

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148348.6 + phase: 0
(552 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
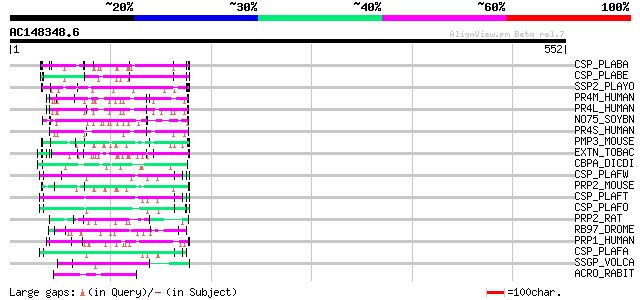
Score E
Sequences producing significant alignments: (bits) Value
CSP_PLABA (P23093) Circumsporozoite protein precursor (CS) 70 1e-11
CSP_PLABE (P06915) Circumsporozoite protein precursor (CS) 70 1e-11
SSP2_PLAYO (Q01443) Sporozoite surface protein 2 precursor 69 2e-11
PR4M_HUMAN (P10161) Basic salivary proline-rich protein 4 allele... 66 3e-10
PR4L_HUMAN (P10162) Basic salivary proline-rich protein 4 allele... 65 5e-10
NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75) (NGM-75) 65 5e-10
PR4S_HUMAN (P10163) Basic salivary proline-rich protein 4 allele... 64 1e-09
PMP3_MOUSE (P05143) Proline-rich protein MP-3 (Fragment) 62 4e-09
EXTN_TOBAC (P13983) Extensin precursor (Cell wall hydroxyproline... 61 7e-09
CBPA_DICDI (P35085) Calcium-binding protein 61 9e-09
CSP_PLAFW (P08307) Circumsporozoite protein precursor (CS) 60 1e-08
PRP2_MOUSE (P05142) Proline-rich protein MP-2 precursor 59 3e-08
CSP_PLAFT (P13814) Circumsporozoite protein precursor (CS) 58 6e-08
CSP_PLAFO (P19597) Circumsporozoite protein precursor (CS) 58 6e-08
PRP2_RAT (P10164) Acidic proline-rich protein PRP25 precursor (F... 58 7e-08
RB97_DROME (Q02926) Ribonucleoprotein RB97D 57 1e-07
PRP1_HUMAN (P04280) Basic salivary proline-rich protein 1 precur... 57 1e-07
CSP_PLAFA (P02893) Circumsporozoite protein precursor (CS) 57 1e-07
SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor ... 56 2e-07
ACRO_RABIT (P48038) Acrosin precursor (EC 3.4.21.10) 56 2e-07
>CSP_PLABA (P23093) Circumsporozoite protein precursor (CS)
Length = 347
Score = 70.5 bits (171), Expect = 1e-11
Identities = 45/111 (40%), Positives = 50/111 (44%), Gaps = 14/111 (12%)
Query: 75 QNPNFRPPQPPQNPNYRQQPPPQNPNFRPP-----PPPQNPNFRPPPPPQNPNFRQPIRQ 129
+N + P PP NPN PPP NPN PP PPP NPN PPP NPN P
Sbjct: 85 RNNKLKQPPPPPNPN---DPPPPNPNDPPPPNPNDPPPPNPN---DPPPPNPN--DPPPP 136
Query: 130 NPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPN-NQFQTPNVQEQAPPPPS 179
N N PP N N +P N N PN N PN + PP P+
Sbjct: 137 NANDPPPPNANDPAPPNANDPAPPNANDPAPPNANDPPPPNANDPPPPNPN 187
Score = 68.9 bits (167), Expect = 3e-11
Identities = 46/115 (40%), Positives = 49/115 (42%), Gaps = 15/115 (13%)
Query: 74 PQNPNFRPPQ-----PPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPP---PPQNPNFRQ 125
P NPN PP PP NPN PPP NPN PPP NPN PPP P PN
Sbjct: 95 PPNPNDPPPPNPNDPPPPNPN---DPPPPNPN---DPPPPNPNDPPPPNANDPPPPNAND 148
Query: 126 PIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPN-NQFQTPNVQEQAPPPPS 179
P N N P N N +P N N PN N PN + PP P+
Sbjct: 149 PAPPNANDPAPPNANDPAPPNANDPPPPNANDPPPPNPNDPAPPNANDPPPPNPN 203
Score = 63.9 bits (154), Expect = 1e-09
Identities = 51/148 (34%), Positives = 57/148 (38%), Gaps = 17/148 (11%)
Query: 42 HQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNF 101
++++QPP N + N P D P NPN PP NPN PPP N N
Sbjct: 87 NKLKQPPPPPNPNDPPPPNPNDPPPPNPNDPPPPNPN---DPPPPNPN---DPPPPNAND 140
Query: 102 RPPP---PPQNPNFRPPPPPQ-----NPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNG 153
PPP P PN P PP PN P N N PP PN N +P
Sbjct: 141 PPPPNANDPAPPNANDPAPPNANDPAPPNANDPPPPNANDPPPPNPNDPAPPNANDPPPP 200
Query: 154 NLNQFQNP---NNQFQTPNVQEQAPPPP 178
N N P NN P Q Q P P
Sbjct: 201 NPNDPAPPQGNNNPQPQPRPQPQPQPQP 228
Score = 61.6 bits (148), Expect = 5e-09
Identities = 38/99 (38%), Positives = 44/99 (44%), Gaps = 8/99 (8%)
Query: 84 PPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPP-----PPPQNPNFRQPIRQNPNFQPPNG 138
P N ++ +N + PPPP NPN PP PPP NPN P NPN PP
Sbjct: 72 PEGKKNEKKNKIERNNKLKQPPPPPNPNDPPPPNPNDPPPPNPN--DPPPPNPNDPPPPN 129
Query: 139 PNRGINQNQWNPQNGNLNQFQNPN-NQFQTPNVQEQAPP 176
PN N +P N N PN N PN + APP
Sbjct: 130 PNDPPPPNANDPPPPNANDPAPPNANDPAPPNANDPAPP 168
Score = 60.1 bits (144), Expect = 1e-08
Identities = 53/156 (33%), Positives = 59/156 (36%), Gaps = 18/156 (11%)
Query: 32 IPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQ-QDWNPQNPNFRPPQ-----PP 85
I N RL + ++ +N P P D P NPN PP PP
Sbjct: 60 IRNTVNRLLADAPEGKKNEKKNKIERNNKLKQPPPPPNPNDPPPPNPNDPPPPNPNDPPP 119
Query: 86 QNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPP----PQNPNFRQPIRQNPNFQPPNGPNR 141
NPN PPP NP PPPP N PPPP P PN P N N P N
Sbjct: 120 PNPN---DPPPPNP-NDPPPPNAND---PPPPNANDPAPPNANDPAPPNANDPAPPNAND 172
Query: 142 GINQNQWNPQNGNLNQFQNPN-NQFQTPNVQEQAPP 176
N +P N N PN N PN + APP
Sbjct: 173 PPPPNANDPPPPNPNDPAPPNANDPPPPNPNDPAPP 208
Score = 59.3 bits (142), Expect = 3e-08
Identities = 52/151 (34%), Positives = 55/151 (35%), Gaps = 22/151 (14%)
Query: 40 PPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP---QPPQNPNYRQQPPP 96
PP PP + N + P P D P NPN PP P PN PP
Sbjct: 94 PPPNPNDPPPPNPNDPPPPNPNDPPPPNPN-DPPPPNPNDPPPPNANDPPPPNANDPAPP 152
Query: 97 QNPNFRPP----PPPQNPNFRPP-----PPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQ 147
+ PP P P N N PP PPP NPN P N N PP PN
Sbjct: 153 NANDPAPPNANDPAPPNANDPPPPNANDPPPPNPN--DPAPPNANDPPPPNPNDPA---- 206
Query: 148 WNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P GN N P Q Q P Q Q P P
Sbjct: 207 --PPQGNNNPQPQPRPQPQ-PQPQPQPQPQP 234
Score = 57.0 bits (136), Expect = 1e-07
Identities = 49/140 (35%), Positives = 56/140 (40%), Gaps = 19/140 (13%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQ----HFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNP 88
PN PP+ PP + N + N NA P +P PN P PP NP
Sbjct: 128 PNPNDPPPPNANDPPPPNANDPAPPNANDPAPPNANDPAPPNANDPPPPNANDPPPP-NP 186
Query: 89 N-----YRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGI 143
N PPP NPN P PP N N +P P PQ QP + P QP P
Sbjct: 187 NDPAPPNANDPPPPNPN-DPAPPQGNNNPQPQPRPQPQPQPQP-QPQPQPQPQPRP---- 240
Query: 144 NQNQWNPQNGNLNQFQNPNN 163
Q PQ G N +N NN
Sbjct: 241 ---QPQPQPGGNNNNKNNNN 257
Score = 44.7 bits (104), Expect = 6e-04
Identities = 36/94 (38%), Positives = 40/94 (42%), Gaps = 11/94 (11%)
Query: 31 AIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQN--- 87
A PN PP+ PP + N + AP D P NPN P PPQ
Sbjct: 158 APPNANDPAPPNANDPPPPNANDPPPPNPNDPAPPN--ANDPPPPNPN--DPAPPQGNNN 213
Query: 88 --PNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQ 119
P R QP PQ P +P P PQ P RP P PQ
Sbjct: 214 PQPQPRPQPQPQ-PQPQPQPQPQ-PQPRPQPQPQ 245
>CSP_PLABE (P06915) Circumsporozoite protein precursor (CS)
Length = 339
Score = 70.1 bits (170), Expect = 1e-11
Identities = 44/114 (38%), Positives = 49/114 (42%), Gaps = 12/114 (10%)
Query: 75 QNPNFRPPQPPQNPNYRQQPPPQNPNFRPP-----PPPQNPNFRPPP---PPQNPNFRQP 126
+N + P PP NPN PPP NPN PP PPP NPN PPP P PN P
Sbjct: 85 RNNKLKQPPPPPNPN---DPPPPNPNDPPPPNPNDPPPPNPNDPPPPNANDPPPPNANDP 141
Query: 127 IRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPN-NQFQTPNVQEQAPPPPS 179
N N P N N +P N N PN N PN + PP P+
Sbjct: 142 APPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNANDPPPPNPN 195
Score = 65.9 bits (159), Expect = 3e-10
Identities = 52/148 (35%), Positives = 58/148 (39%), Gaps = 14/148 (9%)
Query: 34 NEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQ-QDWNPQNPNFRPPQPPQNPNYRQ 92
N RL P ++ N+ +N P P D P NPN PP NPN
Sbjct: 62 NTVNRLLPMLRRKKNEKKNEKIERNNKLKQPPPPPNPNDPPPPNPN---DPPPPNPN--- 115
Query: 93 QPPPQNPNFRPPPPPQNPNFRPPP---PPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWN 149
PPP NPN PPP N N PPP P PN P N N P N N +
Sbjct: 116 DPPPPNPN---DPPPPNANDPPPPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNAND 172
Query: 150 PQNGNLNQFQNPN-NQFQTPNVQEQAPP 176
P N N PN N PN + APP
Sbjct: 173 PAPPNANDPAPPNANDPPPPNPNDPAPP 200
Score = 55.8 bits (133), Expect = 3e-07
Identities = 47/139 (33%), Positives = 54/139 (38%), Gaps = 17/139 (12%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQ----HFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQN- 87
PN PP+ PP + N + N NA P +P PN P PP
Sbjct: 120 PNPNDPPPPNANDPPPPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNAN 179
Query: 88 ---PNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGIN 144
P PPP NPN P PP N N +P P PQ QP + P QP P
Sbjct: 180 DPAPPNANDPPPPNPN-DPAPPQGNNNPQPQPRPQPQPQPQP-QPQPQPQPQPRP----- 232
Query: 145 QNQWNPQNGNLNQFQNPNN 163
Q PQ G N +N NN
Sbjct: 233 --QPQPQPGGNNNNKNNNN 249
Score = 44.7 bits (104), Expect = 6e-04
Identities = 32/96 (33%), Positives = 37/96 (38%), Gaps = 7/96 (7%)
Query: 31 AIPNEYGRLPPHQVQQPPSDHNQ----HFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQ 86
A PN PP+ P + N + N NA P +P PN P PPQ
Sbjct: 142 APPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNANDPAPPNANDPPPPNPNDPAPPQ 201
Query: 87 NPNYRQQPPPQNPNFRPPPPPQ---NPNFRPPPPPQ 119
N Q P P +P P PQ P RP P PQ
Sbjct: 202 GNNNPQPQPRPQPQPQPQPQPQPQPQPQPRPQPQPQ 237
>SSP2_PLAYO (Q01443) Sporozoite surface protein 2 precursor
Length = 827
Score = 69.3 bits (168), Expect = 2e-11
Identities = 54/143 (37%), Positives = 62/143 (42%), Gaps = 16/143 (11%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPP-PQNP 99
P+ P + +N N+ N N P P NP NPN P P NPN P P NP
Sbjct: 312 PNNPNNPNNPNNP--NNPNNPNNPNN-PNNPNNPNNPN--NPNNPNNPNNPNNPNNPNNP 366
Query: 100 NF----RPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPN-RGINQNQ-WNPQNG 153
N P P NPN P NP R P R+NPN PN PN N N+ NP
Sbjct: 367 NNPNNPNNPNNPNNPNDPSNPNNPNPKKRNPKRRNPNKPKPNKPNPNKPNPNEPSNPNKP 426
Query: 154 NLNQFQNPNNQFQTPNVQEQAPP 176
N N+ NPN PN E + P
Sbjct: 427 NPNEPSNPNK----PNPNEPSNP 445
Score = 65.1 bits (157), Expect = 5e-10
Identities = 58/154 (37%), Positives = 67/154 (42%), Gaps = 27/154 (17%)
Query: 32 IPNEYGRLP--PHQVQQP--PSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQN 87
IPN+ P P + P P+D N N +N N P NP NPN P P N
Sbjct: 289 IPNKIPEKPSNPEEPVNPNDPNDPNNPNNPNNPNN-----PNNPNNPNNPN--NPNNPNN 341
Query: 88 PNYRQQPP-PQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQN 146
PN P P NPN P P NPN P P NPN NPN PN PN N N
Sbjct: 342 PNNPNNPNNPNNPN--NPNNPNNPN--NPNNPNNPN-------NPN--NPNNPNDPSNPN 388
Query: 147 QWNPQNGNLNQFQNPNN-QFQTPNVQEQAPPPPS 179
NP+ N + +NPN + PN + P PS
Sbjct: 389 NPNPKKRNPKR-RNPNKPKPNKPNPNKPNPNEPS 421
Score = 64.3 bits (155), Expect = 8e-10
Identities = 47/143 (32%), Positives = 59/143 (40%), Gaps = 17/143 (11%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPP-QNPNYRQQPPPQNP 99
P+ P + +N N+ N N P P NP NPN + P +NPN + +P NP
Sbjct: 357 PNNPNNPNNPNNP--NNPNNPNNPNN-PNDPSNPNNPNPKKRNPKRRNPN-KPKPNKPNP 412
Query: 100 NFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQ 159
N P P NPN P P NPN PN P+ PN+ NP N N+
Sbjct: 413 NKPNPNEPSNPNKPNPNEPSNPN-------KPNPNEPSNPNKPNPNEPSNPNKPNPNEPL 465
Query: 160 NPN-----NQFQTPNVQEQAPPP 177
NPN N+ PN P
Sbjct: 466 NPNEPSNPNEPSNPNAPSNPNEP 488
Score = 63.5 bits (153), Expect = 1e-09
Identities = 45/141 (31%), Positives = 57/141 (39%), Gaps = 8/141 (5%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPN 100
P+ ++ P N + N N P + NP PN P P NPN P NPN
Sbjct: 390 PNPKKRNPKRRNPNKPKPNKPNPNKPNPNEPSNPNKPN---PNEPSNPNKPNPNEPSNPN 446
Query: 101 FRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQN--GNLNQF 158
P P NPN P P NPN + N P+ PN N N+ + N N N+
Sbjct: 447 KPNPNEPSNPNKPNPNEPLNPNEPSNPNEPSNPNAPSNPNEPSNPNEPSNPNEPSNPNEP 506
Query: 159 QNPN---NQFQTPNVQEQAPP 176
NPN N + N E + P
Sbjct: 507 SNPNEPSNPKKPSNPNEPSNP 527
Score = 57.8 bits (138), Expect = 7e-08
Identities = 49/152 (32%), Positives = 64/152 (41%), Gaps = 13/152 (8%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQ 92
PNE +P + + + N + N P P + NP PN P P NPN
Sbjct: 417 PNEPSNPNKPNPNEPSNPNKPNPNEPSNPNKPN--PNEPSNPNKPN---PNEPLNPNEPS 471
Query: 93 QP-PPQNPNF-RPPPPPQNPNF-RPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQ-W 148
P P NPN P P NPN P P NPN + N + P+ PN N N+
Sbjct: 472 NPNEPSNPNAPSNPNEPSNPNEPSNPNEPSNPNEPSNPNEPSNPKKPSNPNEPSNPNEPL 531
Query: 149 NP-QNGNLNQFQNPN---NQFQTPNVQEQAPP 176
NP + N N+ NPN N + N +E + P
Sbjct: 532 NPNEPSNPNEPSNPNEPSNPEEPSNPKEPSNP 563
Score = 57.0 bits (136), Expect = 1e-07
Identities = 48/152 (31%), Positives = 64/152 (41%), Gaps = 14/152 (9%)
Query: 33 PNEYGRLPPHQVQQP--PSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNY 90
PN L P++ P PS+ N N + N P + NP P+ P P NPN
Sbjct: 459 PNPNEPLNPNEPSNPNEPSNPNAPSNPNEPSN-----PNEPSNPNEPS--NPNEPSNPNE 511
Query: 91 RQQPP-PQNPNFRPPP-PPQNPNFRPPP-PPQNPNFRQPIRQNPNFQPPNGPNRGINQNQ 147
P P NPN P P NPN P P NPN + N + P+ PN N +
Sbjct: 512 PSNPKKPSNPNEPSNPNEPLNPNEPSNPNEPSNPNEPSNPEEPSNPKEPSNPNEPSNPEE 571
Query: 148 WNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
NP+ + + P+N + N +E P PS
Sbjct: 572 PNPEEP--SNPKEPSNPEEPINPEELNPKEPS 601
Score = 54.7 bits (130), Expect = 6e-07
Identities = 49/152 (32%), Positives = 64/152 (41%), Gaps = 20/152 (13%)
Query: 33 PNEYGRLPPHQVQQP--PSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRP--PQPPQNP 88
PNE P++ P PS+ N+ N + N + P P NPN P P P NP
Sbjct: 485 PNEPSN--PNEPSNPNEPSNPNEPSNPNEPSNP--KKPSNPNEPSNPN-EPLNPNEPSNP 539
Query: 89 NYRQQP-PPQNPNFRP-PPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQN 146
N P P NP P P NPN P P+ PN +P NP + P+ P IN
Sbjct: 540 NEPSNPNEPSNPEEPSNPKEPSNPN--EPSNPEEPNPEEP--SNP--KEPSNPEEPINPE 593
Query: 147 QWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
+ NP+ + + NP N +E P P
Sbjct: 594 ELNPKEPSNPEESNPKEPI---NPEESNPKEP 622
Score = 51.2 bits (121), Expect = 7e-06
Identities = 46/116 (39%), Positives = 49/116 (41%), Gaps = 16/116 (13%)
Query: 68 PQQDWNPQNPNFRP-----PQPPQNPNYRQQP-PPQNPNFRPPPPPQNPNFRPPPPPQNP 121
PQ P PN P P+ P NPN P P NPN P P NPN P P NP
Sbjct: 281 PQIPIPPVIPNKIPEKPSNPEEPVNPNDPNDPNNPNNPN--NPNNPNNPN--NPNNPNNP 336
Query: 122 NFRQPIRQNPNFQPPNGPNRGINQNQWNPQN-GNLNQFQNPNNQFQTPNVQEQAPP 176
N P N N PN PN N N NP N N N NPNN N + + P
Sbjct: 337 N--NPNNPN-NPNNPNNPNNPNNPN--NPNNPNNPNNPNNPNNPNNPNNPNDPSNP 387
Score = 47.4 bits (111), Expect = 1e-04
Identities = 43/149 (28%), Positives = 58/149 (38%), Gaps = 9/149 (6%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQ 92
PNE + PS+ + N + N P++ NP+ P+ P+ P NP
Sbjct: 539 PNEPSNPNEPSNPEEPSNPKEPSNPNEPSNPEEPNPEEPSNPKEPS--NPEEPINPEELN 596
Query: 93 QPPPQNPNFRPPPPPQNPNFRPPPPPQNP--NFRQPIRQNPNFQPPNGPNRGINQNQWNP 150
P NP P P NP P P NP N I Q+ +P N N I P
Sbjct: 597 PKEPSNPEESNPKEPINPEESNPKEPINPEDNENPLIIQDEPIEPRNDSN-VIPILPIIP 655
Query: 151 QNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
Q GN P+N + P+ E P P+
Sbjct: 656 QKGN----NIPSNLPENPSDSEVEYPRPN 680
Score = 33.9 bits (76), Expect = 1.1
Identities = 39/159 (24%), Positives = 54/159 (33%), Gaps = 30/159 (18%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQ--------NPNFRPPQP 84
PNE P + PS+ + N N P++ NP+ NP P+
Sbjct: 563 PNEPSN-PEEPNPEEPSNPKEPSNPEEPINPEELNPKEPSNPEESNPKEPINPEESNPKE 621
Query: 85 PQNPNYRQQP---------PPQNPNFRPPPP--PQNPNFRPPPPPQNPNFRQPIRQNPNF 133
P NP + P P + N P P PQ N P P+NP+ + PN
Sbjct: 622 PINPEDNENPLIIQDEPIEPRNDSNVIPILPIIPQKGNNIPSNLPENPSDSEVEYPRPND 681
Query: 134 QPPNG----------PNRGINQNQWNPQNGNLNQFQNPN 162
N PN I NP G+ + P+
Sbjct: 682 NGENSNNTMKSKKNIPNEPIPSPGDNPYKGHEERIPKPH 720
>PR4M_HUMAN (P10161) Basic salivary proline-rich protein 4 allele M
(Salivary proline-rich protein Po) (Parotid o protein)
[Contains: Peptide P-D] (Fragment)
Length = 238
Score = 65.9 bits (159), Expect = 3e-10
Identities = 54/154 (35%), Positives = 63/154 (40%), Gaps = 19/154 (12%)
Query: 40 PPHQ---VQQPPSDHNQHFNHHNTQNAPGRFPQQDWN-----PQNPNFRPPQPPQNPNYR 91
PPH + PP NQ P R P Q N P P PPQ N
Sbjct: 42 PPHPGKPERPPPQGGNQSQGPPPHPGKPERPPPQGGNQSQGPPPTPGKPEGPPPQGGNQS 101
Query: 92 QQPPPQ--NPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWN 149
Q PPP P RPPP N + RPPPPP P P N + PP P +
Sbjct: 102 QGPPPHPGKPE-RPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQGPPPHPGKPEGP---P 157
Query: 150 PQNGNLNQ-FQNPNNQFQTPNVQE----QAPPPP 178
PQ GN ++ ++P + Q P QE Q PPPP
Sbjct: 158 PQEGNKSRSARSPPGKPQGPPQQEGNKPQGPPPP 191
Score = 53.9 bits (128), Expect = 1e-06
Identities = 46/127 (36%), Positives = 54/127 (42%), Gaps = 18/127 (14%)
Query: 65 GRFPQQDWNPQNPNFRP--PQ--PPQNPNYRQQPPPQNPNFRPPPPPQ--NPNFRPPPPP 118
GR PQ PQ P P PQ PPQ N Q PPP +P PPPQ N + PPP P
Sbjct: 8 GRRPQGGNQPQRPPPPPGKPQGPPPQGGNQSQGPPP-HPGKPERPPPQGGNQSQGPPPHP 66
Query: 119 QNPNFRQPIRQNPNFQPPNGPNR-------GINQNQWNPQNGNLNQFQNPNNQFQTPNVQ 171
P P N + PP P + G NQ+Q P + + P Q N
Sbjct: 67 GKPERPPPQGGNQSQGPPPTPGKPEGPPPQGGNQSQGPPPHPGKPERPPP----QGGNQS 122
Query: 172 EQAPPPP 178
+ PPPP
Sbjct: 123 HRPPPPP 129
Score = 51.2 bits (121), Expect = 7e-06
Identities = 49/142 (34%), Positives = 54/142 (37%), Gaps = 24/142 (16%)
Query: 47 PPSDHNQHFNHHNTQNAPGRFPQQDWN----PQNPNFRPPQPP-QNPNYRQQPPPQNPNF 101
PP NQ P R P Q N P P +P +PP Q N Q PPP +P
Sbjct: 94 PPQGGNQSQGPPPHPGKPERPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQGPPP-HPGK 152
Query: 102 RPPPPPQNPN-----FRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLN 156
PPPQ N PP PQ P Q P PP G +G P GN
Sbjct: 153 PEGPPPQEGNKSRSARSPPGKPQGP--PQQEGNKPQGPPPPGKPQGP-----PPPGGNPQ 205
Query: 157 QFQNPNNQFQTPNVQEQAPPPP 178
Q Q P P + Q PPPP
Sbjct: 206 QPQAP------PAGKPQGPPPP 221
Score = 49.3 bits (116), Expect = 3e-05
Identities = 42/145 (28%), Positives = 57/145 (38%), Gaps = 19/145 (13%)
Query: 37 GRLPP--HQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQP 94
GR P +Q Q+PP + Q P + Q P +P +PP + Q
Sbjct: 8 GRRPQGGNQPQRPPPPPGK------PQGPPPQGGNQSQGPPPHPGKPERPPPQGGNQSQG 61
Query: 95 PPQNPNFRPPPPPQ--NPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQN 152
PP +P PPPQ N + PPP P P P N + PP P + + PQ
Sbjct: 62 PPPHPGKPERPPPQGGNQSQGPPPTPGKPEGPPPQGGNQSQGPPPHPGK---PERPPPQG 118
Query: 153 GNLNQFQNPNNQFQTPNVQEQAPPP 177
GN + P P + + PPP
Sbjct: 119 GNQSHRPPP------PPGKPERPPP 137
Score = 48.9 bits (115), Expect = 3e-05
Identities = 36/101 (35%), Positives = 44/101 (42%), Gaps = 14/101 (13%)
Query: 40 PPHQVQQ---PPSDHNQHFNHHNTQNAPGRFPQQDWN-PQNPNFRPPQPPQNPNYRQQPP 95
PPH + PP + N+ + + P PQQ+ N PQ P PP PQ P PP
Sbjct: 147 PPHPGKPEGPPPQEGNKSRSARSPPGKPQGPPQQEGNKPQGPP--PPGKPQGP-----PP 199
Query: 96 PQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
P +P PP PPPPPQ +P R QPP
Sbjct: 200 PGGNPQQPQAPPAGKPQGPPPPPQG---GRPPRPAQGQQPP 237
Score = 46.6 bits (109), Expect = 2e-04
Identities = 41/119 (34%), Positives = 47/119 (39%), Gaps = 24/119 (20%)
Query: 40 PPHQ---VQQPPSDHNQHFNHHNTQNAPGRFPQQDWN--------PQNPNFRPPQP---- 84
PPH + PP NQ P R P Q N P P PPQ
Sbjct: 105 PPHPGKPERPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQGPPPHPGKPEGPPPQEGNKS 164
Query: 85 --PQNPNYRQQPPPQNPNFRP--PPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGP 139
++P + Q PPQ +P PPPP P PPPP NP +QP Q P P GP
Sbjct: 165 RSARSPPGKPQGPPQQEGNKPQGPPPPGKPQ-GPPPPGGNP--QQP--QAPPAGKPQGP 218
Score = 43.9 bits (102), Expect = 0.001
Identities = 37/104 (35%), Positives = 43/104 (40%), Gaps = 21/104 (20%)
Query: 84 PPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGI 143
PP P R+ P N RPPPPP P PPPQ N Q +P +P P +G
Sbjct: 2 PPGKPQGRR-PQGGNQPQRPPPPPGKPQ---GPPPQGGNQSQGPPPHPG-KPERPPPQGG 56
Query: 144 NQNQW-----------NPQNGNLNQFQNPNNQFQTPNVQEQAPP 176
NQ+Q PQ GN +Q P TP E PP
Sbjct: 57 NQSQGPPPHPGKPERPPPQGGNQSQGPPP-----TPGKPEGPPP 95
Score = 42.7 bits (99), Expect = 0.002
Identities = 29/83 (34%), Positives = 35/83 (41%), Gaps = 9/83 (10%)
Query: 94 PPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNG 153
PPP P R P P RPPPPP P P N + PP P + + PQ G
Sbjct: 1 PPPGKPQGRRPQGGNQPQ-RPPPPPGKPQGPPPQGGNQSQGPPPHPGK---PERPPPQGG 56
Query: 154 NLNQFQNPNNQFQTPNVQEQAPP 176
N +Q P+ P E+ PP
Sbjct: 57 NQSQGPPPH-----PGKPERPPP 74
Score = 42.4 bits (98), Expect = 0.003
Identities = 31/82 (37%), Positives = 32/82 (38%), Gaps = 17/82 (20%)
Query: 38 RLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQ 97
R PP + Q PP G PQ P P PP P NP Q PP
Sbjct: 168 RSPPGKPQGPPQQE-------------GNKPQGPPPPGKPQ-GPPPPGGNPQQPQAPPAG 213
Query: 98 NPNFRPPPPPQNPNFRPPPPPQ 119
P PPPPPQ RPP P Q
Sbjct: 214 KPQ-GPPPPPQGG--RPPRPAQ 232
>PR4L_HUMAN (P10162) Basic salivary proline-rich protein 4 allele L
(Salivary proline-rich protein Po) (Parotid o protein)
[Contains: Peptide P-D] (Fragment)
Length = 276
Score = 65.1 bits (157), Expect = 5e-10
Identities = 53/154 (34%), Positives = 65/154 (41%), Gaps = 19/154 (12%)
Query: 40 PPHQVQ---QPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP-----QPPQNPNYR 91
PPH + +PP +Q T P P Q N PP +PPQ N
Sbjct: 80 PPHPGKPESRPPQGGHQSQGPPPTPGKPEGPPPQGGNQSQGTPPPPGKPEGRPPQGGNQS 139
Query: 92 QQPPPQ--NPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWN 149
Q PPP P RPPP N + RPPPPP P P N + PP P +
Sbjct: 140 QGPPPHPGKPE-RPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQGPPPHPGKPEGP---P 195
Query: 150 PQNGNLNQ-FQNPNNQFQTPNVQE----QAPPPP 178
PQ GN ++ ++P + Q P QE Q PPPP
Sbjct: 196 PQEGNKSRSARSPPGKPQGPPQQEGNKPQGPPPP 229
Score = 53.5 bits (127), Expect = 1e-06
Identities = 53/171 (30%), Positives = 68/171 (38%), Gaps = 36/171 (21%)
Query: 37 GRLPP--HQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP-QPPQNPNYRQQ 93
GR P +Q Q+PP + Q P + Q P P +P +PPQ N Q
Sbjct: 4 GRRPQGGNQPQRPPPPPGK------PQGPPPQGGNQSQGPPPPPGKPEGRPPQGGNQSQG 57
Query: 94 PPPQNPNFRPPPPPQ--NPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNR-------GIN 144
PPP +P PPPQ N + PPP P P R P + + PP P + G N
Sbjct: 58 PPP-HPGKPERPPPQGGNQSQGPPPHPGKPESRPPQGGHQSQGPPPTPGKPEGPPPQGGN 116
Query: 145 QNQWN-----------PQNGNLNQFQNPN------NQFQTPNVQEQAPPPP 178
Q+Q PQ GN +Q P+ Q N + PPPP
Sbjct: 117 QSQGTPPPPGKPEGRPPQGGNQSQGPPPHPGKPERPPPQGGNQSHRPPPPP 167
Score = 52.0 bits (123), Expect = 4e-06
Identities = 51/150 (34%), Positives = 59/150 (39%), Gaps = 25/150 (16%)
Query: 40 PPHQVQ-QPPSDHNQHFNHHNTQNAPGRFPQQDWN----PQNPNFRPPQPP-QNPNYRQQ 93
PP + + +PP NQ P R P Q N P P +P +PP Q N Q
Sbjct: 124 PPGKPEGRPPQGGNQSQGPPPHPGKPERPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQG 183
Query: 94 PPPQNPNFRPPPPPQNPN-----FRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQW 148
PPP +P PPPQ N PP PQ P Q P PP G +G
Sbjct: 184 PPP-HPGKPEGPPPQEGNKSRSARSPPGKPQGP--PQQEGNKPQGPPPPGKPQGP----- 235
Query: 149 NPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P GN Q Q P P + Q PPPP
Sbjct: 236 PPPGGNPQQPQAP------PAGKPQGPPPP 259
Score = 50.8 bits (120), Expect = 9e-06
Identities = 37/106 (34%), Positives = 44/106 (40%), Gaps = 10/106 (9%)
Query: 77 PNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
P R PQ P R PPP P PPP N + PPPPP P R P N + PP
Sbjct: 2 PQGRRPQGGNQPQ-RPPPPPGKPQ-GPPPQGGNQSQGPPPPPGKPEGRPPQGGNQSQGPP 59
Query: 137 NGPNRGINQNQWNPQNGNLNQFQ-----NPNNQFQTPNVQEQAPPP 177
P + + PQ GN +Q P ++ Q Q PPP
Sbjct: 60 PHPGK---PERPPPQGGNQSQGPPPHPGKPESRPPQGGHQSQGPPP 102
Score = 48.9 bits (115), Expect = 3e-05
Identities = 36/101 (35%), Positives = 44/101 (42%), Gaps = 14/101 (13%)
Query: 40 PPHQVQQ---PPSDHNQHFNHHNTQNAPGRFPQQDWN-PQNPNFRPPQPPQNPNYRQQPP 95
PPH + PP + N+ + + P PQQ+ N PQ P PP PQ P PP
Sbjct: 185 PPHPGKPEGPPPQEGNKSRSARSPPGKPQGPPQQEGNKPQGPP--PPGKPQGP-----PP 237
Query: 96 PQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
P +P PP PPPPPQ +P R QPP
Sbjct: 238 PGGNPQQPQAPPAGKPQGPPPPPQG---GRPPRPAQGQQPP 275
Score = 42.4 bits (98), Expect = 0.003
Identities = 31/82 (37%), Positives = 32/82 (38%), Gaps = 17/82 (20%)
Query: 38 RLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQ 97
R PP + Q PP G PQ P P PP P NP Q PP
Sbjct: 206 RSPPGKPQGPPQQE-------------GNKPQGPPPPGKPQ-GPPPPGGNPQQPQAPPAG 251
Query: 98 NPNFRPPPPPQNPNFRPPPPPQ 119
P PPPPPQ RPP P Q
Sbjct: 252 KPQ-GPPPPPQGG--RPPRPAQ 270
>NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75) (NGM-75)
Length = 309
Score = 65.1 bits (157), Expect = 5e-10
Identities = 50/161 (31%), Positives = 72/161 (44%), Gaps = 34/161 (21%)
Query: 33 PNEYGRLPPHQVQ----QPPSDHNQHFNHHNTQNAPGRFPQQDWNPQN---PNFRPPQPP 85
P EY LPPH+ QPP + H N P P + P P + PP
Sbjct: 86 PPEY--LPPHEKPPPEYQPPHEKPPHENPPPEHQPPHEKPPEHQPPHEKPPPEYEPPHEK 143
Query: 86 QNPNYR---QQPPP--QNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPN 140
P Y+ ++PPP Q P+ +PPP Q P+ +PPP Q P+ + P Q P+ +PP
Sbjct: 144 PPPEYQPPHEKPPPEYQPPHEKPPPEYQPPHEKPPPEHQPPHEKPPEHQPPHEKPP---- 199
Query: 141 RGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAP---PPP 178
++ P + + P ++Q P QE+ P PPP
Sbjct: 200 -----PEYQPPH------EKPPPEYQPP--QEKPPHEKPPP 227
Score = 57.8 bits (138), Expect = 7e-08
Identities = 43/149 (28%), Positives = 67/149 (44%), Gaps = 15/149 (10%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP--QPPQNPNY 90
P EY LPP + PP H P P + P P +PP +PP++
Sbjct: 74 PPEY--LPPPHEKPPPEYLPPHEKPPPEYQPPHEKPPHENPP--PEHQPPHEKPPEHQPP 129
Query: 91 RQQPPP--QNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQW 148
++PPP + P+ +PPP Q P+ +PPP Q P+ + P P +QPP+ +Q
Sbjct: 130 HEKPPPEYEPPHEKPPPEYQPPHEKPPPEYQPPHEKPP----PEYQPPHEKPPPEHQPPH 185
Query: 149 NPQNGNLNQFQNPNNQFQTPNVQEQAPPP 177
+ + P ++Q P+ + PPP
Sbjct: 186 EKPPEHQPPHEKPPPEYQPPH---EKPPP 211
Score = 54.3 bits (129), Expect = 8e-07
Identities = 38/122 (31%), Positives = 56/122 (45%), Gaps = 24/122 (19%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP---QPPQNPN 89
P EY PPH+ +PP +H P P ++ P P+ +PP QPPQ
Sbjct: 167 PPEYQ--PPHE--KPPPEHQPPHEKPPEHQPPHEKPPPEYQP--PHEKPPPEYQPPQEKP 220
Query: 90 YRQQPPP--QNPNFRPPPPPQNPNFRPPP----------PPQNPNFRQP---IRQNPNFQ 134
++PPP Q P+ +PPP Q P+ +PPP P P + +P + P+ +
Sbjct: 221 PHEKPPPEYQPPHEKPPPEHQPPHEKPPPVYPPPYEKPPPVYEPPYEKPPPVVYPPPHEK 280
Query: 135 PP 136
PP
Sbjct: 281 PP 282
Score = 49.3 bits (116), Expect = 3e-05
Identities = 33/117 (28%), Positives = 49/117 (41%), Gaps = 22/117 (18%)
Query: 40 PPHQVQ---------------QPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPP-- 82
PP + Q +PP +H P P ++ P P+ +PP
Sbjct: 155 PPPEYQPPHEKPPPEYQPPHEKPPPEHQPPHEKPPEHQPPHEKPPPEYQP--PHEKPPPE 212
Query: 83 -QPPQNPNYRQQPPP--QNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
QPPQ ++PPP Q P+ +PPP Q P+ +PPP P + P P ++ P
Sbjct: 213 YQPPQEKPPHEKPPPEYQPPHEKPPPEHQPPHEKPPPVYPPPYEKPPPVYEPPYEKP 269
Score = 48.9 bits (115), Expect = 3e-05
Identities = 38/118 (32%), Positives = 55/118 (46%), Gaps = 20/118 (16%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQN----PNFRPP--QPPQ 86
P EY PPH+ + PP H P P ++ P + P +PP +PP+
Sbjct: 134 PPEYE--PPHE-KPPPEYQPPHEKPPPEYQPPHEKPPPEYQPPHEKPPPEHQPPHEKPPE 190
Query: 87 NPNYRQQPPP--QNPNFRPPP---PPQNPN--FRPPPPPQNPNFRQPIRQNPNFQPPN 137
+ ++PPP Q P+ +PPP PPQ +PPP Q P+ + P P QPP+
Sbjct: 191 HQPPHEKPPPEYQPPHEKPPPEYQPPQEKPPHEKPPPEYQPPHEKPP----PEHQPPH 244
Score = 48.1 bits (113), Expect = 6e-05
Identities = 37/144 (25%), Positives = 60/144 (40%), Gaps = 17/144 (11%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNY----RQQPPP 96
P +++PP+ F P +P +P P ++PP P Y ++PPP
Sbjct: 35 PPPIEKPPTYEPPPFYK------PPYYPPPVHHPP-PEYQPPHEKTPPEYLPPPHEKPPP 87
Query: 97 QN--PNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPN-GPNRGINQNQWNPQNG 153
+ P+ +PPP Q P+ +PP P + P + P QPP+ P P
Sbjct: 88 EYLPPHEKPPPEYQPPHEKPPHENPPPEHQPPHEKPPEHQPPHEKPPPEYEPPHEKPPPE 147
Query: 154 NLNQFQNPNNQFQTPNVQEQAPPP 177
+ P ++Q P+ + PPP
Sbjct: 148 YQPPHEKPPPEYQPPH---EKPPP 168
Score = 46.2 bits (108), Expect = 2e-04
Identities = 33/108 (30%), Positives = 43/108 (39%), Gaps = 12/108 (11%)
Query: 33 PNEYGRLPPHQVQ----QPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNP 88
P EY PPH+ QPP + H P P + P P+ +PP P P
Sbjct: 199 PPEYQ--PPHEKPPPEYQPPQEKPPHEKPPPEYQPPHEKPPPEHQP--PHEKPP-PVYPP 253
Query: 89 NYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
Y + PP P + PPP P PP P P+ + P + PP
Sbjct: 254 PYEKPPPVYEPPYEKPPPVVYPPPHEKPPIYEP---PPLEKPPVYNPP 298
Score = 37.0 bits (84), Expect = 0.13
Identities = 27/106 (25%), Positives = 42/106 (39%), Gaps = 9/106 (8%)
Query: 79 FRPPQPPQNPNYRQQPPPQNPNFRPPP---PPQNPNFRPPPPPQNPNFRQPIRQN--PNF 133
F P P + P + PP P + PPP PP P ++PP P + P + P +
Sbjct: 32 FYEPPPIEKPPTYEPPPFYKPPYYPPPVHHPP--PEYQPPHEKTPPEYLPPPHEKPPPEY 89
Query: 134 QPPN--GPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPP 177
PP+ P ++ P + Q P+ + + PPP
Sbjct: 90 LPPHEKPPPEYQPPHEKPPHENPPPEHQPPHEKPPEHQPPHEKPPP 135
Score = 35.4 bits (80), Expect = 0.39
Identities = 29/99 (29%), Positives = 43/99 (43%), Gaps = 22/99 (22%)
Query: 88 PNYRQQPPP--QNPNFRPPP---PPQNPN--FRPPPPPQNPNFRQPIRQNPNFQPPNGPN 140
P + +PPP + P + PPP PP P PPP Q P+ + P P + PP P+
Sbjct: 29 PRFFYEPPPIEKPPTYEPPPFYKPPYYPPPVHHPPPEYQPPHEKTP----PEYLPP--PH 82
Query: 141 RGINQNQWNPQNGNLNQFQNPNNQFQTPNVQ--EQAPPP 177
P L + P ++Q P+ + + PPP
Sbjct: 83 E-------KPPPEYLPPHEKPPPEYQPPHEKPPHENPPP 114
>PR4S_HUMAN (P10163) Basic salivary proline-rich protein 4 allele S
precursor (Salivary proline-rich protein Po) (Parotid o
protein) [Contains: Protein N1; Glycosylated protein A]
Length = 247
Score = 63.5 bits (153), Expect = 1e-09
Identities = 52/151 (34%), Positives = 62/151 (40%), Gaps = 15/151 (9%)
Query: 40 PPHQVQ-QPPSDHNQHFNHHNTQNAPGRFPQQDWN-----PQNPNFRPPQPPQNPNYRQ- 92
PP + Q PP NQ P P Q N P +P PPQ N Q
Sbjct: 53 PPGKPQGPPPQGGNQSQGPPPPPGKPEGRPPQGGNQSQGPPPHPGKPERPPPQGGNQSQG 112
Query: 93 QPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQN 152
PPP RPPP N + RPPPPP P P N + PP P + PQ
Sbjct: 113 TPPPPGKPERPPPQGGNQSHRPPPPPGKPERPPPQGGNQSQGPPPHPGKPEGP---PPQE 169
Query: 153 GNLNQ-FQNPNNQFQTPNVQE----QAPPPP 178
GN ++ ++P + Q P QE Q PPPP
Sbjct: 170 GNKSRSARSPPGKPQGPPQQEGNKPQGPPPP 200
Score = 54.7 bits (130), Expect = 6e-07
Identities = 52/152 (34%), Positives = 59/152 (38%), Gaps = 27/152 (17%)
Query: 40 PPHQ---VQQPPSDHNQHFNHHNTQNAPGRFPQQDWN----PQNPNFRPPQPP-QNPNYR 91
PPH + PP NQ P R P Q N P P +P +PP Q N
Sbjct: 93 PPHPGKPERPPPQGGNQSQGTPPPPGKPERPPPQGGNQSHRPPPPPGKPERPPPQGGNQS 152
Query: 92 QQPPPQNPNFRPPPPPQNPN-----FRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQN 146
Q PPP +P PPPQ N PP PQ P Q P PP G +G
Sbjct: 153 QGPPP-HPGKPEGPPPQEGNKSRSARSPPGKPQGP--PQQEGNKPQGPPPPGKPQGP--- 206
Query: 147 QWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P GN Q Q+P P + Q PPPP
Sbjct: 207 --PPAGGNPQQPQDP------PAGKPQGPPPP 230
Score = 51.2 bits (121), Expect = 7e-06
Identities = 38/108 (35%), Positives = 44/108 (40%), Gaps = 11/108 (10%)
Query: 77 PNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
P R PQ P R PPP P PPP N + PPPPP P R P N + PP
Sbjct: 36 PEGRRPQGGNQPQ-RPPPPPGKPQ-GPPPQGGNQSQGPPPPPGKPEGRPPQGGNQSQGPP 93
Query: 137 NGPNRGINQNQWNPQNGNLNQFQNP------NNQFQTPNVQEQAPPPP 178
P + + PQ GN +Q P Q N + PPPP
Sbjct: 94 PHPGK---PERPPPQGGNQSQGTPPPPGKPERPPPQGGNQSHRPPPPP 138
Score = 45.8 bits (107), Expect = 3e-04
Identities = 34/101 (33%), Positives = 42/101 (40%), Gaps = 14/101 (13%)
Query: 40 PPHQVQQ---PPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPP 96
PPH + PP + N+ + + P PQQ+ N PQ P P Q PPP
Sbjct: 156 PPHPGKPEGPPPQEGNKSRSARSPPGKPQGPPQQEGNK-------PQGPPPPGKPQGPPP 208
Query: 97 QNPN-FRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPP 136
N +P PP PPPPPQ +P R QPP
Sbjct: 209 AGGNPQQPQDPPAGKPQGPPPPPQG---GRPPRPAQGQQPP 246
Score = 39.7 bits (91), Expect = 0.021
Identities = 30/82 (36%), Positives = 31/82 (37%), Gaps = 17/82 (20%)
Query: 38 RLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQ 97
R PP + Q PP G PQ P P PP NP Q PP
Sbjct: 177 RSPPGKPQGPPQQE-------------GNKPQGPPPPGKPQ-GPPPAGGNPQQPQDPPAG 222
Query: 98 NPNFRPPPPPQNPNFRPPPPPQ 119
P PPPPPQ RPP P Q
Sbjct: 223 KPQ-GPPPPPQGG--RPPRPAQ 241
>PMP3_MOUSE (P05143) Proline-rich protein MP-3 (Fragment)
Length = 296
Score = 62.0 bits (149), Expect = 4e-09
Identities = 55/160 (34%), Positives = 59/160 (36%), Gaps = 18/160 (11%)
Query: 32 IPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYR 91
IPN+ R PP Q P + R PQ P P RPPQ P P
Sbjct: 8 IPNQ--RPPPSGSQPRPPVNGSQQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGP 65
Query: 92 Q------QPPPQNPNFRP---PPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPNGP 139
Q PPP P RP PPPP P RP PPPP P R P Q P PP GP
Sbjct: 66 QPRPPQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQQRPP--QGP--PPPGGP 121
Query: 140 NRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
+ Q P Q P Q PPPP+
Sbjct: 122 QQRPPQGPPPPGGPQPRPPQGPPPPAGPQPRPPQGPPPPA 161
Score = 60.8 bits (146), Expect = 9e-09
Identities = 55/155 (35%), Positives = 58/155 (36%), Gaps = 20/155 (12%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPG----RFPQQDWNPQNPNFRPPQPPQNP 88
P G P Q PP PG R PQ P P RPPQ P P
Sbjct: 59 PPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQQRPPQGPPPP 118
Query: 89 NYRQQPPPQNPNFRPPPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQ 145
QQ PPQ PPPP P RP PPPP P R P Q P PP GP+ +
Sbjct: 119 GGPQQRPPQG-----PPPPGGPQPRPPQGPPPPAGPQPRPP--QGP--PPPAGPH--LRP 167
Query: 146 NQWNPQNGNLNQF--QNPNNQFQTPNVQEQAPPPP 178
Q P G Q Q+P Q PPPP
Sbjct: 168 TQGPPPTGGPQQRYPQSPPPPGGPQPRPPQGPPPP 202
Score = 58.5 bits (140), Expect = 4e-08
Identities = 44/116 (37%), Positives = 47/116 (39%), Gaps = 16/116 (13%)
Query: 75 QNPNFRPP------QPPQNPNYRQQPPPQNPNFRP---PPPPQNPNFRP---PPPPQNPN 122
Q PN RPP +PP N + + PPP P RP PPPP P RP PPPP P
Sbjct: 7 QIPNQRPPPSGSQPRPPVNGSQQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQ 66
Query: 123 FRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
R P Q P PP GP Q P Q P Q PPPP
Sbjct: 67 PRPP--QGP--PPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQQRPPQGPPPP 118
Score = 55.8 bits (133), Expect = 3e-07
Identities = 53/153 (34%), Positives = 59/153 (37%), Gaps = 30/153 (19%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPG------RFPQQDWNPQNPNFRPPQPPQNPNYRQQP 94
P Q PP H TQ P R+PQ +P P P+PPQ P P
Sbjct: 151 PRPPQGPPPPAGPHLRP--TQGPPPTGGPQQRYPQ---SPPPPGGPQPRPPQGP-----P 200
Query: 95 PPQNPNFRPP--PPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWN 149
PP P+ RP PPP P RP PPP P R P Q P PP GP Q
Sbjct: 201 PPGGPHPRPTQGPPPTGPQPRPTQGPPPTGGPQQRPP--QGP--PPPGGPQPRPPQGPPP 256
Query: 150 PQNGNLNQFQNPN---NQFQTPNV--QEQAPPP 177
P Q P+ QTP + Q PPP
Sbjct: 257 PTGPQPRPTQGPHPTGGPQQTPPLAGNPQGPPP 289
Score = 49.7 bits (117), Expect = 2e-05
Identities = 48/130 (36%), Positives = 52/130 (39%), Gaps = 25/130 (19%)
Query: 66 RFPQQDWNPQNPNFRPPQ---PPQNPNYR--QQPPPQN------PNFRPPPPPQNPNFRP 114
R PQ P P RPPQ PP P+ R Q PPP P + PPPP P RP
Sbjct: 138 RPPQGPPPPAGPQPRPPQGPPPPAGPHLRPTQGPPPTGGPQQRYP--QSPPPPGGPQPRP 195
Query: 115 ---PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQF--QNPNNQFQTPN 169
PPPP P+ R Q P PP GP Q P G Q Q P
Sbjct: 196 PQGPPPPGGPHPRP--TQGP---PPTGPQP--RPTQGPPPTGGPQQRPPQGPPPPGGPQP 248
Query: 170 VQEQAPPPPS 179
Q PPPP+
Sbjct: 249 RPPQGPPPPT 258
Score = 46.6 bits (109), Expect = 2e-04
Identities = 38/112 (33%), Positives = 41/112 (35%), Gaps = 9/112 (8%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQ--PPQNPNY 90
P G P Q PP H TQ P PQ P PQ PPQ P
Sbjct: 185 PPPPGGPQPRPPQGPPPPGGPH--PRPTQGPPPTGPQPRPTQGPPPTGGPQQRPPQGPPP 242
Query: 91 RQQPPPQNPNFRPPPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPNGP 139
P P+ P + PPPP P RP P P P P+ NP PP P
Sbjct: 243 PGGPQPRPP--QGPPPPTGPQPRPTQGPHPTGGPQQTPPLAGNPQGPPPGRP 292
Score = 38.5 bits (88), Expect = 0.046
Identities = 32/92 (34%), Positives = 34/92 (36%), Gaps = 14/92 (15%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPG------RFPQQDWNPQNPNFRPPQPPQNPNYRQQP 94
P Q PP Q TQ P R PQ P P RPPQ P P Q
Sbjct: 207 PRPTQGPPPTGPQP---RPTQGPPPTGGPQQRPPQGPPPPGGPQPRPPQGPPPPTGPQPR 263
Query: 95 PPQNPN-----FRPPPPPQNPNFRPPPPPQNP 121
P Q P+ + PP NP PP PQ P
Sbjct: 264 PTQGPHPTGGPQQTPPLAGNPQGPPPGRPQGP 295
>EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein)
Length = 620
Score = 61.2 bits (147), Expect = 7e-09
Identities = 49/150 (32%), Positives = 60/150 (39%), Gaps = 22/150 (14%)
Query: 40 PPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQP---PQNPNYRQQPPP 96
PP PP + Q T + P P P P + PP P P P Y Q PPP
Sbjct: 412 PPPTYSPPPPTYAQPPPLPPTYSPP---PPAYSPPPPPTYSPPPPTYSPPPPAYAQPPPP 468
Query: 97 QNPNFRPP-----PPPQNPNFRPPPP---PQNPNFRQPIRQNPNFQPPNGPNRGINQNQW 148
P + PP PPP +P + PPPP P P F P + + PP P+R Q
Sbjct: 469 P-PTYSPPPPAYSPPPPSPIYSPPPPQVQPLPPTFSPPPPRRIHLPPP--PHR-----QP 520
Query: 149 NPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P Q +P P Q +PPPP
Sbjct: 521 RPPTPTYGQPPSPPTFSPPPPRQIHSPPPP 550
Score = 57.8 bits (138), Expect = 7e-08
Identities = 40/119 (33%), Positives = 48/119 (39%), Gaps = 13/119 (10%)
Query: 28 STSAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQP--- 84
S S P Y PP PPS + + P + P P + PP P
Sbjct: 360 SYSPPPPTYLPPPPPSSPPPPSFSPPPPTYEQSPPPPPAYSPP--LPAPPTYSPPPPTYS 417
Query: 85 PQNPNYRQQPPPQNPNFRPPP----PPQNPNFRPPPP---PQNPNFRQPIRQNPNFQPP 136
P P Y Q PPP P + PPP PP P + PPPP P P + QP P + PP
Sbjct: 418 PPPPTYAQ-PPPLPPTYSPPPPAYSPPPPPTYSPPPPTYSPPPPAYAQPPPPPPTYSPP 475
Score = 57.4 bits (137), Expect = 1e-07
Identities = 37/124 (29%), Positives = 46/124 (36%), Gaps = 24/124 (19%)
Query: 68 PQQDWNPQNPNFRPPQPPQN----------PNYRQQPPPQNPNFRPPPPPQNPNFRPPPP 117
P ++P P + PP PP + P Y Q PPP P PP P P + PPPP
Sbjct: 357 PPPSYSPPPPTYLPPPPPSSPPPPSFSPPPPTYEQSPPP--PPAYSPPLPAPPTYSPPPP 414
Query: 118 ---PQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQA 174
P P + QP P + PP ++P P P Q
Sbjct: 415 TYSPPPPTYAQPPPLPPTYSPPPPAYSPPPPPTYSPP---------PPTYSPPPPAYAQP 465
Query: 175 PPPP 178
PPPP
Sbjct: 466 PPPP 469
Score = 56.2 bits (134), Expect = 2e-07
Identities = 45/147 (30%), Positives = 60/147 (40%), Gaps = 21/147 (14%)
Query: 42 HQVQQPPSDHNQHFNHH---NTQNAPGRF--PQQDWNPQNPNFRPPQPPQNPNYRQQPPP 96
HQ Q P H + H Q +P R P PQ P + PP P Y Q P P
Sbjct: 223 HQPQPPTHRHAPPTHRHAPPTHQPSPLRHLPPSPRRQPQPPTYSPPPPA----YAQSPQP 278
Query: 97 QNPNFRPPPP-----PQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQ 151
+P + PPPP P +P + PPPP +P+ P P F PP P ++P
Sbjct: 279 -SPTYSPPPPTYSPPPPSPIYSPPPPAYSPS--PPPTPTPTFSPP--PPAYSPPPTYSPP 333
Query: 152 NGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P++ +P +PPPP
Sbjct: 334 PP--TYLPLPSSPIYSPPPPVYSPPPP 358
Score = 55.5 bits (132), Expect = 4e-07
Identities = 35/118 (29%), Positives = 45/118 (37%), Gaps = 6/118 (5%)
Query: 32 IPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQP---PQNP 88
+P + PP ++ PP H Q T P P P PP P P+ P
Sbjct: 498 LPPTFSPPPPRRIHLPPPPHRQPRPPTPTYGQPPSPPTFSPPPPRQIHSPPPPHWQPRTP 557
Query: 89 NYRQQPPPQNPNFRPPPPPQNPNFRPP---PPPQNPNFRQPIRQNPNFQPPNGPNRGI 143
PP P F PPP Q + PP P P P + QP + PP+ P G+
Sbjct: 558 TPTYGQPPSPPTFSAPPPRQIHSPPPPHRQPRPPTPTYGQPPSPPTTYSPPSPPPYGL 615
Score = 54.7 bits (130), Expect = 6e-07
Identities = 51/166 (30%), Positives = 64/166 (37%), Gaps = 25/166 (15%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAP-GRFPQQDWNPQNPNFR----PP---QPPQNP 88
G LP H Q+PPS + H P G+ P P P+ PP QPP P
Sbjct: 132 GHLPSHG-QRPPSPSHGHAPPSGGHTPPRGQHPPSHRRPSPPSRHGHPPPPTYAQPPPTP 190
Query: 89 NYRQQPPPQNPNFRPPPPPQNPNFRPPPP-----PQNPNFRQ--PIRQN--PNFQP---- 135
Y P Q P PPPP + P PP PQ P R P ++ P QP
Sbjct: 191 IYSPSPQVQPPPTYSPPPPTHVQPTPSPPSRGHQPQPPTHRHAPPTHRHAPPTHQPSPLR 250
Query: 136 --PNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
P P R ++P Q P+ + +P +PPPPS
Sbjct: 251 HLPPSPRRQPQPPTYSPPPPAYAQSPQPSPTY-SPPPPTYSPPPPS 295
Score = 53.9 bits (128), Expect = 1e-06
Identities = 36/119 (30%), Positives = 50/119 (41%), Gaps = 18/119 (15%)
Query: 74 PQNPNFRPP----QPPQNPNYRQQPPPQNPNFRPPPPPQNP---NFRPPP------PPQN 120
P +P + PP PP P+Y PPP P + PPPPP +P +F PPP PP
Sbjct: 341 PSSPIYSPPPPVYSPPPPPSY--SPPP--PTYLPPPPPSSPPPPSFSPPPPTYEQSPPPP 396
Query: 121 PNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
P + P+ P + PP P + + P + P +PPPP+
Sbjct: 397 PAYSPPLPAPPTYSPP-PPTYSPPPPTYAQPPPLPPTYSPPPPAYSPPPPPTYSPPPPT 454
Score = 50.1 bits (118), Expect = 2e-05
Identities = 38/150 (25%), Positives = 49/150 (32%), Gaps = 43/150 (28%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQ 92
P Y PP + PP +P P ++P P + PP P
Sbjct: 286 PPTYSPPPPSPIYSPPPPAYSP--------SPPPTPTPTFSPPPPAYSPPPTYSPPPPTY 337
Query: 93 QPPPQNPNFRPPP----PPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQW 148
P P +P + PPP PP P++ PPPP P P+F PP
Sbjct: 338 LPLPSSPIYSPPPPVYSPPPPPSYSPPPPTYLPPPPPSSPPPPSFSPP------------ 385
Query: 149 NPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
P EQ+PPPP
Sbjct: 386 -------------------PPTYEQSPPPP 396
Score = 41.6 bits (96), Expect = 0.005
Identities = 43/191 (22%), Positives = 72/191 (37%), Gaps = 32/191 (16%)
Query: 1 MLSSSSSFFSLKLRSNYIHGNHFCKTLSTSAIPNEYGRLPPHQ-VQQPPSDHNQHFNHHN 59
+L S++ + + +G + +++ P+ G PP PPS + H
Sbjct: 12 LLQSTAILSLVAAEATTQYGGYLPPPVTSQPPPSSIGLSPPSAPTTTPPSRGHVPSPRH- 70
Query: 60 TQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPP------QNPNFR 113
AP P+ + P + PP P +R PP F PPP P P++
Sbjct: 71 ---AP---PRHAYPPPSHGHLPPSVGGPPPHRGHLPPSR-GFNPPPSPVISPSHPPPSYG 123
Query: 114 PPPPPQNPNF-----RQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTP 168
PPP P ++P + PP+G + P G P+++ +P
Sbjct: 124 APPPSHGPGHLPSHGQRPPSPSHGHAPPSGGH--------TPPRGQ----HPPSHRRPSP 171
Query: 169 NVQEQAPPPPS 179
+ PPPP+
Sbjct: 172 PSRHGHPPPPT 182
>CBPA_DICDI (P35085) Calcium-binding protein
Length = 467
Score = 60.8 bits (146), Expect = 9e-09
Identities = 50/154 (32%), Positives = 60/154 (38%), Gaps = 24/154 (15%)
Query: 28 STSAIPNEYGRLPPHQVQQPPSDHNQH--FNHHNTQNAPGRFPQQDWNPQNPNFRPPQPP 85
ST P G+ PP Q P S+ + APG++P PQ P PPQ P
Sbjct: 35 STPGAPGAPGQYPPQQPGAPGSNLPPYPGTQQPGAPGAPGQYP-----PQQPGQYPPQQP 89
Query: 86 QNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQ 145
P Q PPQ P +P PPQ P PPQ P QP P +
Sbjct: 90 GAPG---QYPPQQPG-QPGYPPQQPGQSGQYPPQQPG-----------QPGYPPQQPGAP 134
Query: 146 NQWNPQNGNLNQF--QNPNNQFQTPNVQEQAPPP 177
Q+ PQ G Q+ Q P Q P Q+ PP
Sbjct: 135 GQYPPQQGQPGQYPPQQPGQPGQYPPQQQGQYPP 168
Score = 53.9 bits (128), Expect = 1e-06
Identities = 49/157 (31%), Positives = 55/157 (34%), Gaps = 28/157 (17%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQ 92
P + G+ PP Q QP Q APG++P Q P PPQ P
Sbjct: 110 PGQSGQYPPQQPGQPGYPPQQ-------PGAPGQYPPQQGQPGQ------YPPQQPGQPG 156
Query: 93 QPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIR--QN----------PNFQPPNGPN 140
Q PPQ PP P P PP P P P + QN P PP G
Sbjct: 157 QYPPQQQGQYPPQQPGQPGAYPPQQPGQPGAYPPQQGVQNTLAKTGAPGQPGVPPPQGAY 216
Query: 141 RGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPP 177
G Q PQ G Q P + P Q A PP
Sbjct: 217 PG--QPGVPPQQGAYPGQQPPMGAY-PPQGQPGAYPP 250
Score = 50.4 bits (119), Expect = 1e-05
Identities = 50/162 (30%), Positives = 59/162 (35%), Gaps = 36/162 (22%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYR- 91
P G+ PP Q Q P APG++P Q P P + P QP Q+ Y
Sbjct: 70 PGAPGQYPPQQPGQYPPQQ---------PGAPGQYPPQQ--PGQPGYPPQQPGQSGQYPP 118
Query: 92 QQP-----PPQNPNFRPPPPPQ--NPNFRPPPPPQNPNFRQPIRQ---------NPNFQP 135
QQP PPQ P PPQ P PP P P P +Q P P
Sbjct: 119 QQPGQPGYPPQQPGAPGQYPPQQGQPGQYPPQQPGQPGQYPPQQQGQYPPQQPGQPGAYP 178
Query: 136 PNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPP 177
P P + + PQ G QN + P Q PPP
Sbjct: 179 PQQPGQ---PGAYPPQQG----VQNTLAKTGAPG-QPGVPPP 212
>CSP_PLAFW (P08307) Circumsporozoite protein precursor (CS)
Length = 442
Score = 60.1 bits (144), Expect = 1e-08
Identities = 57/150 (38%), Positives = 59/150 (39%), Gaps = 11/150 (7%)
Query: 34 NEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRP-PQPPQNPNYRQ 92
NE R P H+ + P D N N N P P D N NPN P P NPN
Sbjct: 112 NEKLRKPKHKKLKQPGDGNPDPNA-NPNVDPNANPNVDPNA-NPNANPNANPNANPNANP 169
Query: 93 QPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGINQN-QW 148
P NPN P P NPN P P NPN NPN P PN N N
Sbjct: 170 NANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNVDPNANPNANPNANPNA 227
Query: 149 NPQ-NGNLNQFQNPN-NQFQTPNVQEQAPP 176
NP N N N NPN N PN A P
Sbjct: 228 NPNANPNANPNANPNANPNANPNANPNANP 257
Score = 52.0 bits (123), Expect = 4e-06
Identities = 51/156 (32%), Positives = 64/156 (40%), Gaps = 15/156 (9%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQ---NPNFRP-PQP 84
+A PN + P+ + N + N + NA P P + N NPN P P
Sbjct: 202 NANPNANPNVDPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP 261
Query: 85 PQNPNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQP---PNG 138
NPN P NPN P P NPN P P NPN NPN P PN
Sbjct: 262 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNK 319
Query: 139 PNRGINQ--NQWNPQNGNLNQFQNPNNQFQTPNVQE 172
N+G Q N N N N+++ N NN + N +E
Sbjct: 320 NNQGNGQGHNMPNDPNRNVDENANANNAVKNNNNEE 355
Score = 51.2 bits (121), Expect = 7e-06
Identities = 48/139 (34%), Positives = 54/139 (38%), Gaps = 14/139 (10%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 194 NANPNANPNANPNANPNVDPNANPNANPNANPNANPNANPNANPNA-NPNANPNANPNAN 252
Query: 88 PNYRQQPPPQNPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGINQ 145
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 253 PNAN---PNANPNANPNANPNANPNANPNANPNANPN------ANPNANPNANPNANPNA 303
Query: 146 NQWNPQNGNLNQFQNPNNQ 164
N N N N N NNQ
Sbjct: 304 NPNANPNANPNANPNKNNQ 322
Score = 50.4 bits (119), Expect = 1e-05
Identities = 52/157 (33%), Positives = 59/157 (37%), Gaps = 10/157 (6%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 182 NANPNANPNANPNANPNANPNANPNANPNVDPNANPNANPNANPNA-NPNANPNANPNAN 240
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 241 PNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPN 298
Query: 145 QN-QWNPQ-NGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
N NP N N N NPN Q P P+
Sbjct: 299 ANPNANPNANPNANPNANPNKNNQGNGQGHNMPNDPN 335
Score = 44.3 bits (103), Expect = 8e-04
Identities = 40/133 (30%), Positives = 52/133 (39%), Gaps = 13/133 (9%)
Query: 52 NQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPN 111
N + N+ N N +D + ++ N + + P +++ P + N P P NPN
Sbjct: 82 NGNNNNGNNNNGDNGREGKDEDKRDGNNEDNEKLRKPKHKKLKQPGDGN---PDPNANPN 138
Query: 112 FRPPPPPQ-----NPNFRQPIRQNPNFQPPNGPNRGINQN-QWNPQ-NGNLNQFQNPN-N 163
P P NPN NPN P PN N N NP N N N NPN N
Sbjct: 139 VDPNANPNVDPNANPNANP--NANPNANPNANPNANPNANPNANPNANPNANPNANPNAN 196
Query: 164 QFQTPNVQEQAPP 176
PN A P
Sbjct: 197 PNANPNANPNANP 209
>PRP2_MOUSE (P05142) Proline-rich protein MP-2 precursor
Length = 261
Score = 59.3 bits (142), Expect = 3e-08
Identities = 57/163 (34%), Positives = 57/163 (34%), Gaps = 23/163 (14%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPG----RFPQQDWNPQNPNFRPPQPPQNP 88
P G P Q PP PG R PQ P P RPPQ P P
Sbjct: 52 PPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPP 111
Query: 89 NYRQQ------PPPQNPNFRP---PPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPP 136
QQ PPP P RP PPPP P RP PPPP P R P Q P PP
Sbjct: 112 GGPQQRPPQGPPPPGGPQPRPPQGPPPPGGPQLRPPQGPPPPAGPQPRPP--QGP--PPP 167
Query: 137 NGPNRGINQNQWNPQNG-NLNQFQNPNNQFQTPNVQEQAPPPP 178
GP Q P G Q P Q PPPP
Sbjct: 168 AGPQP--RPPQGPPTTGPQPRPTQGPPPTGGPQQQPPQGPPPP 208
Score = 58.2 bits (139), Expect = 6e-08
Identities = 44/116 (37%), Positives = 47/116 (39%), Gaps = 16/116 (13%)
Query: 75 QNPNFRPP------QPPQNPNYRQQPPPQNPNFRP---PPPPQNPNFRP---PPPPQNPN 122
Q PN RPP +PP N + + PPP P RP PPPP P RP PPPP P
Sbjct: 28 QIPNQRPPPSGFQPRPPVNGSQQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGPQ 87
Query: 123 FRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
R P Q P PP GP Q P Q P Q PPPP
Sbjct: 88 PRPP--QGP--PPPGGPQPRPPQGPPPPGGPQQRPPQGPPPPGGPQPRPPQGPPPP 139
Score = 58.2 bits (139), Expect = 6e-08
Identities = 52/150 (34%), Positives = 56/150 (36%), Gaps = 12/150 (8%)
Query: 32 IPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYR 91
IPN+ R PP Q P + R PQ P P RPPQ P P
Sbjct: 29 IPNQ--RPPPSGFQPRPPVNGSQQGPPPPGGPQPRPPQGPPPPGGPQPRPPQGPPPPGGP 86
Query: 92 QQPPPQNPNFRPPP--PPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWN 149
Q PPQ P PPP P P + PPPP P R P Q P PP GP Q
Sbjct: 87 QPRPPQGP---PPPGGPQPRPP-QGPPPPGGPQQRPP--QGP--PPPGGPQPRPPQGPPP 138
Query: 150 PQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
P L Q P Q PPPP+
Sbjct: 139 PGGPQLRPPQGPPPPAGPQPRPPQGPPPPA 168
Score = 55.1 bits (131), Expect = 5e-07
Identities = 48/144 (33%), Positives = 54/144 (37%), Gaps = 15/144 (10%)
Query: 45 QQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQ---QPPPQNPNF 101
Q P + N + P F Q P N + + P PP P R PPP P
Sbjct: 17 QGPREELQNQIQIPNQRPPPSGF--QPRPPVNGSQQGPPPPGGPQPRPPQGPPPPGGPQP 74
Query: 102 RP---PPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNL 155
RP PPPP P RP PPPP P R P Q P PP GP + Q P
Sbjct: 75 RPPQGPPPPGGPQPRPPQGPPPPGGPQPRPP--QGP--PPPGGPQQRPPQGPPPPGGPQP 130
Query: 156 NQFQNPNNQFQTPNVQEQAPPPPS 179
Q P Q PPPP+
Sbjct: 131 RPPQGPPPPGGPQLRPPQGPPPPA 154
Score = 51.6 bits (122), Expect = 5e-06
Identities = 53/164 (32%), Positives = 56/164 (33%), Gaps = 22/164 (13%)
Query: 33 PNEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPG----RFPQQDWNPQNPNFRPPQ---PP 85
P G P Q PP PG R PQ P P RPPQ PP
Sbjct: 94 PPPPGGPQPRPPQGPPPPGGPQQRPPQGPPPPGGPQPRPPQGPPPPGGPQLRPPQGPPPP 153
Query: 86 QNPNYRQQ---PPPQNPNFRPP--PPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQPPN 137
P R PPP P RPP PP P RP PPP P + P Q P PP
Sbjct: 154 AGPQPRPPQGPPPPAGPQPRPPQGPPTTGPQPRPTQGPPPTGGPQQQPP--QGP--PPPG 209
Query: 138 GPNRGINQNQWNPQNGNLNQFQNP---NNQFQTPNVQEQAPPPP 178
GP Q P + Q P QTP + PP
Sbjct: 210 GPQPRPPQGPPPPGGPQPSPTQGPPPTGGPQQTPPLAGNTQGPP 253
Score = 38.9 bits (89), Expect = 0.035
Identities = 33/105 (31%), Positives = 38/105 (35%), Gaps = 22/105 (20%)
Query: 41 PHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPN 100
P Q PP+ Q P + P PQ QPPQ P P P+ P
Sbjct: 172 PRPPQGPPTT--------GPQPRPTQGPPPTGGPQQ------QPPQGPPPPGGPQPRPP- 216
Query: 101 FRPPPPPQNPNFRP---PPPPQNPNFRQPIRQNPNFQP---PNGP 139
+ PPPP P P PPP P P+ N P P GP
Sbjct: 217 -QGPPPPGGPQPSPTQGPPPTGGPQQTPPLAGNTQGPPQGRPQGP 260
Score = 38.9 bits (89), Expect = 0.035
Identities = 34/100 (34%), Positives = 37/100 (37%), Gaps = 26/100 (26%)
Query: 66 RFPQQDWNPQNPNFRPPQ--------------PPQNPNYRQQPP--PQNPNFRPPPPPQN 109
R PQ P P RPPQ PP +QQPP P P P PPQ
Sbjct: 159 RPPQGPPPPAGPQPRPPQGPPTTGPQPRPTQGPPPTGGPQQQPPQGPPPPGGPQPRPPQG 218
Query: 110 PNFRPPP--PPQNPNFRQPIRQNPNFQPP-----NGPNRG 142
P PPP P +P P P PP GP +G
Sbjct: 219 P---PPPGGPQPSPTQGPPPTGGPQQTPPLAGNTQGPPQG 255
>CSP_PLAFT (P13814) Circumsporozoite protein precursor (CS)
Length = 424
Score = 58.2 bits (139), Expect = 6e-08
Identities = 57/152 (37%), Positives = 59/152 (38%), Gaps = 11/152 (7%)
Query: 34 NEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRP-PQPPQNPNYRQ 92
NE R P H+ + P D N N N P P D N NPN P P NPN
Sbjct: 102 NETLRKPKHKKLKQPGDGNPDPNA-NPNVDPNANPNVDPNA-NPNVDPNANPNANPNANP 159
Query: 93 QPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQN--PNFQPPNGPNRGINQN- 146
P NPN P P NPN P P NPN N PN P PN N N
Sbjct: 160 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP 219
Query: 147 QWNPQ-NGNLNQFQNPN-NQFQTPNVQEQAPP 176
NP N N N NPN N PN A P
Sbjct: 220 NANPNANPNANPNANPNANPNANPNANPNANP 251
Score = 52.0 bits (123), Expect = 4e-06
Identities = 48/140 (34%), Positives = 54/140 (38%), Gaps = 8/140 (5%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 168 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNANPNAN 226
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 227 PNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPN 284
Query: 145 QNQWNPQNGNLNQFQNPNNQ 164
N N N N N NNQ
Sbjct: 285 ANPNANPNANPNANPNKNNQ 304
Score = 51.6 bits (122), Expect = 5e-06
Identities = 51/153 (33%), Positives = 63/153 (40%), Gaps = 13/153 (8%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 188 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNANPNAN 246
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQP---PNGPNR 141
PN P NPN P P NPN P P NPN NPN P PN N+
Sbjct: 247 PNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNKNNQ 304
Query: 142 GINQ--NQWNPQNGNLNQFQNPNNQFQTPNVQE 172
G Q N N N N+++ N NN + N +E
Sbjct: 305 GNGQGHNMPNDPNRNVDENANANNAVKNNNNEE 337
Score = 51.2 bits (121), Expect = 7e-06
Identities = 52/157 (33%), Positives = 59/157 (37%), Gaps = 10/157 (6%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 164 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNANPNAN 222
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 223 PNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPN 280
Query: 145 QN-QWNPQ-NGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
N NP N N N NPN Q P P+
Sbjct: 281 ANPNANPNANPNANPNANPNKNNQGNGQGHNMPNDPN 317
Score = 42.4 bits (98), Expect = 0.003
Identities = 37/124 (29%), Positives = 47/124 (37%), Gaps = 9/124 (7%)
Query: 59 NTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPP 118
N N +D + ++ N + + P +++ P + N P P NPN P P
Sbjct: 79 NNNNGDNNREGKDEDKRDGNNEDNETLRKPKHKKLKQPGDGN---PDPNANPNVDPNANP 135
Query: 119 Q---NPNFRQPIRQNPNFQPPNGPNRGINQN-QWNPQ-NGNLNQFQNPN-NQFQTPNVQE 172
N N NPN P PN N N NP N N N NPN N PN
Sbjct: 136 NVDPNANPNVDPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP 195
Query: 173 QAPP 176
A P
Sbjct: 196 NANP 199
>CSP_PLAFO (P19597) Circumsporozoite protein precursor (CS)
Length = 397
Score = 58.2 bits (139), Expect = 6e-08
Identities = 57/154 (37%), Positives = 60/154 (38%), Gaps = 15/154 (9%)
Query: 34 NEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRP-PQPPQNPNYRQ 92
NE R P H+ + P+D N N N P P D N NPN P P NPN
Sbjct: 83 NEKLRKPKHKKLKQPADGNPDPNA-NPNVDPNANPNVDPNA-NPNVDPNANPNANPNANP 140
Query: 93 QPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGINQN--- 146
P NPN P P NPN P P NPN NPN P PN N N
Sbjct: 141 NANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPNANPNV 198
Query: 147 --QWNPQ-NGNLNQFQNPN-NQFQTPNVQEQAPP 176
NP N N N NPN N PN A P
Sbjct: 199 DPNANPNANPNANPNANPNANPNANPNANPNANP 232
Score = 51.2 bits (121), Expect = 7e-06
Identities = 48/140 (34%), Positives = 54/140 (38%), Gaps = 12/140 (8%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRPPQPPQ-N 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 145 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNVDPNAN 203
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 204 PNANPNANPNANPNANPNANPNANPNANPNANPNANPN------ANPNANPNANPNANPN 257
Query: 145 QNQWNPQNGNLNQFQNPNNQ 164
N N N N N NNQ
Sbjct: 258 ANPNANPNANPNANPNKNNQ 277
Score = 50.8 bits (120), Expect = 9e-06
Identities = 52/157 (33%), Positives = 59/157 (37%), Gaps = 10/157 (6%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRP-PQPPQN 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 137 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNANPNAN 195
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 196 PNVDPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPN 253
Query: 145 QN-QWNPQ-NGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
N NP N N N NPN Q P P+
Sbjct: 254 ANPNANPNANPNANPNANPNKNNQGNGQGHNMPNDPN 290
Score = 50.8 bits (120), Expect = 9e-06
Identities = 50/157 (31%), Positives = 59/157 (36%), Gaps = 13/157 (8%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQ---NPNFRP-PQP 84
+A PN P+ + N + N + NA P P D N NPN P P
Sbjct: 157 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNVDPNANPNANPNANPNANP 216
Query: 85 PQNPNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQP---PNG 138
NPN P NPN P P NPN P P NPN NPN P PN
Sbjct: 217 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNK 274
Query: 139 PNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAP 175
N+G Q P + N N +N N N + P
Sbjct: 275 NNQGNGQGHNMPNDPNRNVDENANANSAVKNNNNEEP 311
>PRP2_RAT (P10164) Acidic proline-rich protein PRP25 precursor
(Fragment)
Length = 172
Score = 57.8 bits (138), Expect = 7e-08
Identities = 48/153 (31%), Positives = 61/153 (39%), Gaps = 33/153 (21%)
Query: 40 PPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNP------NFRPPQP-----PQNP 88
PP +PP++ +Q Q P + P Q PQ+P +PPQP P P
Sbjct: 36 PPGFPPRPPANGSQQ--GPPPQGGPQQSPLQPGKPQDPPPQGSPQQKPPQPGKPQGPPPP 93
Query: 89 NYRQQPPPQNPNFRPPPPPQNPNFRPPPP--PQNPN-FRQPIRQNPNFQPPNGPNRGINQ 145
Q+ PPQ + PPPP P +PP P PQ P P ++ P P GP
Sbjct: 94 GGPQKKPPQPGKPQGPPPPGGPQKKPPQPGKPQGPTPPGGPQQKPPQAGKPQGPPPPGGP 153
Query: 146 NQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
Q PQ GN +Q PPPP
Sbjct: 154 QQKPPQPGN-----------------QQGPPPP 169
Score = 48.9 bits (115), Expect = 3e-05
Identities = 38/123 (30%), Positives = 46/123 (36%), Gaps = 24/123 (19%)
Query: 74 PQNPNFRPPQPPQNPNYRQQPPPQNPNFRP--------PPPPQNPNFRP--------PPP 117
P P F PP+PP N + + PP P P PPP +P +P PPP
Sbjct: 34 PPPPGF-PPRPPANGSQQGPPPQGGPQQSPLQPGKPQDPPPQGSPQQKPPQPGKPQGPPP 92
Query: 118 PQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNV--QEQAP 175
P P + P P PP G + PQ G P Q P + Q P
Sbjct: 93 PGGPQKKPPQPGKPQGPPPPG-----GPQKKPPQPGKPQGPTPPGGPQQKPPQAGKPQGP 147
Query: 176 PPP 178
PPP
Sbjct: 148 PPP 150
Score = 46.6 bits (109), Expect = 2e-04
Identities = 31/81 (38%), Positives = 36/81 (44%), Gaps = 16/81 (19%)
Query: 64 PGRFPQQDWNPQNPNFRPPQP-----PQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPP 118
PG+ PQ P P +PPQP P P QQ PPQ + PPPP P +PP P
Sbjct: 103 PGK-PQGPPPPGGPQKKPPQPGKPQGPTPPGGPQQKPPQAGKPQGPPPPGGPQQKPPQPG 161
Query: 119 QNPNFRQPIRQNPNFQPPNGP 139
+Q P PP GP
Sbjct: 162 N--------QQGP--PPPGGP 172
Score = 40.0 bits (92), Expect = 0.016
Identities = 34/103 (33%), Positives = 41/103 (39%), Gaps = 17/103 (16%)
Query: 86 QNPNYRQQPPPQNPNFRPPPPPQNPNFRPP-------PPPQNPNFRQPIRQ-NPNFQPPN 137
Q P Q Q PN +PPPP P RPP PPPQ + P++ P PP
Sbjct: 17 QEPGDELQILDQTPNQKPPPPGFPP--RPPANGSQQGPPPQGGPQQSPLQPGKPQDPPPQ 74
Query: 138 GPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNV--QEQAPPPP 178
G + Q PQ G P + P + Q PPPP
Sbjct: 75 G-----SPQQKPPQPGKPQGPPPPGGPQKKPPQPGKPQGPPPP 112
Score = 38.5 bits (88), Expect = 0.046
Identities = 27/83 (32%), Positives = 33/83 (39%), Gaps = 13/83 (15%)
Query: 64 PGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRP--------P 115
PG ++ P P P PP P Q+ PPQ + P PP P +P P
Sbjct: 93 PGGPQKKPPQPGKPQ--GPPPPGGP---QKKPPQPGKPQGPTPPGGPQQKPPQAGKPQGP 147
Query: 116 PPPQNPNFRQPIRQNPNFQPPNG 138
PPP P + P N PP G
Sbjct: 148 PPPGGPQQKPPQPGNQQGPPPPG 170
>RB97_DROME (Q02926) Ribonucleoprotein RB97D
Length = 471
Score = 57.0 bits (136), Expect = 1e-07
Identities = 47/132 (35%), Positives = 55/132 (41%), Gaps = 19/132 (14%)
Query: 56 NHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPN--YRQQPPPQN-----PNFR---PPP 105
N Q G+ P N Q P+FRPP PPQ Y+QQPPP PNF PPP
Sbjct: 222 NQQQQQQGGGQQPPNG-NMQAPSFRPPVPPQQAMGPYQQQPPPAPMSAPPPNFNYWGPPP 280
Query: 106 PPQNPNFR-PPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQ 164
P P ++ PPP N P Q + P P G QW + NQ+ P
Sbjct: 281 PAMPPYYQQPPPQQMNGWGPYPPPQQNGWNAPPPPPPG--AQQW-----HANQWGCPPPV 333
Query: 165 FQTPNVQEQAPP 176
Q P V PP
Sbjct: 334 QQVPPVGAVPPP 345
Score = 54.3 bits (129), Expect = 8e-07
Identities = 43/119 (36%), Positives = 53/119 (44%), Gaps = 24/119 (20%)
Query: 45 QQPPSDHNQHFNHHNT---QNAPGRFPQQ------DWNPQNPNFRPPQPPQNPNYRQQPP 95
QQPP+ + Q + Q A G + QQ P N N+ P PP P Y QQPP
Sbjct: 232 QQPPNGNMQAPSFRPPVPPQQAMGPYQQQPPPAPMSAPPPNFNYWGPPPPAMPPYYQQPP 291
Query: 96 PQNPN-FRPPPPPQNPNFRPPPPPQNPNFRQ----------PIRQNP---NFQPPNGPN 140
PQ N + P PPPQ + PPPP P +Q P++Q P PP G N
Sbjct: 292 PQQMNGWGPYPPPQQNGWNAPPPPP-PGAQQWHANQWGCPPPVQQVPPVGAVPPPMGHN 349
Score = 51.6 bits (122), Expect = 5e-06
Identities = 47/143 (32%), Positives = 58/143 (39%), Gaps = 14/143 (9%)
Query: 39 LPPHQVQQPPSDHNQHFNH----HNTQNAPGRFPQ--QDWNPQNPNFRPPQPPQNPNYRQ 92
+PP+ Q PP N + N NAP P Q W+ N PP Q P
Sbjct: 283 MPPYYQQPPPQQMNGWGPYPPPQQNGWNAPPPPPPGAQQWHA-NQWGCPPPVQQVPPVGA 341
Query: 93 QPPPQNPNFRPPPPPQNPNFRPP-PPPQNPNFRQPIRQNPNFQPPNG----PNRGINQNQ 147
PPP N PP P N N PP P P+ +Q Q P QP G N G ++
Sbjct: 342 VPPPMGHNGPPPTAPGNWNMPPPVPGAAPPSHQQQSSQQPTPQPNFGTGYQQNYGGGPSK 401
Query: 148 WNPQNGN-LNQFQ-NPNNQFQTP 168
N N N +N + P N + TP
Sbjct: 402 HNNMNANRMNPYSAGPPNSYHTP 424
>PRP1_HUMAN (P04280) Basic salivary proline-rich protein 1 precursor
(Salivary proline-rich protein) [Contains: Basic peptide
IB-6; Peptide P-H]
Length = 392
Score = 57.0 bits (136), Expect = 1e-07
Identities = 48/144 (33%), Positives = 60/144 (41%), Gaps = 19/144 (13%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWN-PQNPNFRPPQPPQNPNYRQQPP 95
G PP + Q PP ++ + + P P Q N PQ P P PP P Q PP
Sbjct: 131 GPPPPGKPQGPPPQGDKSQSPRSPPGKPQGPPPQGGNQPQGP----PPPPGKP---QGPP 183
Query: 96 PQNPNF-RPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGN 154
PQ N + PPPP P PPPQ + P ++P +P P +G NQ Q P
Sbjct: 184 PQGGNKPQGPPPPGKPQ---GPPPQGDKSQSP--RSPPGKPQGPPPQGGNQPQGPPPPPG 238
Query: 155 LNQFQNPNNQFQTPNVQEQAPPPP 178
P Q + Q PPPP
Sbjct: 239 -----KPQGPPQQGGNRPQGPPPP 257
Score = 56.6 bits (135), Expect = 2e-07
Identities = 47/146 (32%), Positives = 54/146 (36%), Gaps = 7/146 (4%)
Query: 40 PPHQVQQPPSDHNQHFNHHNTQNAP-GRFPQQDWN--PQNPNFRPPQPPQNPNYRQQPPP 96
PP + Q PP P G PQ D + PQ+P +P PP + Q PP
Sbjct: 236 PPGKPQGPPQQGGNRPQGPPPPGKPQGPPPQGDKSRSPQSPPGKPQGPPPQGGNQPQGPP 295
Query: 97 QNPNFRPPPPPQNPNFR--PPPP--PQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQN 152
P PPPQ N PPPP PQ P + + PP P Q NPQ
Sbjct: 296 PPPGKPQGPPPQGGNKPQGPPPPGKPQGPPAQGGSKSQSARAPPGKPQGPPQQEGNNPQG 355
Query: 153 GNLNQFQNPNNQFQTPNVQEQAPPPP 178
NP P Q Q PP P
Sbjct: 356 PPPPAGGNPQQPQAPPAGQPQGPPRP 381
Score = 56.2 bits (134), Expect = 2e-07
Identities = 48/144 (33%), Positives = 60/144 (41%), Gaps = 19/144 (13%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWN-PQNPNFRPPQPPQNPNYRQQPP 95
G PP + Q PP ++ + + P P Q N PQ P P PP P Q PP
Sbjct: 70 GPPPPGKPQGPPPQGDKSRSPRSPPGKPQGPPPQGGNQPQGP----PPPPGKP---QGPP 122
Query: 96 PQNPNF-RPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGN 154
PQ N + PPPP P PPPQ + P ++P +P P +G NQ Q P
Sbjct: 123 PQGGNKPQGPPPPGKPQ---GPPPQGDKSQSP--RSPPGKPQGPPPQGGNQPQGPPPPPG 177
Query: 155 LNQFQNPNNQFQTPNVQEQAPPPP 178
Q P + Q PPPP
Sbjct: 178 KPQGPPPQG-----GNKPQGPPPP 196
Score = 54.7 bits (130), Expect = 6e-07
Identities = 55/174 (31%), Positives = 67/174 (37%), Gaps = 36/174 (20%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWN----PQNPNFRPPQPPQNPNYRQ 92
G PP + Q PP ++ + + P P Q N P P +P PPQ R
Sbjct: 192 GPPPPGKPQGPPPQGDKSQSPRSPPGKPQGPPPQGGNQPQGPPPPPGKPQGPPQQGGNRP 251
Query: 93 Q--PPPQNPNFRPP-----------------PPPQNPN--FRPPPPPQNPNFRQPIRQN- 130
Q PPP P PP PPPQ N PPPPP P P N
Sbjct: 252 QGPPPPGKPQGPPPQGDKSRSPQSPPGKPQGPPPQGGNQPQGPPPPPGKPQGPPPQGGNK 311
Query: 131 PNFQPPNGPNRGINQNQWNPQNGNLNQ-FQNPNNQFQTPNVQE----QAPPPPS 179
P PP G +G Q G+ +Q + P + Q P QE Q PPPP+
Sbjct: 312 PQGPPPPGKPQGP-----PAQGGSKSQSARAPPGKPQGPPQQEGNNPQGPPPPA 360
Score = 54.3 bits (129), Expect = 8e-07
Identities = 40/111 (36%), Positives = 48/111 (43%), Gaps = 15/111 (13%)
Query: 73 NPQNPN----FRPPQPPQNPNYRQQPPPQNPNF-RPPPPPQNPNFRPPPPPQNPNFRQPI 127
NPQ P+ +P PP P Q PPPQ N + PPPP P PPPQ R P
Sbjct: 35 NPQGPSPQGGNKPQGPPPPPGKPQGPPPQGGNKPQGPPPPGKPQ---GPPPQGDKSRSP- 90
Query: 128 RQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPP 178
++P +P P +G NQ Q P Q P + Q PPPP
Sbjct: 91 -RSPPGKPQGPPPQGGNQPQGPPPPPGKPQGPPPQG-----GNKPQGPPPP 135
Score = 52.0 bits (123), Expect = 4e-06
Identities = 46/136 (33%), Positives = 50/136 (35%), Gaps = 24/136 (17%)
Query: 62 NAPGRFPQQDWNPQNP---NFRPPQPPQNPNYRQQPPPQNPNFRPP---------PPPQN 109
N P P PQ P PQ P P Q PPPQ R P PPPQ
Sbjct: 45 NKPQGPPPPPGKPQGPPPQGGNKPQGPPPPGKPQGPPPQGDKSRSPRSPPGKPQGPPPQG 104
Query: 110 PN--FRPPPPPQNPNFRQPIRQN-PNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQ 166
N PPPPP P P N P PP G +G PQ ++P + Q
Sbjct: 105 GNQPQGPPPPPGKPQGPPPQGGNKPQGPPPPGKPQGP-----PPQGDKSQSPRSPPGKPQ 159
Query: 167 TP----NVQEQAPPPP 178
P Q Q PPPP
Sbjct: 160 GPPPQGGNQPQGPPPP 175
Score = 43.9 bits (102), Expect = 0.001
Identities = 44/132 (33%), Positives = 51/132 (38%), Gaps = 31/132 (23%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWN-PQNPNFRP--PQ--PPQNPNYR 91
G PP + Q PP ++ + + P P Q N PQ P P PQ PPQ N
Sbjct: 253 GPPPPGKPQGPPPQGDKSRSPQSPPGKPQGPPPQGGNQPQGPPPPPGKPQGPPPQGGNKP 312
Query: 92 QQPPPQN------------------PNFRPPPPPQ----NPNFRPPPPPQNPNFRQPIRQ 129
Q PPP P +P PPQ NP PPPP N +QP Q
Sbjct: 313 QGPPPPGKPQGPPAQGGSKSQSARAPPGKPQGPPQQEGNNPQ--GPPPPAGGNPQQP--Q 368
Query: 130 NPNFQPPNGPNR 141
P P GP R
Sbjct: 369 APPAGQPQGPPR 380
Score = 42.4 bits (98), Expect = 0.003
Identities = 32/86 (37%), Positives = 38/86 (43%), Gaps = 10/86 (11%)
Query: 37 GRLPPHQVQQPPSDHNQHFNHHNTQNAPGRF---PQQDWNPQNPNFRPPQPPQNPNYRQQ 93
G PP + Q PP+ + + PG+ PQQ+ N NP PP NP Q
Sbjct: 314 GPPPPGKPQGPPAQGGS--KSQSARAPPGKPQGPPQQEGN--NPQGPPPPAGGNPQQPQA 369
Query: 94 PPPQNPNFRPPPPPQNPNFRPPPPPQ 119
PP P PP PPQ RP PPQ
Sbjct: 370 PPAGQPQ-GPPRPPQGG--RPSRPPQ 392
Score = 36.2 bits (82), Expect = 0.23
Identities = 31/99 (31%), Positives = 36/99 (36%), Gaps = 11/99 (11%)
Query: 40 PPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNP 99
PP + Q PP P P Q + PP PQ P P +
Sbjct: 297 PPGKPQGPPPQGGNKPQGPPPPGKPQGPPAQGGSKSQSARAPPGKPQGP-----PQQEGN 351
Query: 100 NFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNG 138
N + PPPP N P PQ P QP Q P +PP G
Sbjct: 352 NPQGPPPPAGGN---PQQPQAPPAGQP--QGPP-RPPQG 384
Score = 35.0 bits (79), Expect = 0.51
Identities = 19/68 (27%), Positives = 27/68 (38%)
Query: 61 QNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQN 120
Q P + + + + P +P PPQ Q PP P P P +P PP+
Sbjct: 322 QGPPAQGGSKSQSARAPPGKPQGPPQQEGNNPQGPPPPAGGNPQQPQAPPAGQPQGPPRP 381
Query: 121 PNFRQPIR 128
P +P R
Sbjct: 382 PQGGRPSR 389
>CSP_PLAFA (P02893) Circumsporozoite protein precursor (CS)
Length = 412
Score = 57.0 bits (136), Expect = 1e-07
Identities = 57/153 (37%), Positives = 59/153 (38%), Gaps = 21/153 (13%)
Query: 34 NEYGRLPPHQVQQPPSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRP-PQPPQNPNYRQ 92
NE R P H+ + P D N N N P P D N NPN P P NPN
Sbjct: 102 NEKLRKPKHKKLKQPGDGNPDPNA-NPNVDPNANPNVDPNA-NPNVDPNANPNANPNAN- 158
Query: 93 QPPPQNPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGINQN---- 146
P NPN P P NPN P P NPN NPN P PN N N
Sbjct: 159 --PNANPNANPNANPNANPNANPNANPNANPN------ANPNANPNANPNANPNANPNVD 210
Query: 147 -QWNPQ-NGNLNQFQNPN-NQFQTPNVQEQAPP 176
NP N N N NPN N PN A P
Sbjct: 211 PNANPNANPNANPNANPNANPNANPNANPNANP 243
Score = 51.6 bits (122), Expect = 5e-06
Identities = 48/140 (34%), Positives = 54/140 (38%), Gaps = 8/140 (5%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQNPNFRPPQPPQ-N 87
+A PN P+ + N + N + NA P P + N NPN P P N
Sbjct: 156 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNA-NPNANPNVDPNAN 214
Query: 88 PNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQNPNFQPPNGPNRGIN 144
PN P NPN P P NPN P P NPN NPN P PN N
Sbjct: 215 PNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP--NANPNANPNANPNANPN 272
Query: 145 QNQWNPQNGNLNQFQNPNNQ 164
N N N N N NNQ
Sbjct: 273 ANPNANPNANPNANPNKNNQ 292
Score = 51.2 bits (121), Expect = 7e-06
Identities = 52/158 (32%), Positives = 63/158 (38%), Gaps = 15/158 (9%)
Query: 30 SAIPNEYGRLPPHQVQQPPSDHNQHFNHHNTQNA-PGRFPQQDWNPQ---NPNFRP-PQP 84
+A PN P+ + N + N + NA P P D N NPN P P
Sbjct: 168 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNVDPNANPNANPNANPNANP 227
Query: 85 PQNPNYRQQPPPQ-NPNFRPPPPPQ-NPNFRPPPPPQ-NPNFRQPIRQN--PNFQP---P 136
NPN P NPN P P NPN P P NPN N PN P P
Sbjct: 228 NANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP 287
Query: 137 NGPNRGINQ--NQWNPQNGNLNQFQNPNNQFQTPNVQE 172
N N+G Q N N N N+++ N NN + N +E
Sbjct: 288 NKNNQGNGQGHNMPNDPNRNVDENANANNAVKNNNNEE 325
Score = 41.6 bits (96), Expect = 0.005
Identities = 37/124 (29%), Positives = 47/124 (37%), Gaps = 9/124 (7%)
Query: 59 NTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPP 118
N N +D + ++ N + + P +++ P + N P P NPN P P
Sbjct: 79 NNNNGDNGREGKDEDKRDGNNEDNEKLRKPKHKKLKQPGDGN---PDPNANPNVDPNANP 135
Query: 119 Q---NPNFRQPIRQNPNFQPPNGPNRGINQN-QWNPQ-NGNLNQFQNPN-NQFQTPNVQE 172
N N NPN P PN N N NP N N N NPN N PN
Sbjct: 136 NVDPNANPNVDPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANPNANP 195
Query: 173 QAPP 176
A P
Sbjct: 196 NANP 199
>SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor
(SSG 185)
Length = 485
Score = 56.2 bits (134), Expect = 2e-07
Identities = 33/95 (34%), Positives = 41/95 (42%), Gaps = 3/95 (3%)
Query: 48 PSDHNQHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQP---PQNPNYRQQPPPQNPNFRPP 104
P+ +N A R P +P+ P+ PP P P P PPP P+ PP
Sbjct: 222 PAPNNSPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPP 281
Query: 105 PPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGP 139
PPP P PPPPP R+P +P PP P
Sbjct: 282 PPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSP 316
Score = 52.8 bits (125), Expect = 2e-06
Identities = 36/117 (30%), Positives = 45/117 (37%), Gaps = 24/117 (20%)
Query: 63 APGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPN 122
AP P + RPP PP +P PPP +P+ PPPPP P PPPPP +P
Sbjct: 223 APNNSPLPPSPQPTASSRPPSPPPSPR-PPSPPPPSPS--PPPPPPPPPPPPPPPPPSPP 279
Query: 123 FRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPS 179
P P PP +P+ + P+ PPPPS
Sbjct: 280 PPPPPPPPPPPPPPP---------------------PSPSPPRKPPSPSPPVPPPPS 315
Score = 40.8 bits (94), Expect = 0.009
Identities = 20/49 (40%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Query: 74 PQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPN 122
P P+ PP PP P PPP P+ PP P +P+ PPPP P+
Sbjct: 273 PPPPSPPPPPPPPPP---PPPPPPPPSPSPPRKPPSPSPPVPPPPSPPS 318
Score = 36.2 bits (82), Expect = 0.23
Identities = 19/58 (32%), Positives = 24/58 (40%)
Query: 82 PQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRPPPPPQNPNFRQPIRQNPNFQPPNGP 139
P PN P N + PP P + RPP PP +P P +P+ PP P
Sbjct: 209 PTGLSGPNVNPIGPAPNNSPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPPPPP 266
Score = 34.3 bits (77), Expect = 0.87
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Query: 74 PQNPNFRPPQPPQNPNYRQQPPPQNPNFRPPPPPQNPNFRP 114
P P PP PP +P+ ++PP +P PPPP P+ P
Sbjct: 283 PPPPPPPPPPPPPSPSPPRKPPSPSPPV--PPPPSPPSVLP 321
>ACRO_RABIT (P48038) Acrosin precursor (EC 3.4.21.10)
Length = 431
Score = 56.2 bits (134), Expect = 2e-07
Identities = 34/84 (40%), Positives = 40/84 (47%), Gaps = 10/84 (11%)
Query: 44 VQQPPSDHNQ-HFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFR 102
V QPPS H +F H + P P +P+ RPPQPP Q PPP P
Sbjct: 310 VHQPPSAHPPWYFQHASGPPHPHPHPHP-----HPHPRPPQPPA----AQAPPPPPPPPP 360
Query: 103 PPPPPQNPNFRPPPPPQNPNFRQP 126
PPPPP P PPPPP + + P
Sbjct: 361 PPPPPPPPPPPPPPPPPPASTKPP 384
Score = 36.6 bits (83), Expect = 0.17
Identities = 26/70 (37%), Positives = 30/70 (42%), Gaps = 8/70 (11%)
Query: 75 QNPNFRPP---QPPQNPNYRQQPPPQNPNFRPPPPP--QNPNFRPPPPPQNPNFRQPIRQ 129
Q P+ PP Q P + P +P+ RPP PP Q P PPPPP P P
Sbjct: 312 QPPSAHPPWYFQHASGPPHPHPHPHPHPHPRPPQPPAAQAP---PPPPPPPPPPPPPPPP 368
Query: 130 NPNFQPPNGP 139
P PP P
Sbjct: 369 PPPPPPPPPP 378
Score = 36.2 bits (82), Expect = 0.23
Identities = 30/81 (37%), Positives = 33/81 (40%), Gaps = 11/81 (13%)
Query: 70 QDWNPQNPNFRPP---QPPQN--PNYRQQ---PPPQNPNFRPPP---PPQNPNFRPPPPP 118
Q P P R P QPP P Y Q PP +P+ P P PPQ P + PPPP
Sbjct: 296 QPATPTPPTTRSPGVHQPPSAHPPWYFQHASGPPHPHPHPHPHPHPRPPQPPAAQAPPPP 355
Query: 119 QNPNFRQPIRQNPNFQPPNGP 139
P P P PP P
Sbjct: 356 PPPPPPPPPPPPPPPPPPPPP 376
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 75,768,051
Number of Sequences: 164201
Number of extensions: 3843623
Number of successful extensions: 38377
Number of sequences better than 10.0: 1037
Number of HSP's better than 10.0 without gapping: 386
Number of HSP's successfully gapped in prelim test: 702
Number of HSP's that attempted gapping in prelim test: 18738
Number of HSP's gapped (non-prelim): 5754
length of query: 552
length of database: 59,974,054
effective HSP length: 115
effective length of query: 437
effective length of database: 41,090,939
effective search space: 17956740343
effective search space used: 17956740343
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148348.6


