

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
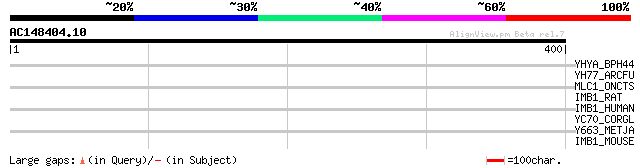
Query= AC148404.10 + phase: 0
(400 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YHYA_BPH44 (P15317) Hypothetical 65 kDa protein in hyaluronidase... 33 1.7
YH77_ARCFU (O28497) Hypothetical UPF0135 protein AF1777 31 4.9
MLC1_ONCTS (P17640) Pro-MCH 1 precursor [Contains: Neuropeptide-... 31 4.9
IMB1_RAT (P52296) Importin beta-1 subunit (Karyopherin beta-1 su... 31 4.9
IMB1_HUMAN (Q14974) Importin beta-1 subunit (Karyopherin beta-1 ... 31 6.4
YC70_CORGL (P42531) Hypothetical protein Cgl1270/cg1434 30 8.4
Y663_METJA (Q58077) Hypothetical protein MJ0663 30 8.4
IMB1_MOUSE (P70168) Importin beta-1 subunit (Karyopherin beta-1 ... 30 8.4
>YHYA_BPH44 (P15317) Hypothetical 65 kDa protein in hyaluronidase
region
Length = 593
Score = 32.7 bits (73), Expect = 1.7
Identities = 28/85 (32%), Positives = 39/85 (44%), Gaps = 9/85 (10%)
Query: 14 FFIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRV-FNFDVAKI 72
F +F +FT A ++S+S I +L A + +WDLN + FN D A I
Sbjct: 389 FRTLFAKNIFTTSVQAVTTSASK------ITGGVLSATNGASRWDLNSANIDFNRD-ATI 441
Query: 73 RFGTSQNYLFRIGSSKNNFTVKFSD 97
F + N L R S N V FS+
Sbjct: 442 NFNSKNNALVR-KSGTNTAFVHFSN 465
>YH77_ARCFU (O28497) Hypothetical UPF0135 protein AF1777
Length = 245
Score = 31.2 bits (69), Expect = 4.9
Identities = 14/36 (38%), Positives = 24/36 (65%), Gaps = 1/36 (2%)
Query: 290 ILQDRLSGLIKANIKAYAGVKFPLELERDVGNNATL 325
I++ R+ L+++N+ YA PL+ R+VGNNA +
Sbjct: 77 IVKRRIEFLLRSNLSLYAA-HLPLDAHREVGNNAVI 111
>MLC1_ONCTS (P17640) Pro-MCH 1 precursor [Contains:
Neuropeptide-glutamic acid-valine (NEV) (Neuropeptide
E-V); Melanin-concentrating hormone (MCH)]
Length = 132
Score = 31.2 bits (69), Expect = 4.9
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 3/65 (4%)
Query: 336 SVERVWFEVMARVEDSRLKPLSIK--KVKPFIESDSVSWANLMSNLSYTKLRPVLLPPE- 392
++E+ + + RVE S P S++ K + +DS W NL L + KLR P+
Sbjct: 34 ALEQDTLDSLLRVEVSENSPDSVRGRSSKIVLLADSGLWMNLNRGLPFYKLRAAAAGPDR 93
Query: 393 ALTLD 397
ALTLD
Sbjct: 94 ALTLD 98
>IMB1_RAT (P52296) Importin beta-1 subunit (Karyopherin beta-1
subunit) (Nuclear factor P97)
Length = 875
Score = 31.2 bits (69), Expect = 4.9
Identities = 21/70 (30%), Positives = 32/70 (45%)
Query: 329 PDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVL 388
PDWR R + + ++ E ++LKPL I+ + IE + ++T R
Sbjct: 377 PDWRYRDAAVMAFGSILEGPEPNQLKPLVIQAMPTLIELMKDPSVVVRDTTAWTVGRICE 436
Query: 389 LPPEALTLDV 398
L PEA DV
Sbjct: 437 LLPEAAINDV 446
>IMB1_HUMAN (Q14974) Importin beta-1 subunit (Karyopherin beta-1
subunit) (Nuclear factor P97) (Importin 90)
Length = 876
Score = 30.8 bits (68), Expect = 6.4
Identities = 22/70 (31%), Positives = 32/70 (45%)
Query: 329 PDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVL 388
PDWR R + + ++ E S+LKPL I+ + IE + ++T R
Sbjct: 378 PDWRYRDAAVMAFGCILEGPEPSQLKPLVIQAMPTLIELMKDPSVVVRDTAAWTVGRICE 437
Query: 389 LPPEALTLDV 398
L PEA DV
Sbjct: 438 LLPEAAINDV 447
>YC70_CORGL (P42531) Hypothetical protein Cgl1270/cg1434
Length = 491
Score = 30.4 bits (67), Expect = 8.4
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Query: 164 IVGKGITVEVR-RAREISFYYQSDLDLQRNGSVICSNQKNEFWPFLQSMCVPLIPIRIIG 222
IV +G +E+ RA +I Q + + + + +C N N + S V +IPI ++G
Sbjct: 101 IVVEGTVIEITMRATKIRTIAQETVIIPNSTAKVCINNSNNW-----SRAVVVIPIPMLG 155
Query: 223 SASLIAYVARN 233
S ++ +AR+
Sbjct: 156 SENITDVIARS 166
>Y663_METJA (Q58077) Hypothetical protein MJ0663
Length = 494
Score = 30.4 bits (67), Expect = 8.4
Identities = 18/45 (40%), Positives = 28/45 (62%), Gaps = 3/45 (6%)
Query: 337 VERVWFEVMARVEDSRLKPLSIKKVKPFIESDSV---SWANLMSN 378
VE V +++++ E+ +LKP SIK++K F E+ V SW SN
Sbjct: 273 VESVREKLLSKTENIQLKPKSIKELKEFFENLDVKNSSWIYKNSN 317
>IMB1_MOUSE (P70168) Importin beta-1 subunit (Karyopherin beta-1
subunit) (Nuclear factor P97) (Pore targeting complex 97
kDa subunit) (PTAC97) (SCG)
Length = 876
Score = 30.4 bits (67), Expect = 8.4
Identities = 21/70 (30%), Positives = 32/70 (45%)
Query: 329 PDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVL 388
PDWR R + + ++ E ++LKPL I+ + IE + ++T R
Sbjct: 378 PDWRYRDAAVVAFGSILEGPEPNQLKPLVIQAMPTLIELMKDPSVVVRDTTAWTVGRICE 437
Query: 389 LPPEALTLDV 398
L PEA DV
Sbjct: 438 LLPEAAINDV 447
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,365,448
Number of Sequences: 164201
Number of extensions: 1783442
Number of successful extensions: 4120
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 4120
Number of HSP's gapped (non-prelim): 8
length of query: 400
length of database: 59,974,054
effective HSP length: 112
effective length of query: 288
effective length of database: 41,583,542
effective search space: 11976060096
effective search space used: 11976060096
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148404.10


