATGILHRL---------------PPAEEQPELMNKFSRA 205
+ F+ +W+ N A MM P+ +++ L P E +P +K + A
Sbjct: 142 VSTFMGLWISNSAGTAMMCPIVKALVNELDTNKIFPVYMTQEEEPVEEGEPPHPSKITVA 201
Query: 206 VILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAK-PISFNSWFFFGFPVAVLLLLC 264
+ YA+ IGG+ TL GTG NL+ G++ P + I+F ++ F+ P+ V++ +
Sbjct: 202 FYAGIAYASSIGGLGTLIGTGTNLVFRGIYTERFPTSTVEITFANFMFYSIPLMVIVNVT 261
Query: 265 FWCILCLIYVRKGSSRALS-------------DYLDRALLKRDLEALGPMSFAEKMVLCV 311
I+ + G R S ++ L +R ++ LGPMS E +
Sbjct: 262 L-VIIAFLITHMGLFRPNSKTGKIIAEANTNRKLMEDVLRQRHID-LGPMSCHEIQMAIA 319
Query: 312 FGLLIVLWMTRRISDDLPGWGVLFH-GLVGDGSISVLAAVLLFIIPNM------------ 358
F +IVL +TR+ +PGW L + +VG S +L+F +P
Sbjct: 320 FAFMIVLLITRK-PGFVPGWSDLINRKVVGSASGLSFIVLLIFALPTQYTFFKYCCGKGP 378
Query: 359 --KQNGEKLMDWN-ECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLA 415
Q + ++ W + +PW L+ LLG GFALA + +GL ++SKA++ L +P +
Sbjct: 379 FTAQAIDAILSWEYVLRNIPWGLLFLLGGGFALAVASRETGLNIMISKAMQVLIGLPNIV 438
Query: 416 IVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPT 475
+ +L + + F +N A +++P+L +++ + +HPL+L +P + ++LP
Sbjct: 439 VQSITFVLANFFSAF-NANVVVANIVLPILCEMSLALELHPLILTLPACLGISMVYFLPV 497
Query: 476 STPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLL 520
STP N + HI+ K G+ + G+SV + T G ++
Sbjct: 498 STPPNAIVTQYAHIKTKYFACCGIVPTIIGISVALVNTNTWGLII 542
>S131_RAT (Q07782) Solute carrier family 13, member 1 (Renal
sodium/sulfate cotransporter) (Na(+)/sulfate
cotransporter) (NaSi-1)
Length = 595
Score = 209 bits (531), Expect = 2e-53
Identities = 118/336 (35%), Positives = 196/336 (58%), Gaps = 26/336 (7%)
Query: 204 RAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLL 263
+ + L + Y++ IGG++T+TGT NLI + + P+ + ++F SWF F FPVAV+LLL
Sbjct: 238 KLMCLCIAYSSTIGGLTTITGTSTNLIFSEHFNTRYPDCRCLNFGSWFLFSFPVAVILLL 297
Query: 264 CFWCILCLIYV--------RKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLL 315
W L +++ + G ++ L + ++K++ E LGPM + E + L +F ++
Sbjct: 298 LSWIWLQWLFLGFNFKEMFKCGKTKTLKEKACAEVIKQEYEKLGPMRYQEIVTLVIFIVM 357
Query: 316 IVLWMTRRISDDLPGWGVLFH---GLVGDGSISVLAAVLLFIIP-----NMKQNGE---- 363
+LW +R + GW VLF G V D +++++A +L F+IP M G+
Sbjct: 358 ALLWFSRD-PGFVTGWSVLFSEYPGYVTDSTVALVAGILFFLIPAKKLTKMTSTGDIIAF 416
Query: 364 ---KLMDWNECKK-LPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPA 419
L+ W E + +PW++ +L+G GFALADG Q SGL+ + L L +P I+
Sbjct: 417 DYSPLITWKEFQSFMPWDIAILVGGGFALADGCQVSGLSSWIGSKLSPLGSLPVWLIILI 476
Query: 420 VSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPS 479
SL+ + +TE + SN AT T+L P+L +A +HV+PL +++P + T FAF LP + P
Sbjct: 477 SSLIVTSLTE-VASNPATITILFPILSPLAEAIHVNPLHILLPSTLCTSFAFLLPVANPP 535
Query: 480 NVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPT 515
N + F+ GH+++ DM+K G+ + + GV+V+ L M T
Sbjct: 536 NAIVFSYGHLKVIDMVKAGLGVNILGVAVVMLGMFT 571
Score = 112 bits (280), Expect = 2e-24
Identities = 57/169 (33%), Positives = 98/169 (57%), Gaps = 7/169 (4%)
Query: 33 LNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVF----TWWITNAVPLPVTSM 88
+ L N+ + L ++ +LV L P A++ T+WIT A+PL +T++
Sbjct: 1 MKLLNYAFVYRRFLLVVFTVLVLLPLPLIIRSKEAECAYILFVIATFWITEALPLSITAL 60
Query: 89 CPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDP 148
P +FP+FGI S+ VA +Y D L++G LA ++E++N+H+R+AL + ++
Sbjct: 61 LPGLMFPMFGIMSSTHVASAYFKDFHLLLIGVICLATSIEKWNLHKRIALRMVMMV---G 117
Query: 149 LNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPE 197
+NP+ L LG ++T F+SMWL N +TA M+MP+ + ++ AE + E
Sbjct: 118 VNPAWLTLGFMSSTAFLSMWLSNTSTAAMVMPIVEAVAQQITSAEAEAE 166
>S131_MOUSE (Q9JHI4) Solute carrier family 13, member 1 (Renal
sodium/sulfate cotransporter) (Na(+)/sulfate
cotransporter) (NaSi-1)
Length = 594
Score = 209 bits (531), Expect = 2e-53
Identities = 119/335 (35%), Positives = 197/335 (58%), Gaps = 25/335 (7%)
Query: 204 RAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLL 263
+ + L+V Y++ IGG++T+TGT NLI + + P+ + ++F SWF F FPVA++LLL
Sbjct: 238 KLMCLSVAYSSTIGGLTTITGTSTNLIFSEHFNTRYPDCRCLNFGSWFLFSFPVALILLL 297
Query: 264 CFWCILCLIYV-------RKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLI 316
W L +Y+ + G ++ L + ++K++ E LGPM + E + L +F ++
Sbjct: 298 LSWIWLQWLYLGFDFKMFKCGKTKTLKEKACAKVIKQEYEKLGPMRYQEIVTLVIFIVMA 357
Query: 317 VLWMTRRISDDLPGWGVLFH---GLVGDGSISVLAAVLLFIIP-----NMKQNGE----- 363
+LW +R + GW VLF G V D +++++A +L F+IP M GE
Sbjct: 358 LLWFSRD-PGFVTGWSVLFSEYPGYVTDSTVALVAGILFFLIPAKKVTKMTSAGEIIAFD 416
Query: 364 --KLMDWNECKK-LPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAV 420
L+ W E + +PW++ +L+G GFALADG Q SGL++ + L L +P I+
Sbjct: 417 YTPLITWKEFQSFMPWDIAILVGGGFALADGCQVSGLSNWIGSKLSPLGSLPVRLIILIS 476
Query: 421 SLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSN 480
SL+ + +TE + SN AT T+L P+L +A + V+PL +++P + T FAF LP + P N
Sbjct: 477 SLIVTSLTE-VASNPATITILFPILSPLAEAIQVNPLQILLPSTLCTSFAFLLPVANPPN 535
Query: 481 VVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPT 515
+ F+ GH+++ DM+K G+ + + GV+V+ L M T
Sbjct: 536 AIVFSYGHLKVIDMVKAGLGVNILGVAVVMLGMFT 570
Score = 110 bits (275), Expect = 9e-24
Identities = 55/169 (32%), Positives = 99/169 (58%), Gaps = 7/169 (4%)
Query: 33 LNLQNFYIILGPLLSIIICLLVKLDAP----TTSTKMLGVIAWVFTWWITNAVPLPVTSM 88
+ L N+ ++ L ++ +LV L P T + ++ + +WIT A+PL +T++
Sbjct: 1 MKLLNYALVYRRFLLVVFTILVFLPLPLIIRTKEAQCAYILFVIAIFWITEALPLSITAL 60
Query: 89 CPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDP 148
P +FP+FGI + VA +Y D L++G LA ++E++N+H+R+AL + ++
Sbjct: 61 LPGLMFPMFGIMRSSQVASAYFKDFHLLLIGVICLATSIEKWNLHKRIALRMVMMV---G 117
Query: 149 LNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPE 197
+NP+ L LG ++T F+SMWL N +TA M+MP+ + ++ AE + E
Sbjct: 118 VNPAWLTLGFMSSTAFLSMWLSNTSTAAMVMPIVEAVAQQIISAEAEAE 166
>NAD2_CAEEL (P32739) Sodium-dependent high-affinity dicarboxylate
transporter 2 (Na(+)/dicarboxylate cotransporter 2)
(NaDC-2) (ceNaDC2)
Length = 551
Score = 203 bits (517), Expect = 8e-52
Identities = 157/535 (29%), Positives = 256/535 (47%), Gaps = 58/535 (10%)
Query: 35 LQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLF 94
++ ++LGPL+++ + + L I ++ T+WI A P+ VTS+ PL L+
Sbjct: 10 IKKLLVLLGPLVAVPLLFF------GPEYRCLFSIIFLSTYWIGEAFPIGVTSLFPLALY 63
Query: 95 PLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNV-TLVFCGDPLNPSM 153
P+ I + ++ Y D I L + + I+A+AVE +HRR+AL + T V P+
Sbjct: 64 PILQIVPSKQISPVYFKDSIVLFMCTLIMAMAVEATGLHRRIALKLLTKVGAKQPV---- 119
Query: 154 LLLGLCATTFFVSMWLHNVATAVMMMPVATGIL-------------HRLP-PAEEQPELM 199
+LLG T F+S ++ + A +M P A +L HR P P + +
Sbjct: 120 MLLGFMCITSFISFFVSDTACTALMCPTAVALLMSMSDAVQHLKEDHRKPKPPPDDATVA 179
Query: 200 NK------------FSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAK-PIS 246
K F +A+IL +A+ IGG + +T TG NL+ PE + ++
Sbjct: 180 EKLRIDDMTPQDAGFCKALILACAHASLIGGTAIITSTGPNLVFRENIHKRYPEGQVTMT 239
Query: 247 FNSWFFFGFPVAVLLLLCFWCILCLIYV----------RKGSSRALSDYLDRALLKRDLE 296
+ W F P + LL + IL ++ R A L ++ E
Sbjct: 240 YLQWMVFAIPPMFVYLLASYIILVCYFMGPSTFARWFERPSKEEAHLKKLIEKNIQTMYE 299
Query: 297 ALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLF--HGLVGDGSISVLAAVLLFI 354
LG +S+ EK V F LLI W++R PGWG L + D VL + +LF+
Sbjct: 300 DLGDVSWGEKSVFVFFILLIGSWISRD-PGFTPGWGDLLPHRNFISDSVSGVLISCILFV 358
Query: 355 IPNMKQNG----EKLMDWNECK-KLPWNLILLLGAGFALADGVQSSGLADVMSKALE-FL 408
P + ++ W + K K W+ LL+GAG+A+++GV SGL+ ++S ++
Sbjct: 359 WPKDPFDPIDPMAPILKWTDMKSKFSWSCTLLIGAGYAISEGVDKSGLSRLISCGMKNIF 418
Query: 409 KDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATE 468
+ L + V+ + I+TEF SN +T ++ +P+ +A +M VHPL L +P +A
Sbjct: 419 VGMSSLPLQLTVTTIIVIMTEF-ASNVSTGSIFIPISLGVAESMGVHPLYLALPTTVACS 477
Query: 469 FAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLLLSL 523
FAF LP STP N V + T I + +M+ G L +A + + +L M T + SL
Sbjct: 478 FAFMLPISTPPNAVVYDTKVISMVEMIVCGFLLNIACILITSLNMNTWTYFIFSL 532
>S131_HUMAN (Q9BZW2) Solute carrier family 13, member 1 (Renal
sodium/sulfate cotransporter) (Na(+)/sulfate
cotransporter) (hNaSi-1)
Length = 595
Score = 197 bits (501), Expect = 6e-50
Identities = 109/328 (33%), Positives = 187/328 (56%), Gaps = 26/328 (7%)
Query: 208 LTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWC 267
L + Y++ IGG++T+TGT NLI + + P+ + ++F SWF F FP A+++LL W
Sbjct: 242 LCIAYSSTIGGLTTITGTSTNLIFAEYFNTRYPDCRCLNFGSWFTFSFPAALIILLLSWI 301
Query: 268 ILCLIYV--------RKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLW 319
L +++ + G ++ + ++K++ + LGP+ + E + L +F ++ +LW
Sbjct: 302 WLQWLFLGFNFKEMFKCGKTKTVQQKACAEVIKQEYQKLGPIRYQEIVTLVLFIIMALLW 361
Query: 320 MTRRISDDLPGWGVLFH---GLVGDGSISVLAAVLLFIIP-----NMKQNGE-------K 364
+R +PGW LF G D ++++L +L F+IP GE
Sbjct: 362 FSRD-PGFVPGWSALFSEYPGFATDSTVALLIGLLFFLIPAKTLTKTTPTGEIVAFDYSP 420
Query: 365 LMDWNECKK-LPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLL 423
L+ W E + +PW++ +L+G GFALADG + SGL+ + L L +P I+ SL+
Sbjct: 421 LITWKEFQSFMPWDIAILVGGGFALADGCEESGLSKWIGNKLSPLGSLPAWLIILISSLM 480
Query: 424 CSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVG 483
+ +TE + SN AT TL +P+L +A +HV+PL ++IP + T FAF LP + P N +
Sbjct: 481 VTSLTE-VASNPATITLFLPILSPLAEAIHVNPLYILIPSTLCTSFAFLLPVANPPNAIV 539
Query: 484 FATGHIEIKDMLKVGVPLKVAGVSVLAL 511
F+ GH+++ DM+K G+ + + GV+V+ L
Sbjct: 540 FSYGHLKVIDMVKAGLGVNIVGVAVVML 567
Score = 108 bits (269), Expect = 4e-23
Identities = 55/165 (33%), Positives = 98/165 (59%), Gaps = 7/165 (4%)
Query: 37 NFYIILGPLLSIIICLLVKLDAP----TTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLF 92
++ ++ L ++ +LV L P T + + V T+W+T A+PL VT++ P
Sbjct: 5 SYILVYRRFLFVVFTVLVLLPLPIVLHTKEAECAYTLFVVATFWLTEALPLSVTALLPSL 64
Query: 93 LFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPS 152
+ P+FGI + VA +Y D L++G LA ++E++N+H+R+AL + ++ +NP+
Sbjct: 65 MLPMFGIMPSKKVASAYFKDFHLLLIGVICLATSIEKWNLHKRIALKMVMMV---GVNPA 121
Query: 153 MLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPE 197
L LG ++T F+SMWL N +TA M+MP+A ++ ++ AE + E
Sbjct: 122 WLTLGFMSSTAFLSMWLSNTSTAAMVMPIAEAVVQQIINAEAEVE 166
>S134_HUMAN (Q9UKG4) Solute carrier family 13, member 4
(Na(+)/sulfate cotransporter SUT-1)
Length = 627
Score = 189 bits (479), Expect = 2e-47
Identities = 114/373 (30%), Positives = 196/373 (51%), Gaps = 43/373 (11%)
Query: 189 LPPAEEQPELMNKFS--------RAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAP 240
L P+ + +L K+ + + L++ Y+ IGG++T+ GT +LI + + + P
Sbjct: 244 LTPSPRKQKLNRKYRSHHDQMICKCLSLSISYSATIGGLTTIIGTSTSLIFLEHFNNQYP 303
Query: 241 EAKPISFNSWFFFGFPVAVLLLLCFWCIL------------CLIYVRKGSSRALSDYLDR 288
A+ ++F +WF F FP+++++L+ W + C + +K + R + L
Sbjct: 304 AAEVVNFGTWFLFSFPISLIMLVVSWFWMHWLFLGCNFKETCSLSKKKKTKR---EQLSE 360
Query: 289 ALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH--GLVGDGSISV 346
++ + E LG +S+ E + F L+ VLW TR +PGW F G D ++SV
Sbjct: 361 KRIQEEYEKLGDISYPEMVTGFFFILMTVLWFTRE-PGFVPGWDSFFEKKGYRTDATVSV 419
Query: 347 LAAVLLFIIP------NMKQNGEK---------LMDWNECKK-LPWNLILLLGAGFALAD 390
LLF+IP K +GE ++ W + +K +PW +++L+G G+ALA
Sbjct: 420 FLGFLLFLIPAKKPCFGKKNDGENQEHSLGTEPIITWKDFQKTMPWEIVILVGGGYALAS 479
Query: 391 GVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAI 450
G +SSGL+ + + L +P A+ +L SI+TEF+ SN AT T+ +P+L ++
Sbjct: 480 GSKSSGLSTWIGNQMLSLSSLPPWAVTLLACILVSIVTEFV-SNPATITIFLPILCSLSE 538
Query: 451 NMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLA 510
MH++PL +IP + FA LP P N + F+ GH +IKDM+K G+ + V G+ ++
Sbjct: 539 TMHINPLYTLIPVTMCISFAVMLPVGNPPNAIVFSYGHCQIKDMVKAGLGVNVIGLVIVM 598
Query: 511 LLMPTLGKLLLSL 523
+ + T G L L
Sbjct: 599 VAINTWGVSLFHL 611
Score = 97.8 bits (242), Expect = 6e-20
Identities = 52/155 (33%), Positives = 86/155 (54%), Gaps = 8/155 (5%)
Query: 46 LSIIIC---LLVKLDA--PTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIA 100
L +++C LL+ L P++ V+ +W++ AVPL ++ P FL+P FG+
Sbjct: 13 LLLVVCVPLLLLPLPVLHPSSEASCAYVLIVTAVYWVSEAVPLGAAALVPAFLYPFFGVL 72
Query: 101 SADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCA 160
++ VA Y + L++G +A AVE++N+H+R+AL + L+ P MLLL
Sbjct: 73 RSNEVAAEYFKNTTLLLVGVICVAAAVEKWNLHKRIALRMVLM---AGAKPGMLLLCFMC 129
Query: 161 TTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQ 195
T +SMWL N +T M+MP+ +L L AE++
Sbjct: 130 CTTLLSMWLSNTSTTAMVMPIVEAVLQELVSAEDE 164
>Y672_METJA (Q58086) Hypothetical protein MJ0672
Length = 432
Score = 154 bits (388), Expect = 7e-37
Identities = 119/439 (27%), Positives = 214/439 (48%), Gaps = 55/439 (12%)
Query: 76 WITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRR 135
W +PLPVTS+ + GI + + +I L LG F+LA A++ +N+ +
Sbjct: 40 WFFELLPLPVTSLAIPIMAVFLGIFNLKEALTYFAHPIIFLFLGGFMLAQALKNHNLDKF 99
Query: 136 LALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQ 195
+A L+ G + L+ L A +F+SMW+ N + ++++P+A G+LH+
Sbjct: 100 IAYK--LLNYGKDFKTTCFLMFLSA--YFLSMWISNTSATLILLPIALGLLHK------- 148
Query: 196 PELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGF 255
K ++L V Y+ IGGI+T+ G+ N ++A F SWF GF
Sbjct: 149 --KTGKLRDFLLLGVAYSASIGGIATIIGSPPN--------AIASSYLDYGFFSWFKVGF 198
Query: 256 PVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEK--MVLCVFG 313
P+++LL L C L L Y+ Y + + K D+ M + +L +F
Sbjct: 199 PISLLLFLI--CTLTL-YI----------YFKKWIPKEDIAIQARMELSRNAYKLLVIFV 245
Query: 314 LLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKK 373
L+ LW+ ISD L +F+ D I++ A +LLF+ L++ N+ KK
Sbjct: 246 LIASLWI---ISDYL---SEIFNVQYFDSVIAIFAIILLFVF--------NLVEVNDFKK 291
Query: 374 LPWNLILLLGAGFALADGVQSSGLADVMS-KALEFLKDVPYLAIVPAVSLLCSIITEFIT 432
+ W ++L G L + SG +S K + L ++ + ++ V + I+T FI
Sbjct: 292 IDWGTLILFGGALCLGGVIVKSGANTFLSEKLIAILGNLTPIVLLFLVVTITIILTNFI- 350
Query: 433 SNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIK 492
SN ++VP+L+ +++ + L+L + G++ +F LP TP N + ++ G ++ +
Sbjct: 351 SNTGLTGIIVPILFGVSLGIPKEILILAV--GMSASCSFILPVGTPPNAIVYSEG-VKKE 407
Query: 493 DMLKVGVPLKVAGVSVLAL 511
+M+K+G+ L + +V+ L
Sbjct: 408 EMMKIGMILSILSAAVITL 426
>Y608_HAEIN (Q57486) Hypothetical protein HI0608
Length = 461
Score = 152 bits (385), Expect = 2e-36
Identities = 116/451 (25%), Positives = 222/451 (48%), Gaps = 40/451 (8%)
Query: 66 LGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILAL 125
L ++A++ W++ A+ + +T++ L G+ S + D I L G F LA
Sbjct: 40 LALLAFIAVLWLSEALHVTITALLVPLLAVALGLVSTKQALVGFADPTIFLFFGGFSLAT 99
Query: 126 AVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGI 185
A+ + + +A + + G N + ++ L T F+SMW+ N ATA MM+P+A GI
Sbjct: 100 ALHIQKLDKLIANKIMALARG---NLFIAVIYLFLITAFLSMWMSNTATAAMMLPLAMGI 156
Query: 186 LHRLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPI 245
L +L ++ + V+L + Y+ IGG+ TL G+ N I+ +
Sbjct: 157 LSQLDREKDHNTYV-----FVLLGIAYSASIGGMGTLVGSPPNAIVASNLN--------L 203
Query: 246 SFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAE 305
+F+ W ++G P+ ++LL IL +I+ K +L+ ++E + PM
Sbjct: 204 TFSDWLWYGLPIMIILLPLMIGILYIIFKPKL-------HLNFEQTFENIE-MNPMRI-- 253
Query: 306 KMVLCVFGLLIVLWM-TRRISDDLPGWGVLFHGLVG-DGSISVLAAVLLFIIPNMKQNGE 363
+ +F ++ + W+ + +I+ + G L + D +++LAA+++
Sbjct: 254 -LTFIIFPVIALTWIFSGKINPFISGLLGLQKNIASFDSIVALLAAIVIC--------ST 304
Query: 364 KLMDWNECKK-LPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPA-VS 421
+ W + + W +++L G G L+ ++ SG + +++ ++ F+ D + ++ V+
Sbjct: 305 GVASWKQIQSNTDWGVLMLFGGGLTLSAVLKDSGASKILADSIVFMIDGQHFYLIGLLVA 364
Query: 422 LLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNV 481
+TEF TSN A+A LLVP+ IA ++ + + L + GI AF LP +TP N
Sbjct: 365 AFIIFLTEF-TSNTASAALLVPIFISIAQSLGMPEIGLALIIGIGASCAFMLPVATPPNA 423
Query: 482 VGFATGHIEIKDMLKVGVPLKVAGVSVLALL 512
+ F +G ++ +M+KVG L + V V+A +
Sbjct: 424 IVFGSGQVKQSEMVKVGFLLNLVCVVVIATM 454
>YN86_YEAST (P27514) Hypothetical 99.5 kDa protein in URK1-SMM1
intergenic region
Length = 894
Score = 127 bits (319), Expect = 7e-29
Identities = 116/470 (24%), Positives = 203/470 (42%), Gaps = 64/470 (13%)
Query: 34 NLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTS-MCPLF 92
+L F +I ++++ L + ++ + W T +PL VTS M PL
Sbjct: 428 SLLKFLLITSCFIALLTFNLTPFTQDSLQKNCFAILIYASLLWATETIPLFVTSLMIPLL 487
Query: 93 LF------------PLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNV 140
+ P+ S+ + + VI L+LG F LA A+ +YN+ + L+
Sbjct: 488 IVVFPVIKDPITSQPMSPRDSSQFILSTMWSSVIMLLLGGFTLAAALSKYNIAKVLS--- 544
Query: 141 TLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELMN 200
T + NP +LL FVSMW+ NVA V+ + +L LP
Sbjct: 545 THILASAGTNPHFILLTNMFVALFVSMWVSNVAAPVLCYSIVQPLLRTLPRN-------C 597
Query: 201 KFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVL 260
+++A+IL + A+ IGG+S+ + N+ IG+ + P S+ WF PV +
Sbjct: 598 SYAKALILGIALASNIGGMSSPIASPQNIFSIGIM-----DPSP-SWAEWFMIALPVCFI 651
Query: 261 LLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLW- 319
++ W +L + + + + + L + R P + + V V + IVLW
Sbjct: 652 CVMAIWVLLIITFPPEPNVKILQLHPSR----------DPFTLKQWFVTLVCIITIVLWC 701
Query: 320 MTRRISDDLPGWGVLFHGLVGD-GSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNL 378
++ +IS G+ G+ G IS++ V+ F G L+ ++ W +
Sbjct: 702 LSNQIS-----------GIFGEMGIISIIPIVVFF--------GTGLLTSDDFNNFMWTI 742
Query: 379 ILLLGAGFALADGVQSSGLADVMSKALEF-LKDVPYLAIVPAVSLLCSIITEFITSNDAT 437
++L G L V SSGL M++ ++ ++ P IV L+ ++ F+ S+
Sbjct: 743 VVLAMGGTTLGKAVSSSGLLSTMAQLIKAQVEHEPIFIIVLIFGLVILVMATFV-SHTVA 801
Query: 438 ATLLVPLLYHIAINMHV--HPLLLMIPGGIATEFAFWLPTSTPSNVVGFA 485
A ++VPL+ I N+ H LL++ + A LPTS NV +
Sbjct: 802 AMIIVPLMSEIGSNLPSGDHSRLLIVIAALLCSSAMGLPTSGFPNVTAIS 851
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,636,292
Number of Sequences: 164201
Number of extensions: 2539471
Number of successful extensions: 8037
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 7834
Number of HSP's gapped (non-prelim): 94
length of query: 530
length of database: 59,974,054
effective HSP length: 115
effective length of query: 415
effective length of database: 41,090,939
effective search space: 17052739685
effective search space used: 17052739685
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148482.2


