

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
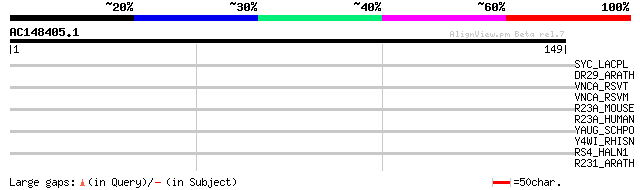
Query= AC148405.1 + phase: 0
(149 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SYC_LACPL (Q88YX9) Cysteinyl-tRNA synthetase (EC 6.1.1.16) (Cyst... 30 2.5
DR29_ARATH (Q9SZA7) Probable disease resistance protein At4g33300 30 2.5
VNCA_RSVT (Q00844) Major noncapsid protein (NCP) (20.5 kDa prote... 29 3.3
VNCA_RSVM (Q01209) Major noncapsid protein (NCP) (20.5 kDa prote... 29 4.3
R23A_MOUSE (P54726) UV excision repair protein RAD23 homolog A (... 29 4.3
R23A_HUMAN (P54725) UV excision repair protein RAD23 homolog A (... 29 4.3
YAUG_SCHPO (Q10169) Hypothetical protein C26A3.16 in chromosome I 28 5.7
Y4WI_RHISN (P55687) Hypothetical 59.0 kDa protein y4wI 28 5.7
RS4_HALN1 (Q9HQJ6) 30S ribosomal protein S4P 28 9.7
R231_ARATH (Q84L33) Putative DNA repair protein RAD23-1 (RAD23-l... 28 9.7
>SYC_LACPL (Q88YX9) Cysteinyl-tRNA synthetase (EC 6.1.1.16)
(Cysteine--tRNA ligase) (CysRS)
Length = 470
Score = 29.6 bits (65), Expect = 2.5
Identities = 22/72 (30%), Positives = 30/72 (41%), Gaps = 2/72 (2%)
Query: 76 KPEKISWPLVWGQFCLCYEGQKLVTETDYLRD-YGIKDGDQ-LRFIRHVSNVCNFQRKPK 133
KP +ISW WG + + V T YL D + I G Q L F H + + + K
Sbjct: 191 KPGEISWESPWGAGRPGWHIECSVMSTKYLGDTFDIHGGGQDLEFPHHENEIAQSEAKTG 250
Query: 134 KRTVNLKQHGRY 145
K N H +
Sbjct: 251 KTFANYWLHNGF 262
>DR29_ARATH (Q9SZA7) Probable disease resistance protein At4g33300
Length = 855
Score = 29.6 bits (65), Expect = 2.5
Identities = 32/111 (28%), Positives = 47/111 (41%), Gaps = 10/111 (9%)
Query: 25 IVDCTLPYD---KLPSQNLTLTVLKLDS-SSFQVEVAKTATVAVLKQAVEAAFRHKPEKI 80
+ DC P+D KL + T LD +SF+ T V+ K E F + E +
Sbjct: 265 VPDCNFPFDGARKLVILDDVWTTQALDRLTSFKFPGCTTLVVSRSK-LTEPKFTYDVEVL 323
Query: 81 SWPLVWGQFCLCYEGQKLVTE---TDYLRDYGIKDGDQLRF--IRHVSNVC 126
S FCLC GQK + D ++ + I+ LR + V+N C
Sbjct: 324 SEDEAISLFCLCAFGQKSIPLGFCKDLVKQHNIQSFSILRVLCLAQVANEC 374
>VNCA_RSVT (Q00844) Major noncapsid protein (NCP) (20.5 kDa
protein) (Protein S) (Stripe disease-specific protein)
Length = 178
Score = 29.3 bits (64), Expect = 3.3
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 8/69 (11%)
Query: 13 RRSLSIQLSSMIIVDCTLPYDKLP---SQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAV 69
+R++ + + ++ +D TL YD LP S N+TL LK V + +LK V
Sbjct: 5 QRTIEVSVGPIVGLDYTLLYDTLPETVSDNITLPDLKDPE-----RVTEDTKKLILKGCV 59
Query: 70 EAAFRHKPE 78
A+ H E
Sbjct: 60 YIAYHHPLE 68
>VNCA_RSVM (Q01209) Major noncapsid protein (NCP) (20.5 kDa
protein) (Protein S) (Stripe disease-specific protein)
Length = 178
Score = 28.9 bits (63), Expect = 4.3
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 8/69 (11%)
Query: 13 RRSLSIQLSSMIIVDCTLPYDKLP---SQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAV 69
+R++ + + ++ +D TL YD LP S N+TL LK V + +LK V
Sbjct: 5 QRTVEVSVGPIVGLDYTLLYDTLPETVSDNITLPDLKDPE-----RVTEDTKKLILKGCV 59
Query: 70 EAAFRHKPE 78
A+ H E
Sbjct: 60 YIAYHHPLE 68
>R23A_MOUSE (P54726) UV excision repair protein RAD23 homolog A
(mHR23A)
Length = 363
Score = 28.9 bits (63), Expect = 4.3
Identities = 23/73 (31%), Positives = 42/73 (57%), Gaps = 7/73 (9%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLV 99
+T+T+ L +F++ + TV VLK+ +EA + + ++P V GQ L Y G K++
Sbjct: 3 VTITLKTLQQQTFKIRMEPDETVKVLKEKIEA----EKGRDAFP-VAGQ-KLIYAG-KIL 55
Query: 100 TETDYLRDYGIKD 112
++ +RDY I +
Sbjct: 56 SDDVPIRDYHIDE 68
>R23A_HUMAN (P54725) UV excision repair protein RAD23 homolog A
(hHR23A)
Length = 363
Score = 28.9 bits (63), Expect = 4.3
Identities = 23/73 (31%), Positives = 42/73 (57%), Gaps = 7/73 (9%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLV 99
+T+T+ L +F++ + TV VLK+ +EA + + ++P V GQ L Y G K++
Sbjct: 3 VTITLKTLQQQTFKIRMEPDETVKVLKEKIEA----EKGRDAFP-VAGQ-KLIYAG-KIL 55
Query: 100 TETDYLRDYGIKD 112
++ +RDY I +
Sbjct: 56 SDDVPIRDYRIDE 68
>YAUG_SCHPO (Q10169) Hypothetical protein C26A3.16 in chromosome I
Length = 354
Score = 28.5 bits (62), Expect = 5.7
Identities = 20/82 (24%), Positives = 38/82 (45%), Gaps = 10/82 (12%)
Query: 39 NLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKL 98
N++LT+ + + V V ++V LK+A+ + E+ L Y G+ L
Sbjct: 3 NISLTIKAANDQKYAVTVDSESSVLALKEAIAPVADIEKERQR---------LIYAGRVL 53
Query: 99 VTETDYLRDYGIKDGDQLRFIR 120
E + L+ Y I+DG + ++
Sbjct: 54 KDE-ESLKTYKIQDGHSIHLVK 74
>Y4WI_RHISN (P55687) Hypothetical 59.0 kDa protein y4wI
Length = 513
Score = 28.5 bits (62), Expect = 5.7
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 7/39 (17%)
Query: 73 FRHKPEKISWPL--VWGQFCLCYEGQKLVTETDYLRDYG 109
+R+KPE WPL +W ++ + ++ D LRD+G
Sbjct: 103 YRYKPESAPWPLSIMWDEYM-----RDMLRHRDLLRDHG 136
>RS4_HALN1 (Q9HQJ6) 30S ribosomal protein S4P
Length = 170
Score = 27.7 bits (60), Expect = 9.7
Identities = 12/42 (28%), Positives = 28/42 (66%), Gaps = 2/42 (4%)
Query: 93 YEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKK 134
Y+G+++ E+D L YG+K+ ++L R S + +++R+ ++
Sbjct: 18 YQGERIAEESDLLSRYGLKNKEEL--WRAQSELRDYRREARR 57
>R231_ARATH (Q84L33) Putative DNA repair protein RAD23-1 (RAD23-like
protein 1) (AtRAD23-1)
Length = 371
Score = 27.7 bits (60), Expect = 9.7
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLV 99
+ LTV L S F++ V + T+ +K+ +E K ++P GQ L + G+ L
Sbjct: 1 MKLTVKTLKGSHFEIRVLPSDTIMAVKKNIE----DSQGKDNYPC--GQQLLIHNGKVLK 54
Query: 100 TETDYLRDYGIKDG 113
ET + + ++G
Sbjct: 55 DETSLVENKVTEEG 68
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,429,950
Number of Sequences: 164201
Number of extensions: 543462
Number of successful extensions: 1290
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1290
Number of HSP's gapped (non-prelim): 10
length of query: 149
length of database: 59,974,054
effective HSP length: 100
effective length of query: 49
effective length of database: 43,553,954
effective search space: 2134143746
effective search space used: 2134143746
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148405.1


