

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.12 + phase: 0 /pseudo
(102 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
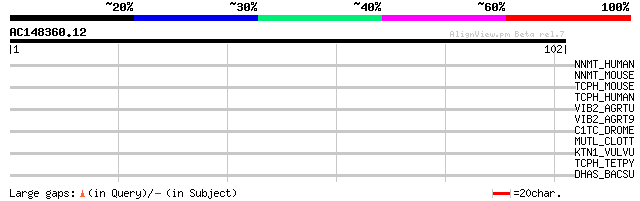
Score E
Sequences producing significant alignments: (bits) Value
NNMT_HUMAN (P40261) Nicotinamide N-methyltransferase (EC 2.1.1.1) 30 0.79
NNMT_MOUSE (O55239) Nicotinamide N-methyltransferase (EC 2.1.1.1) 29 2.3
TCPH_MOUSE (P80313) T-complex protein 1, eta subunit (TCP-1-eta)... 28 3.0
TCPH_HUMAN (Q99832) T-complex protein 1, eta subunit (TCP-1-eta)... 28 3.0
VIB2_AGRTU (P09776) VirB2 protein precursor 28 3.9
VIB2_AGRT9 (P05351) VirB2 protein precursor 28 3.9
C1TC_DROME (O96553) C-1-tetrahydrofolate synthase, cytoplasmic (... 28 3.9
MUTL_CLOTT (Q05491) MutL protein 28 5.1
KTN1_VULVU (O97961) Kinectin 27 6.7
TCPH_TETPY (P54409) T-complex protein 1, eta subunit (TCP-1-eta)... 27 8.7
DHAS_BACSU (Q04797) Aspartate-semialdehyde dehydrogenase (EC 1.2... 27 8.7
>NNMT_HUMAN (P40261) Nicotinamide N-methyltransferase (EC 2.1.1.1)
Length = 264
Score = 30.4 bits (67), Expect = 0.79
Identities = 12/31 (38%), Positives = 19/31 (60%)
Query: 19 KFGCRKSLKGDILEQISAAKALFQLISACSA 49
K C +KGD+L I + ++QL+SAC +
Sbjct: 47 KIFCLDGVKGDLLIDIGSGPTIYQLLSACES 77
>NNMT_MOUSE (O55239) Nicotinamide N-methyltransferase (EC 2.1.1.1)
Length = 264
Score = 28.9 bits (63), Expect = 2.3
Identities = 11/31 (35%), Positives = 20/31 (64%)
Query: 19 KFGCRKSLKGDILEQISAAKALFQLISACSA 49
K C ++KG++L I + ++QL+SAC +
Sbjct: 47 KIFCLGAVKGELLIDIGSGPTIYQLLSACES 77
>TCPH_MOUSE (P80313) T-complex protein 1, eta subunit (TCP-1-eta)
(CCT-eta)
Length = 544
Score = 28.5 bits (62), Expect = 3.0
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 36 AAKALFQLISACSASVG-GSKSIALLAPKAMKEVKSLVD 73
AAK L + + A VG G+ S+ LLA + +K+VK V+
Sbjct: 75 AAKTLVDIAKSQDAEVGDGTTSVTLLAAEFLKQVKPYVE 113
>TCPH_HUMAN (Q99832) T-complex protein 1, eta subunit (TCP-1-eta)
(CCT-eta) (HIV-1 Nef interacting protein)
Length = 543
Score = 28.5 bits (62), Expect = 3.0
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 36 AAKALFQLISACSASVG-GSKSIALLAPKAMKEVKSLVD 73
AAK L + + A VG G+ S+ LLA + +K+VK V+
Sbjct: 75 AAKTLVDIAKSQDAEVGDGTTSVTLLAAEFLKQVKPYVE 113
>VIB2_AGRTU (P09776) VirB2 protein precursor
Length = 121
Score = 28.1 bits (61), Expect = 3.9
Identities = 16/54 (29%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Query: 31 LEQISAAKALFQLISACSASVGGSK--SIALLAPKAMKEVKSLVDMILGFMSIC 82
L ++S + A+ ++IS+C+ S+GG+ SI+ P A + D +IC
Sbjct: 11 LNRLSLSNAMMRVISSCAPSLGGAMAWSISSCGPAAAQSAGGGTDPATMVNNIC 64
>VIB2_AGRT9 (P05351) VirB2 protein precursor
Length = 121
Score = 28.1 bits (61), Expect = 3.9
Identities = 16/54 (29%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Query: 31 LEQISAAKALFQLISACSASVGGSK--SIALLAPKAMKEVKSLVDMILGFMSIC 82
L ++S + A+ ++IS+C+ S+GG+ SI+ P A + D +IC
Sbjct: 11 LNRLSLSNAMMRVISSCAPSLGGAMAWSISSCGPAAAQSAGGGTDPATMVNNIC 64
>C1TC_DROME (O96553) C-1-tetrahydrofolate synthase, cytoplasmic
(C1-THF synthase) [Includes: Methylenetetrahydrofolate
dehydrogenase (EC 1.5.1.5); Methenyltetrahydrofolate
cyclohydrolase (EC 3.5.4.9); Formyltetrahydrofolate
synthetase (EC 6.3.4.3)]
Length = 968
Score = 28.1 bits (61), Expect = 3.9
Identities = 13/29 (44%), Positives = 19/29 (64%)
Query: 59 LLAPKAMKEVKSLVDMILGFMSICCSKIS 87
+L+P A ++VK L D G + IC SK+S
Sbjct: 873 VLSPAAEEKVKRLTDAGFGNLPICMSKVS 901
>MUTL_CLOTT (Q05491) MutL protein
Length = 462
Score = 27.7 bits (60), Expect = 5.1
Identities = 19/59 (32%), Positives = 30/59 (50%), Gaps = 2/59 (3%)
Query: 10 DVIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIAL-LAPKAMKE 67
D+ + S I G K+ + + EQ+ + F ACS++ GG K IA+ L P+ E
Sbjct: 32 DITTIESDIMVGFNKAYE-KLTEQLEGKEVNFVKKLACSSAAGGLKMIAIGLVPELTAE 89
>KTN1_VULVU (O97961) Kinectin
Length = 1330
Score = 27.3 bits (59), Expect = 6.7
Identities = 24/69 (34%), Positives = 33/69 (47%), Gaps = 6/69 (8%)
Query: 29 DILEQISAAKALF-----QLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICC 83
DILEQ A KAL Q+ + SASV + ++A K K++K D +
Sbjct: 573 DILEQNEALKALIQQFHSQIAAQTSASVLAEELHKVIAEKD-KQIKQTEDSLANEHDHLT 631
Query: 84 SKISEEKDL 92
SK E KD+
Sbjct: 632 SKEEELKDI 640
>TCPH_TETPY (P54409) T-complex protein 1, eta subunit (TCP-1-eta)
(CCT-eta)
Length = 558
Score = 26.9 bits (58), Expect = 8.7
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 36 AAKALFQLISACSASVG-GSKSIALLAPKAMKEVKSLVD 73
AAK L + A VG G+ S+ LLA + +KE K+ ++
Sbjct: 74 AAKTLVDIAKAQDDEVGDGTTSVCLLAGELLKESKNFIE 112
>DHAS_BACSU (Q04797) Aspartate-semialdehyde dehydrogenase (EC
1.2.1.11) (ASA dehydrogenase) (ASADH)
Length = 346
Score = 26.9 bits (58), Expect = 8.7
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Query: 21 GCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGF 78
G + + KG L A+ F+ ++ S GGS S A LAP+A+K ++D F
Sbjct: 44 GTKVTFKGQELTVQEASPESFEGVNIALFSAGGSVSQA-LAPEAVKRGAIVIDNTSAF 100
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.135 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,516,589
Number of Sequences: 164201
Number of extensions: 296809
Number of successful extensions: 774
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 771
Number of HSP's gapped (non-prelim): 11
length of query: 102
length of database: 59,974,054
effective HSP length: 78
effective length of query: 24
effective length of database: 47,166,376
effective search space: 1131993024
effective search space used: 1131993024
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148360.12


