

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.3 - phase: 0
(301 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
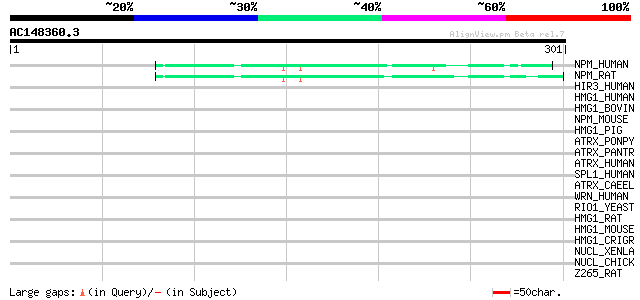
Score E
Sequences producing significant alignments: (bits) Value
NPM_HUMAN (P06748) Nucleophosmin (NPM) (Nucleolar phosphoprotein... 45 2e-04
NPM_RAT (P13084) Nucleophosmin (NPM) (Nucleolar phosphoprotein B... 44 6e-04
HIR3_HUMAN (Q9BW71) HIRA-interacting protein 3 42 0.001
HMG1_HUMAN (P09429) High mobility group protein 1 (HMG-1) 42 0.002
HMG1_BOVIN (P10103) High mobility group protein 1 (HMG-1) 42 0.002
NPM_MOUSE (Q61937) Nucleophosmin (NPM) (Nucleolar phosphoprotein... 42 0.002
HMG1_PIG (P12682) High mobility group protein 1 (HMG-1) 42 0.002
ATRX_PONPY (Q7YQM3) Transcriptional regulator ATRX (X-linked hel... 41 0.004
ATRX_PANTR (Q7YQM4) Transcriptional regulator ATRX (X-linked hel... 41 0.004
ATRX_HUMAN (P46100) Transcriptional regulator ATRX (X-linked hel... 41 0.004
SPL1_HUMAN (Q14515) SPARC-like protein 1 precursor (High endothe... 40 0.005
ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog (X-li... 40 0.007
WRN_HUMAN (Q14191) Werner syndrome helicase 40 0.009
RIO1_YEAST (Q12196) RIO1 protein 40 0.009
HMG1_RAT (P63159) High mobility group protein 1 (HMG-1) (Amphote... 39 0.012
HMG1_MOUSE (P63158) High mobility group protein 1 (HMG-1) 39 0.012
HMG1_CRIGR (P07156) High mobility group protein 1 (HMG-1) (Fragm... 39 0.012
NUCL_XENLA (P20397) Nucleolin (Protein C23) 39 0.016
NUCL_CHICK (P15771) Nucleolin (Protein C23) 39 0.016
Z265_RAT (O35986) Zinc finger protein 265 (Zinc finger, splicing) 39 0.021
>NPM_HUMAN (P06748) Nucleophosmin (NPM) (Nucleolar phosphoprotein
B23) (Numatrin) (Nucleolar protein NO38)
Length = 294
Score = 45.1 bits (105), Expect = 2e-04
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>NPM_RAT (P13084) Nucleophosmin (NPM) (Nucleolar phosphoprotein B23)
(Numatrin) (Nucleolar protein NO38)
Length = 292
Score = 43.5 bits (101), Expect = 6e-04
Identities = 54/227 (23%), Positives = 87/227 (37%), Gaps = 30/227 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEDEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
M G APG + + +K + +DD++DED+E+DE D + E+
Sbjct: 137 GMSGKRSAPG----GGNKVPQKKVKLDEDDDEDDEDDEDDEDDDDDDF-------DEEET 185
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ K ++K + DT K+A K + + GK S SK +
Sbjct: 186 EEKVPVKKSVRDTPA---KNAQKSNQN------GKDLKPSTPRSKGQ 223
>HIR3_HUMAN (Q9BW71) HIRA-interacting protein 3
Length = 556
Score = 42.4 bits (98), Expect = 0.001
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 22/106 (20%)
Query: 163 TKPPPVPKDRTPGEPFVAKS-----SKEAEME---KLLKSMEGMPGAPGMKMYSRDDLMK 214
T+ PV K + PG+ V++ S+E+E E + K +EG G +K ++ +
Sbjct: 171 TRKKPVVKKQAPGKASVSRKQAREESEESEAEPVQRTAKKVEGNKGTKSLKESEQES--E 228
Query: 215 KNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIR 260
+ A+ ++ E+E EEE+ K ++ EKGDWK + R
Sbjct: 229 EEILAQKKEQREEEVEEEE------------KEEDEEKGDWKPRTR 262
>HMG1_HUMAN (P09429) High mobility group protein 1 (HMG-1)
Length = 214
Score = 42.0 bits (97), Expect = 0.002
Identities = 23/66 (34%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK E+E+D+EDE+E
Sbjct: 139 DKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEEDEEDEEDEEE 198
Query: 231 EEDEAD 236
EEDE D
Sbjct: 199 EEDEED 204
>HMG1_BOVIN (P10103) High mobility group protein 1 (HMG-1)
Length = 214
Score = 42.0 bits (97), Expect = 0.002
Identities = 23/66 (34%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK E+E+D+EDE+E
Sbjct: 139 DKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEEDEEDEEDEEE 198
Query: 231 EEDEAD 236
EEDE D
Sbjct: 199 EEDEED 204
>NPM_MOUSE (Q61937) Nucleophosmin (NPM) (Nucleolar phosphoprotein
B23) (Numatrin) (Nucleolar protein NO38)
Length = 292
Score = 41.6 bits (96), Expect = 0.002
Identities = 54/227 (23%), Positives = 87/227 (37%), Gaps = 30/227 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEDEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
M G APG ++ K ++EDDDED++++ED+ D + E+
Sbjct: 137 GMSGKRSAPGGG--NKVPQKKVKLDEDDEDDDEDDEDDEDDDD---------DDFDEEET 185
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ K ++K + DT K+A K + + GK S SK +
Sbjct: 186 EEKVPVKKSVRDTPA---KNAQKSNQN------GKDLKPSTPRSKGQ 223
>HMG1_PIG (P12682) High mobility group protein 1 (HMG-1)
Length = 214
Score = 41.6 bits (96), Expect = 0.002
Identities = 23/66 (34%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK E+E+D+EDE+E
Sbjct: 139 DKHPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEEDEEDEEDEEE 198
Query: 231 EEDEAD 236
EEDE D
Sbjct: 199 EEDEED 204
>ATRX_PONPY (Q7YQM3) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP)
Length = 2492
Score = 40.8 bits (94), Expect = 0.004
Identities = 45/199 (22%), Positives = 91/199 (45%), Gaps = 35/199 (17%)
Query: 109 SEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPV 168
S+G+ E KT + +EV G + V+ D D + + +++S++ + + P
Sbjct: 1348 SDGESGEEKKTKPKEHKEVKGRNRRKVSS---EDSEDSDFQESGVSEEVSESEDEQRP-- 1402
Query: 169 PKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE 228
RT +S+K+AE+E+ +S + +K+ K+ E E++ E+E
Sbjct: 1403 ---RT-------RSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEEEKEEE 1452
Query: 229 DEEEDEADFPSKLGKILKSKENEKGDWK------QKIRKGIVDTSTTLKKHATKVSNHIQ 282
+EEE+E + + +E+E D K +KIRK + D K T+ N ++
Sbjct: 1453 EEEEEEEE---------EEEEDENDDSKSPGKGRKKIRKILKD-----DKLRTETQNALK 1498
Query: 283 RWWKGKKTTSSKKNSKSEL 301
+ +K + ++ + +L
Sbjct: 1499 EEEERRKRIAEREREREKL 1517
>ATRX_PANTR (Q7YQM4) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP)
Length = 2492
Score = 40.8 bits (94), Expect = 0.004
Identities = 45/199 (22%), Positives = 91/199 (45%), Gaps = 35/199 (17%)
Query: 109 SEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPV 168
S+G+ E KT + +EV G + V+ D D + + +++S++ + + P
Sbjct: 1348 SDGESGEEKKTKPKEHKEVKGRNRRKVSS---EDSEDSDFQESGVSEEVSESEDEQRP-- 1402
Query: 169 PKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE 228
RT +S+K+AE+E+ +S + +K+ K+ E E++ E+E
Sbjct: 1403 ---RT-------RSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEEEKEEE 1452
Query: 229 DEEEDEADFPSKLGKILKSKENEKGDWK------QKIRKGIVDTSTTLKKHATKVSNHIQ 282
+EEE+E + + +E+E D K +KIRK + D K T+ N ++
Sbjct: 1453 EEEEEEEE---------EEEEDENDDSKSPGKGRKKIRKILKD-----DKLRTETQNALK 1498
Query: 283 RWWKGKKTTSSKKNSKSEL 301
+ +K + ++ + +L
Sbjct: 1499 EEEERRKRIAEREREREKL 1517
>ATRX_HUMAN (P46100) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP) (Znf-HX)
Length = 2492
Score = 40.8 bits (94), Expect = 0.004
Identities = 45/199 (22%), Positives = 91/199 (45%), Gaps = 35/199 (17%)
Query: 109 SEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPV 168
S+G+ E KT + +EV G + V+ D D + + +++S++ + + P
Sbjct: 1348 SDGESGEEKKTKPKEHKEVKGRNRRKVSS---EDSEDSDFQESGVSEEVSESEDEQRP-- 1402
Query: 169 PKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE 228
RT +S+K+AE+E+ +S + +K+ K+ E E++ E+E
Sbjct: 1403 ---RT-------RSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEEEKEEE 1452
Query: 229 DEEEDEADFPSKLGKILKSKENEKGDWK------QKIRKGIVDTSTTLKKHATKVSNHIQ 282
+EEE+E + + +E+E D K +KIRK + D K T+ N ++
Sbjct: 1453 EEEEEEEE---------EEEEDENDDSKSPGKGRKKIRKILKD-----DKLRTETQNALK 1498
Query: 283 RWWKGKKTTSSKKNSKSEL 301
+ +K + ++ + +L
Sbjct: 1499 EEEERRKRIAEREREREKL 1517
>SPL1_HUMAN (Q14515) SPARC-like protein 1 precursor (High
endothelial venule protein) (Hevin) (MAST 9)
Length = 664
Score = 40.4 bits (93), Expect = 0.005
Identities = 34/197 (17%), Positives = 87/197 (43%), Gaps = 11/197 (5%)
Query: 86 EADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPD 145
+ D + ++ L+++E SE Q + + V ++D++ E + + +
Sbjct: 104 DGDLSVNLEYAPSEGTLDIKEDMSEPQEKKLSENTDFLAPGVSSFTDSNQQESITKREEN 163
Query: 146 IDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMK 205
+ NY L+++ + +P ++ +E E E G G
Sbjct: 164 QEQPRNYSHHQLNRSSKHSQGLRDQGNQEQDPNISNGEEEEEKEP------GEVGTHNDN 217
Query: 206 MYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVD 265
+ +L +++ ++ E+D+ D+ +E+D P+++ K ++ E ++G+ +Q+
Sbjct: 218 QERKTELPREHANSKQEEDNTQSDDILEESDQPTQVSK-MQEDEFDQGNQEQEDN----S 272
Query: 266 TSTTLKKHATKVSNHIQ 282
+ +++A+ V+ HIQ
Sbjct: 273 NAEMEEENASNVNKHIQ 289
>ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog
(X-linked nuclear protein-1)
Length = 1359
Score = 40.0 bits (92), Expect = 0.007
Identities = 28/122 (22%), Positives = 54/122 (43%), Gaps = 1/122 (0%)
Query: 180 AKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPS 239
AKS E++ + + + ++ KK + +ED+D DE+ E+
Sbjct: 86 AKSESESDESDEEEDRKKSKSKKKVDQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSK 145
Query: 240 KLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSS-KKNSK 298
K K S+ +E+ + ++K++K + ++KK A + KK+ KK +K
Sbjct: 146 KTKKQTSSESSEESEEERKVKKSKKNKEKSVKKRAETSEESDEDEKPSKKSKKGLKKKAK 205
Query: 299 SE 300
SE
Sbjct: 206 SE 207
Score = 35.4 bits (80), Expect = 0.17
Identities = 37/137 (27%), Positives = 59/137 (43%), Gaps = 24/137 (17%)
Query: 164 KPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENED 223
KPPP + P + A SS+E + ++ E P K R A++E
Sbjct: 48 KPPP---KKRPAKKRKASSSEEDDDDE-----EESPRKSSKKSRKR---------AKSES 90
Query: 224 DDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQR 283
+ ++ DEEED SK K + K+ EK K+ + S ++ +K +
Sbjct: 91 ESDESDEEEDRKKSKSK--KKVDQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSK--- 145
Query: 284 WWKGKKTTSSKKNSKSE 300
K KK TSS+ + +SE
Sbjct: 146 --KTKKQTSSESSEESE 160
Score = 34.3 bits (77), Expect = 0.39
Identities = 34/136 (25%), Positives = 57/136 (41%), Gaps = 18/136 (13%)
Query: 181 KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD---- 236
K SK+ + + K E AP K + K + +E + DE+EEE E+
Sbjct: 219 KKSKKKSKKVVKKESESEDEAPEKKKTEKRKRSKTSSEESSESEKSDEEEEEKESSPKPK 278
Query: 237 --FPSKLGKILKSKENEKGDWK---QKIRKGIV--------DTSTTLKKHATKVSNHIQR 283
P + K+ +E+E+ D + QK ++G V + + A+ V + +
Sbjct: 279 KKKPLAVKKLSSDEESEESDVEVLPQKKKRGAVTLISDSEDEKDQKSESEASDVEEKVSK 338
Query: 284 WWKGKKTTSSKKNSKS 299
K KK SS+ S S
Sbjct: 339 -KKAKKQESSESGSDS 353
Score = 32.7 bits (73), Expect = 1.1
Identities = 48/207 (23%), Positives = 80/207 (38%), Gaps = 19/207 (9%)
Query: 103 ELEEHDSEGQCNSECKTVERACQEVMG-YSDTDVAEYLYSSKPDIDSLTNYLCKDLSKAC 161
E ++ + E S K+ +RA E SD + SK +D K K+
Sbjct: 66 EDDDDEEESPRKSSKKSRKRAKSESESDESDEEEDRKKSKSKKKVDQ------KKKEKSK 119
Query: 162 NTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAEN 221
+ +D E KS K+++ K S E + + + K+ +
Sbjct: 120 KKRTTSSSEDEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKSKKNKEKSVKKR 179
Query: 222 EDDDEDEDEEEDEADFPSKLGKI-LKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNH 280
+ E+ DE+E PSK K LK K + + + + K + + KK K S
Sbjct: 180 AETSEESDEDEK----PSKKSKKGLKKKAKSESESESEDEKEVKKSKKKSKKVVKKESES 235
Query: 281 IQRWWKGKKT-------TSSKKNSKSE 300
+ KKT TSS+++S+SE
Sbjct: 236 EDEAPEKKKTEKRKRSKTSSEESSESE 262
>WRN_HUMAN (Q14191) Werner syndrome helicase
Length = 1432
Score = 39.7 bits (91), Expect = 0.009
Identities = 36/161 (22%), Positives = 75/161 (46%), Gaps = 24/161 (14%)
Query: 83 KKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSS 142
+ +E ++ +DI E L++ E S+ + S+ ++ + + + + Y+ S
Sbjct: 385 ENMERACLMSLDITEH--ELQILEQQSQEEYLSDI--AYKSTEHLSPNDNENDTSYVIES 440
Query: 143 KPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAP 202
D++ + K LS P D +V +S ++ EME +LKS+E +
Sbjct: 441 DEDLEM---EMLKHLS----------PNDNENDTSYVIESDEDLEME-MLKSLENLNSGT 486
Query: 203 GMKMYSRDDLMKKNFGAENEDDDEDED------EEEDEADF 237
+S+ M++N G ++++ED++ EE+D+ DF
Sbjct: 487 VEPTHSKCLKMERNLGLPTKEEEEDDENEANEGEEDDDKDF 527
>RIO1_YEAST (Q12196) RIO1 protein
Length = 484
Score = 39.7 bits (91), Expect = 0.009
Identities = 29/86 (33%), Positives = 45/86 (51%), Gaps = 11/86 (12%)
Query: 210 DDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTT 269
DD + G+E +D++E E ++DE K+LK K++E D K+ ++ D
Sbjct: 410 DDEEDGSSGSEEDDEEEGEYYDDDEP-------KVLKGKKHEDKDLKKLRKQEAKDAKR- 461
Query: 270 LKKHATKVSNHIQRWWKGKKTTSSKK 295
+K TKV HI++ K K T SKK
Sbjct: 462 -EKRKTKVKKHIKK--KLVKKTKSKK 484
>HMG1_RAT (P63159) High mobility group protein 1 (HMG-1)
(Amphoterin) (Heparin-binding protein p30)
Length = 214
Score = 39.3 bits (90), Expect = 0.012
Identities = 21/66 (31%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK ++E+D+EDE+E
Sbjct: 139 DKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEDDEEDEEDEEE 198
Query: 231 EEDEAD 236
EE+E D
Sbjct: 199 EEEEED 204
>HMG1_MOUSE (P63158) High mobility group protein 1 (HMG-1)
Length = 214
Score = 39.3 bits (90), Expect = 0.012
Identities = 21/66 (31%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK ++E+D+EDE+E
Sbjct: 139 DKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEDDEEDEEDEEE 198
Query: 231 EEDEAD 236
EE+E D
Sbjct: 199 EEEEED 204
>HMG1_CRIGR (P07156) High mobility group protein 1 (HMG-1)
(Fragment)
Length = 180
Score = 39.3 bits (90), Expect = 0.012
Identities = 21/66 (31%), Positives = 34/66 (50%)
Query: 171 DRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
D+ P E AK ++ E + +G P A + + KK ++E+D+EDE+E
Sbjct: 105 DKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEDDEEDEEDEEE 164
Query: 231 EEDEAD 236
EE+E D
Sbjct: 165 EEEEED 170
>NUCL_XENLA (P20397) Nucleolin (Protein C23)
Length = 650
Score = 38.9 bits (89), Expect = 0.016
Identities = 26/85 (30%), Positives = 38/85 (44%), Gaps = 6/85 (7%)
Query: 168 VPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDED 227
V K TP + K + E + E M AP +K K AE +D++ED
Sbjct: 137 VAKKPTPAKKPAGKKQESEEEDDEESEDEPMEVAPALKG------KKTAQAAEEDDEEED 190
Query: 228 EDEEEDEADFPSKLGKILKSKENEK 252
+D+EED+ D + G + KE K
Sbjct: 191 DDDEEDDDDEEEQQGSAKRKKEMPK 215
Score = 34.7 bits (78), Expect = 0.30
Identities = 24/96 (25%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Query: 159 KACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFG 218
KA KP P K + + +E+E E ME P G K + +
Sbjct: 135 KAVAKKPTPAKKPAGKKQESEEEDDEESEDEP----MEVAPALKGKKTAQAAEEDDEEED 190
Query: 219 AENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
++E+DD+DE+E++ A ++ K + + K D
Sbjct: 191 DDDEEDDDDEEEQQGSAKRKKEMPKTIPEAKKTKTD 226
Score = 31.2 bits (69), Expect = 3.3
Identities = 27/113 (23%), Positives = 44/113 (38%), Gaps = 11/113 (9%)
Query: 168 VPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDED 227
VP +TP + K A K PG G KK E EDD ++
Sbjct: 45 VPVKKTPAK-------KTATPAKATPGKAATPGKKGATPAKNGKQAKKQESEEEEDDSDE 97
Query: 228 EDEEE----DEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATK 276
E E++ ++ + K +S+E++ + + + K + T KK A K
Sbjct: 98 EAEDQKPIKNKPVAKKAVAKKEESEEDDDDEDESEEEKAVAKKPTPAKKPAGK 150
Score = 30.4 bits (67), Expect = 5.6
Identities = 26/77 (33%), Positives = 41/77 (52%), Gaps = 17/77 (22%)
Query: 182 SSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDE---ADFP 238
S +EAE +K +K+ K ++ + KK E+E+DD+DEDE E+E A P
Sbjct: 95 SDEEAEDQKPIKN----------KPVAKKAVAKKE---ESEEDDDDEDESEEEKAVAKKP 141
Query: 239 SKLGKIL-KSKENEKGD 254
+ K K +E+E+ D
Sbjct: 142 TPAKKPAGKKQESEEED 158
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 38.9 bits (89), Expect = 0.016
Identities = 22/73 (30%), Positives = 39/73 (53%)
Query: 167 PVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDE 226
P K TP + VA +K+A K+ GA K +++ +++ E++++DE
Sbjct: 71 PAKKAVTPAKKAVATPAKKAVAPSPKKAAVVGKGAKNGKNAKKEESEEEDEDDEDDEEDE 130
Query: 227 DEDEEEDEADFPS 239
DE+EE DE + P+
Sbjct: 131 DEEEESDEEEEPA 143
Score = 37.7 bits (86), Expect = 0.035
Identities = 58/239 (24%), Positives = 89/239 (36%), Gaps = 29/239 (12%)
Query: 24 AKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAENVCNLK 83
AKK A+K P K AK+ K SPKK + + + A+N N K
Sbjct: 58 AKKAATPAKKAATPAKKAVTP---AKKAVATPAKKAVAPSPKKAAV--VGKGAKNGKNAK 112
Query: 84 KVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSK 143
K E++ D ++ D E EE D E + K + + V S+
Sbjct: 113 KEESEEEDEDDEDDEEDEDEEEESDEEEEPAVPVKPAAKKSAAAVPAKKPAVVPAKQESE 172
Query: 144 P----DIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMP 199
D + + +A +T P PV K TP + AK+ E+E E+
Sbjct: 173 EEEEEDDEEEDEEDDESEDEAMDTTPAPVKKP-TPAKATPAKAKAESEDEE--------- 222
Query: 200 GAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQK 258
D+ + + +D++EDE+E EDE GK K N+ +K
Sbjct: 223 ----------DEEDEDEDEEDEDDEEEDEEESEDEKPVKEAPGKRKKEMANKSAPEAKK 271
>Z265_RAT (O35986) Zinc finger protein 265 (Zinc finger, splicing)
Length = 332
Score = 38.5 bits (88), Expect = 0.021
Identities = 53/248 (21%), Positives = 95/248 (37%), Gaps = 42/248 (16%)
Query: 81 NLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLY 140
N + + DWI +K + S +C E KT E + G T++ + L
Sbjct: 5 NFRVSDGDWICPD---KKCGNVNFARRTSCNRCGRE-KTTEAKMMKAGG---TEIGKTLA 57
Query: 141 SSKPDIDSLTNYLCKDLSKA----------CNTKPPPVPKDRT---------PGEPFVAK 181
+ S ++ CK S CNT ++RT ++ +
Sbjct: 58 EKSRGLFSANDWQCKTCSNVNWARRSECNMCNTPKYAKLEERTGYGGGFNERENVEYIER 117
Query: 182 SSKEAEMEKL---LKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDED-------EDEE 231
+ E ++ K G P + +D K E ED+DED EDE+
Sbjct: 118 EESDGEYDEFGRKKKKYRGKAVGPASILKEVED---KESEGEEEDEDEDLSKYKLDEDED 174
Query: 232 EDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTT 291
ED+AD SK L + E E + K+ R+ + ++ + +++ S+ + + +
Sbjct: 175 EDDADL-SKYN--LDASEEEDSNKKKSNRRSRSKSRSSHSRSSSRSSSPSSSRSRSRSRS 231
Query: 292 SSKKNSKS 299
S +S+S
Sbjct: 232 RSSSSSQS 239
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,065,513
Number of Sequences: 164201
Number of extensions: 1652156
Number of successful extensions: 16528
Number of sequences better than 10.0: 424
Number of HSP's better than 10.0 without gapping: 199
Number of HSP's successfully gapped in prelim test: 235
Number of HSP's that attempted gapping in prelim test: 12340
Number of HSP's gapped (non-prelim): 1600
length of query: 301
length of database: 59,974,054
effective HSP length: 110
effective length of query: 191
effective length of database: 41,911,944
effective search space: 8005181304
effective search space used: 8005181304
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC148360.3


