

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.2 + phase: 0
(209 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
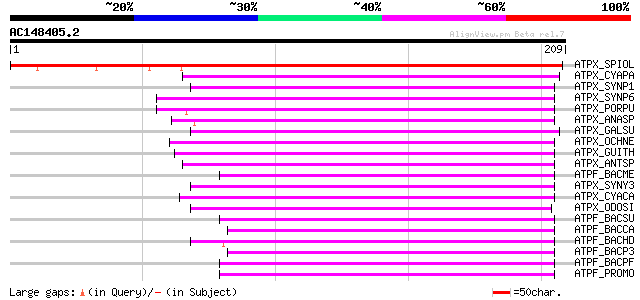
Sequences producing significant alignments: (bits) Value
ATPX_SPIOL (P31853) ATP synthase B' chain, chloroplast precursor... 243 3e-64
ATPX_CYAPA (P48085) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 93 5e-19
ATPX_SYNP1 (Q05367) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 92 7e-19
ATPX_SYNP6 (P08446) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 85 1e-16
ATPX_PORPU (P51245) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 84 2e-16
ATPX_ANASP (P12410) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 82 9e-16
ATPX_GALSU (P35012) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 82 1e-15
ATPX_OCHNE (Q40608) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 80 3e-15
ATPX_GUITH (O78478) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 79 8e-15
ATPX_ANTSP (Q02852) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 77 2e-14
ATPF_BACME (P20601) ATP synthase B chain (EC 3.6.3.14) 74 2e-13
ATPX_SYNY3 (P27183) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 73 4e-13
ATPX_CYACA (Q9TM29) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 73 5e-13
ATPX_ODOSI (Q00823) ATP synthase B' chain (EC 3.6.3.14) (Subunit... 65 9e-11
ATPF_BACSU (P37814) ATP synthase B chain (EC 3.6.3.14) 63 4e-10
ATPF_BACCA (P41014) ATP synthase B chain (EC 3.6.3.14) 60 5e-09
ATPF_BACHD (Q9K6H1) ATP synthase B chain (EC 3.6.3.14) 59 1e-08
ATPF_BACP3 (P09221) ATP synthase B chain precursor (EC 3.6.3.14) 57 3e-08
ATPF_BACPF (P22481) ATP synthase B chain (EC 3.6.3.14) 55 9e-08
ATPF_PROMO (P21904) ATP synthase B chain, sodium ion specific (E... 52 1e-06
>ATPX_SPIOL (P31853) ATP synthase B' chain, chloroplast precursor
(EC 3.6.3.14) (Subunit II)
Length = 222
Score = 243 bits (619), Expect = 3e-64
Identities = 137/222 (61%), Positives = 169/222 (75%), Gaps = 14/222 (6%)
Query: 1 MASMIMASS-KPIIRVPTNSLPSTPKLPSLQT---------ITPNLKLQLSKLKSMTLAA 50
MA+M++ASS K + T ++ PK P L+T + P L LS LKS + A
Sbjct: 1 MANMLVASSSKTLPTTTTTTITPKPKFPLLKTPLLKLSPPQLPPLKHLNLSVLKSAAITA 60
Query: 51 T--SLSFVFAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRD 106
T +LSF+ PSLA EKA+LFDFNLTLPII+ EFLFLMFALDK+Y+TPLG FMD RD
Sbjct: 61 TPLTLSFLLPYPSLAEEIEKASLFDFNLTLPIIMAEFLFLMFALDKIYYTPLGDFMDKRD 120
Query: 107 AEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKK 166
A I+ +L+ V +TS EVK+LEEQANAV+RAARAEI+ ALN+MKKETQ EVE K+AEGRKK
Sbjct: 121 ASIKEQLSGVKDTSSEVKQLEEQANAVMRAARAEISAALNKMKKETQLEVEAKLAEGRKK 180
Query: 167 VDAELQEALANLEKQKEETVKALDSQIAALSQDIVNKVLPVS 208
++ ELQEAL +LE+QKE+T+K+LDSQI+ALS DIV KVLPVS
Sbjct: 181 IEVELQEALGSLEQQKEDTIKSLDSQISALSDDIVKKVLPVS 222
>ATPX_CYAPA (P48085) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 164
Score = 92.8 bits (229), Expect = 5e-19
Identities = 49/142 (34%), Positives = 77/142 (53%)
Query: 66 KAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKE 125
+ LFDF+ TLP+++V+ L LM L+ +++ PL +D R I+ N + E
Sbjct: 18 EGGLFDFDATLPVMMVQLLVLMLILNAVFYKPLIKILDERKEYIQSNFNEAEKCLAQAAE 77
Query: 126 LEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEET 185
L Q + AR + N + E Q V EK+ E +KK D+EL A LE QK+E
Sbjct: 78 LTTQYETKITDARQNASKLTNTTRSEIQRFVSEKLEEAQKKADSELASATNKLELQKDEA 137
Query: 186 VKALDSQIAALSQDIVNKVLPV 207
+K+L+S++ LS I+ K+L +
Sbjct: 138 LKSLESEVQTLSTKILEKLLGI 159
>ATPX_SYNP1 (Q05367) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 159
Score = 92.4 bits (228), Expect = 7e-19
Identities = 47/137 (34%), Positives = 79/137 (57%)
Query: 69 LFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEE 128
+FDF+ TLP++ V+FL L L+ L + PLG +DNRD IR L ++ EL
Sbjct: 22 MFDFDATLPLMAVQFLILTVILNALLYKPLGQALDNRDEYIRTNLQQAKERLQQATELAN 81
Query: 129 QANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKA 188
Q L R E + + + E Q +IA ++ + AEL +A A +++QK+ T++A
Sbjct: 82 QYEQELAYTRREAQAIIEEARAEAQKIATAEIAAAQQALQAELMKAQAEIDQQKQATLQA 141
Query: 189 LDSQIAALSQDIVNKVL 205
L+ Q+++LS+ ++ K+L
Sbjct: 142 LEGQVSSLSEQLLAKLL 158
>ATPX_SYNP6 (P08446) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 158
Score = 85.1 bits (209), Expect = 1e-16
Identities = 44/150 (29%), Positives = 82/150 (54%)
Query: 56 VFAPPSLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNS 115
+ A ++ + LFD + TLP++ V+ L L+F L+ +++ P G +D+RD +RG
Sbjct: 6 ILAAEAVQEAEGGLFDLDATLPLMAVQILVLVFLLNAVFYKPFGKVLDDRDQFVRGGRQD 65
Query: 116 VTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEAL 175
EVK L Q L A R + + + + E +++AE +++ A+ ++A
Sbjct: 66 AKARLAEVKALTAQYEQELAATRKQSQALIAEAQTEAGRIAAQQLAEAQREAQAQREQAQ 125
Query: 176 ANLEKQKEETVKALDSQIAALSQDIVNKVL 205
+++QK ++ALD Q+ ALS I++K+L
Sbjct: 126 QEIDQQKAVALQALDQQVDALSHQILDKLL 155
>ATPX_PORPU (P51245) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 158
Score = 84.0 bits (206), Expect = 2e-16
Identities = 49/151 (32%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 56 VFAPPSLAFE-KAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLN 114
++ P LA E + LFDFN TLP++ ++FL LM L+ +++ P+ +D RD IR L
Sbjct: 2 IYLPLLLAEEIEGGLFDFNGTLPLMALQFLILMLLLNTIFYKPVTKILDERDEYIRTTLT 61
Query: 115 SVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEA 174
+ ++ + EL + L AR + + +K+ Q+ V E I + + + + EA
Sbjct: 62 TASSMLVKADELAAKYEEDLSKARRDAQAKIAASQKDAQSIVSEDIKKAQMNAEKLITEA 121
Query: 175 LANLEKQKEETVKALDSQIAALSQDIVNKVL 205
L QKEE +K L+ Q+ LS I K+L
Sbjct: 122 SKQLNIQKEEALKTLEDQVDTLSDQIKTKLL 152
>ATPX_ANASP (P12410) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 163
Score = 82.0 bits (201), Expect = 9e-16
Identities = 45/148 (30%), Positives = 78/148 (52%), Gaps = 4/148 (2%)
Query: 62 LAFEKAA----LFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVT 117
LA EK A LFD + TLP++ ++FL L L+ + PLG +D R+ +R
Sbjct: 8 LAVEKVAKEGGLFDLDATLPLMAIQFLLLALILNATLYKPLGKAIDGRNEYVRNNQLEAQ 67
Query: 118 NTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALAN 177
+ ++L E L AR + + + E Q EK+A +K+ A+ ++A
Sbjct: 68 ERLSKAEKLAEAYEQELAGARRQAQTIIADAQAEAQKIAAEKVAAAQKEAQAQREQAAGE 127
Query: 178 LEKQKEETVKALDSQIAALSQDIVNKVL 205
+E+QK++ + +L+ Q+ ALS+ I+ K+L
Sbjct: 128 IEQQKQQALASLEQQVDALSRQILEKLL 155
>ATPX_GALSU (P35012) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 157
Score = 81.6 bits (200), Expect = 1e-15
Identities = 42/139 (30%), Positives = 76/139 (54%)
Query: 69 LFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEE 128
LFDF+ TL I +EFL L L+ +Y+ P+ +D+R+ IR LN + ++ EL +
Sbjct: 17 LFDFDATLSFIALEFLLLTSVLNLIYYQPISKVIDSREDYIRENLNKASLYLDQANELTK 76
Query: 129 QANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKA 188
+ L AR E + + E Q V +I++ +K+ +Q ++ EK+K + + +
Sbjct: 77 KYELELITARKEAIKMVTTSQTEAQEFVNAQISQAQKEAQQLIQSSMMQFEKEKNKAIYS 136
Query: 189 LDSQIAALSQDIVNKVLPV 207
L+ Q+ LS+ I NK++ +
Sbjct: 137 LEKQVEQLSEQIKNKLISI 155
>ATPX_OCHNE (Q40608) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 163
Score = 80.5 bits (197), Expect = 3e-15
Identities = 45/145 (31%), Positives = 81/145 (55%)
Query: 61 SLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTS 120
+L+ + LFDFN TLP++ ++F+ L L +++ P+G ++ R+A I G L+ +
Sbjct: 13 ALSEGEGGLFDFNATLPLMALQFILLTVILTFVFYKPIGNLLEEREAYINGNLSDASAKL 72
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK 180
+ EL +Q L+ A+A+ + + E + V ++A+ RK + +++ LE
Sbjct: 73 LQADELCKQYEEQLKDAKADAQSCIADAETEAKQVVALELAQARKDAASLVEQVNKELEA 132
Query: 181 QKEETVKALDSQIAALSQDIVNKVL 205
QKE +K L++QI LSQ I K+L
Sbjct: 133 QKELALKQLEAQIDELSQLIKEKLL 157
>ATPX_GUITH (O78478) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 163
Score = 79.0 bits (193), Expect = 8e-15
Identities = 41/143 (28%), Positives = 77/143 (53%)
Query: 63 AFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEE 122
A + LFDFN TLP++ V+ L M L+ +++ P+ +D R+ IR L ++ +
Sbjct: 15 AESEGGLFDFNATLPLMAVQILLFMVILNAVFYNPVAKVLDEREEYIRKNLTQASDILAK 74
Query: 123 VKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQK 182
+ + +Q L R E + ++ +KE Q V +I + +K + + EA + L QK
Sbjct: 75 AEAITKQYEKDLAQERREAQLIISVAQKEAQDIVALEIKQAQKDTELLVNEATSQLNSQK 134
Query: 183 EETVKALDSQIAALSQDIVNKVL 205
++ + AL+ Q+ L++ I +K+L
Sbjct: 135 QKALSALEDQVNTLTEQIKSKLL 157
>ATPX_ANTSP (Q02852) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 159
Score = 77.4 bits (189), Expect = 2e-14
Identities = 42/140 (30%), Positives = 71/140 (50%)
Query: 66 KAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKE 125
+ LFDFN TLP++ ++FL L L+ +Y+ PLG +D RD I L + + + +
Sbjct: 18 QGGLFDFNATLPLMALQFLALTIILNLIYYKPLGKILDERDEYIANSLTAASAALSKAND 77
Query: 126 LEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEET 185
L ++ L +R + + +++ Q V KI E +K D + L QKE+
Sbjct: 78 LTKRYEQDLAESRKKAQDIIKNAQQDAQNIVSSKIKEAQKDADQLMSNTYDQLNIQKEQA 137
Query: 186 VKALDSQIAALSQDIVNKVL 205
++ L+ Q+ LS I K+L
Sbjct: 138 LQNLEKQVDILSNQIQIKLL 157
>ATPF_BACME (P20601) ATP synthase B chain (EC 3.6.3.14)
Length = 172
Score = 73.9 bits (180), Expect = 2e-13
Identities = 40/126 (31%), Positives = 70/126 (54%)
Query: 80 VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARA 139
+V FL L+ L K F P+ M R+ I G+++ +EE K+L E+ +L+ +R
Sbjct: 23 LVMFLILLALLQKFAFGPVMGIMKKREEHIAGEIDEAEKQNEEAKKLVEEQREILKQSRQ 82
Query: 140 EIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQD 199
E+ V + +K + + EE +A R++ + A +E+QK++ V AL Q+A+LS
Sbjct: 83 EVQVMMENARKSAEDKKEEIVAAAREESERLKAAAKQEIEQQKDQAVAALREQVASLSVL 142
Query: 200 IVNKVL 205
I +KV+
Sbjct: 143 IASKVI 148
>ATPX_SYNY3 (P27183) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 143
Score = 73.2 bits (178), Expect = 4e-13
Identities = 35/137 (25%), Positives = 73/137 (52%)
Query: 69 LFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEE 128
+FDF+ TLP++ ++F+ L F L+ +++ P+ +D R IR + K + +
Sbjct: 1 MFDFDATLPLMALQFVVLAFLLNAIFYKPMNKVLDERADYIRTNEEDARERLAKAKAITQ 60
Query: 129 QANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKA 188
+ + AR + + + E + EKIAE +++ + + A +E Q++ + +
Sbjct: 61 EYEQQITDARRQSQAVIADAQAEARRLAAEKIAEAQRESQRQKETAAQEIEAQRQSALSS 120
Query: 189 LDSQIAALSQDIVNKVL 205
L+ ++AALS I++K+L
Sbjct: 121 LEQEVAALSNQILHKLL 137
>ATPX_CYACA (Q9TM29) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 157
Score = 72.8 bits (177), Expect = 5e-13
Identities = 37/141 (26%), Positives = 75/141 (52%)
Query: 65 EKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVK 124
E LFDF+ TLP++ +FL +M LD ++ P+ + +R+ I L S T SE+ K
Sbjct: 13 ESGGLFDFDATLPLMASQFLLIMLILDITFYKPINKVLKDRENYILKTLESATQISEKTK 72
Query: 125 ELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEE 184
E + V+ ++ E ++ +K +T+ ++ ++ + + + +++ L ++KE+
Sbjct: 73 ETLARYEEVILKSKKESQQLIDSIKTKTEHDIVNELIQTQNSTREFISKSIKELYRKKEQ 132
Query: 185 TVKALDSQIAALSQDIVNKVL 205
T+K L+ LS I K++
Sbjct: 133 TLKVLEEDTENLSDKIYLKLI 153
>ATPX_ODOSI (Q00823) ATP synthase B' chain (EC 3.6.3.14) (Subunit
II)
Length = 156
Score = 65.5 bits (158), Expect = 9e-11
Identities = 37/136 (27%), Positives = 69/136 (50%)
Query: 69 LFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEE 128
LFD N TLP++ ++FL LM L+ + ++PL T ++ R I KL + + EL
Sbjct: 19 LFDINATLPLVAIQFLLLMVLLNVILYSPLLTIIEERKEYILNKLAKASEILSQANELTA 78
Query: 129 QANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKA 188
Q L R E + + +K + +E ++ +K +D L+ +L ++K + +
Sbjct: 79 QYEQELNTVRKEAQLEITNSQKIHKEILEIELNISQKYIDNLLENITDDLLEKKNTALNS 138
Query: 189 LDSQIAALSQDIVNKV 204
LD+ + +L I N++
Sbjct: 139 LDTIVQSLCVQIENRL 154
>ATPF_BACSU (P37814) ATP synthase B chain (EC 3.6.3.14)
Length = 170
Score = 63.2 bits (152), Expect = 4e-10
Identities = 38/126 (30%), Positives = 64/126 (50%)
Query: 80 VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARA 139
++ L L+ L K PL M R+ I G++ S ++E ++L E+ +L+ AR
Sbjct: 21 LLAMLILLALLKKYALGPLLNIMKQREDHIAGEITSAEEKNKEAQQLIEEQRVLLKEARQ 80
Query: 140 EIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQD 199
E + KK + + EE I R + + + A + K+KE+ V AL Q+A+LS
Sbjct: 81 ESQTLIENAKKLGEKQKEEIIQAARAESERLKEAARTEIVKEKEQAVSALREQVASLSVM 140
Query: 200 IVNKVL 205
I +KV+
Sbjct: 141 IASKVI 146
>ATPF_BACCA (P41014) ATP synthase B chain (EC 3.6.3.14)
Length = 162
Score = 59.7 bits (143), Expect = 5e-09
Identities = 31/123 (25%), Positives = 64/123 (51%)
Query: 83 FLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIA 142
F+ L+ L K + PL M R+ I +++ +E ++L E+ +++ +R +
Sbjct: 29 FIILLALLRKFAWQPLMNIMKQREEHIANEIDQAEKRRQEAEKLLEEQRELMKQSRQDDE 88
Query: 143 VALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQDIVN 202
+ +K Q + E+ +A R + + + A +E++KE+ + AL Q+A+LS I +
Sbjct: 89 ALIENARKLAQEQKEQIVASARGQAERVKEAAKKEIEREKEQAMAALREQVASLSVVIAS 148
Query: 203 KVL 205
KV+
Sbjct: 149 KVI 151
>ATPF_BACHD (Q9K6H1) ATP synthase B chain (EC 3.6.3.14)
Length = 162
Score = 58.5 bits (140), Expect = 1e-08
Identities = 37/138 (26%), Positives = 69/138 (49%), Gaps = 1/138 (0%)
Query: 69 LFDFNLTLPII-VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELE 127
+FD N I ++ F L++ L K PL M+ R+ I +++ + +E
Sbjct: 1 MFDINWGSIIYQLIAFCVLLWLLSKFALKPLMGVMEKREQMINDQIDQADKDRKAAQEYL 60
Query: 128 EQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVK 187
EQ + AR E + + KK ++ + +E + R + + + ALA ++++KE+ V
Sbjct: 61 EQQRLAVEKAREEAQEIVQKAKKLSEQQGQEIVEAARAEGERLKEAALAEIQREKEQAVA 120
Query: 188 ALDSQIAALSQDIVNKVL 205
+L Q+A+LS I KV+
Sbjct: 121 SLREQVASLSVLIATKVI 138
>ATPF_BACP3 (P09221) ATP synthase B chain precursor (EC 3.6.3.14)
Length = 163
Score = 57.0 bits (136), Expect = 3e-08
Identities = 32/123 (26%), Positives = 61/123 (49%)
Query: 83 FLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIA 142
F+ L+ L K + PL M R+ I K N +E ++L E+ +++ +R E
Sbjct: 29 FIILLALLRKFAWQPLMNIMKQREEHIATKSTRRKNDRQEAEKLLEEQRELMKQSRQEAQ 88
Query: 143 VALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQDIVN 202
+ + + E+ +A R + + + A +E++KE+ + AL Q+A+LS I +
Sbjct: 89 ALIENAASLAEEQKEQIVASARAEAERVKEAAKKEIEREKEQAMAALREQVASLSVLIAS 148
Query: 203 KVL 205
KV+
Sbjct: 149 KVI 151
>ATPF_BACPF (P22481) ATP synthase B chain (EC 3.6.3.14)
Length = 163
Score = 55.5 bits (132), Expect = 9e-08
Identities = 32/126 (25%), Positives = 61/126 (48%)
Query: 80 VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARA 139
++ F L+F L K PL M+ R+ I +++S ++ + + L AR
Sbjct: 14 LLAFSVLLFFLSKFALKPLLGIMEKREQMINEQISSAEKNRKDSEAFIAEQRQALEQARM 73
Query: 140 EIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQD 199
E + KK ++ + ++ + R + + A+A ++++KE+ V AL Q+A LS
Sbjct: 74 EANEIIQNAKKLSEQQGQDIVKAARNDAERIKESAVAEIQREKEQAVSALREQVAGLSVL 133
Query: 200 IVNKVL 205
I KV+
Sbjct: 134 IATKVI 139
>ATPF_PROMO (P21904) ATP synthase B chain, sodium ion specific (EC
3.6.3.15)
Length = 168
Score = 52.0 bits (123), Expect = 1e-06
Identities = 29/126 (23%), Positives = 57/126 (45%)
Query: 80 VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARA 139
++ FL LMF K + P+ +D R +I L E + +A ++++A+
Sbjct: 19 IINFLILMFFFKKYFQKPIAKVLDARKEKIANDLKQAEIDKEMAAKANGEAQGIVKSAKT 78
Query: 140 EIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQD 199
E L + +K+ E + E + + L+ A +EK KE+ K L ++ L+
Sbjct: 79 EANEMLLRAEKKADERKETILKEANTQREKMLKSAEVEIEKMKEQARKELQLEVTDLAVK 138
Query: 200 IVNKVL 205
+ K++
Sbjct: 139 LAEKMI 144
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.128 0.329
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,542,655
Number of Sequences: 164201
Number of extensions: 726594
Number of successful extensions: 5516
Number of sequences better than 10.0: 468
Number of HSP's better than 10.0 without gapping: 115
Number of HSP's successfully gapped in prelim test: 359
Number of HSP's that attempted gapping in prelim test: 4813
Number of HSP's gapped (non-prelim): 960
length of query: 209
length of database: 59,974,054
effective HSP length: 105
effective length of query: 104
effective length of database: 42,732,949
effective search space: 4444226696
effective search space used: 4444226696
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148405.2


